- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Dimethylmethylene Blue Assay (DMMB)
Published: Vol 4, Iss 18, Sep 20, 2014 DOI: 10.21769/BioProtoc.1236 Views: 64562
Reviewed by: Anonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
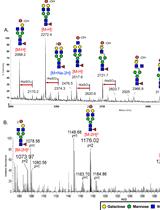

Negative Ion Mode nanoLC-ESI-MS/MS Analyses of Permethylated Sulfated Glycans
Shin-Yi Yu [...] Kay-Hooi Khoo
May 20, 2020 4399 Views
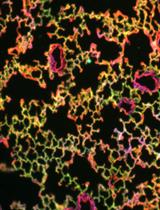
Bioorthogonal Labeling and Chemoselective Functionalization of Lung Extracellular Matrix
Zihan Ling [...] Xi Ren
Feb 20, 2021 4418 Views
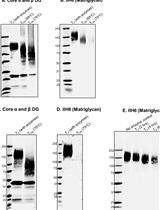
Identification of Matriglycan by Dual Exoglycosidase Digestion of α-Dystroglycan
Ishita Chandel and Kevin P. Campbell
Sep 20, 2023 1360 Views
Abstract
Glycosaminoglycans (GAGs) are long unbranched polysaccharides consisting of repeating disaccharide units composed of a hexosamine (glucosamine or galactosamine) and a hexuronic acid (glucuronic or iduronic acid). Depending on the disaccharide unit the GAGs can be organized into five groups: chondroitin sulfate, dermatan sulfate, heparan sulfate, keratan sulfate and hyaluronic acid. The GAGs are heterogeneous molecules with great variability in molecular mass and both sulfation density and pattern. Spectrophotometric assays to measure the GAG content in biological fluids and tissue/cell extracts are valuable tools. The dye 1,9-dimethylmethylene is a thiazine chromotrope agent that presents a change in the absorption spectrum due to the induction of metachromasia when bound to sulfated GAGs enabling rapid detection of GAGs in solution (Whitley et al., 1989; Chandrasekhar et al., 1987; Farndale et al., 1982). Moreover, there is a window in which a linear curve may be drawn (approximately between 0.5-5 μg of GAGs) enabling the quantification of GAGs in solution.
Keywords: Glycosaminoglycan quantificationMaterials and Reagents
- Dimethylmethylene blue (DMMB) (Sigma-Aldrich, catalog number: 341088 )
- NaCl
- Glycine (Sigma-Aldrich, catalog number: 410225 )
- Glacial acetic acid (Sigma-Aldrich, catalog number: S7653 )
- Tris-Base (Merck KGaA, catalog number: 648310 )
- Bovine chondroitin 4-sulfate as standard (Sigma-Aldrich, catalog number: C9819 )
- DMMB reagent (see Recipes)
Equipment
- Plate mixer (VWR International, catalog number: 89202-332 )
- Cover adhesive (R&D Systems, catalog number: DY992 )
- Microplate reader with 525 nm (BioTek Instruments, catalog number: 11-120-531 )
- 96 well microplate spectrophotometer with 525 nm filter set (Thermo Fisher Scientific, catalog number: 51119200 )
- Microplate shaker (VWR International, catalog number: 97043-608 )
Procedure
- Prepare DMMB reagent and paper filter using Whattman® 3MM. The pH of this solution is around 3.0. To prepare 1 L dye solution, dissolve 16 mg DMMB in 1 L water containing 3.04 g glycine, 1.6 g NaCl and 95 ml of 0.1 M Acetic Acid.
- Prepare standard solution of chondroitin 4 sulfate (500 µg/ml in H2O). Prepare standard curve as stated in the table bellow.
- Pipet the standard stock solution and complete the volume to 20 µl with H2O into the 96 well microplate.
- Pipet 20 µl of each sample into the microplate.
- Add 200 µl of DMMB to each sample and shake the plate of a plate shaker for 5 sec.
- Read the absorbance using a plate reader at 525 nm immediately.
Std (µg/ml)
Vol (µl) of 500 µg/ml std
vol H2O (µl)
vol DMMB (µl)
0
0
20
200
1.25
2.5
17.5
200
2.5
5
15
200
5
10
10
200
7.5
15
5
200
10
20
0
200
Notes
- DMMB assay can normally be performed on samples with high detergent and salt concentrations; however the standard curve should be prepared in the same solution.
- Avoid to performing the assay on samples in high albumin or serum concentrations which may interfere with the assay (Warren, 2000).
- Some groups have reported the interference of DNA in the DMMB assay, however, decreasing the pH to approximately 3 and increasing salt concentrations makes the interference of DNA negligible.
- DMMB requires the length of glycosaminoglycan chain be over a tetrasaccharide.
- DMMB reacts with the sulfate group of the GAG chain and therefore will not work with unsulfated GAGs such as hyaluronic acid.
- Further information can be acquired by utlilizing the carbazole reaction (sensitivity from 1 to 20 μg) to assay the carboxyl groups of the uronic acid for heparin sulfate, chondroitin sulfate and dermatan sulfate and/or anthrone reaction to assay the hexose group for keratan sulfate (Mort and Roughley, 2007).
- The DMMB can also be very efficient when performing chromatography to rapidly assay for fractions containing GAGs (Burton-Wurster et al., 2003).
- The DMMB-GAG complex that is formed results in the immediate formation of turbidity, however this complex starts to precipitate within 10 min, therefore the absorbance measurement should be performed immediately.
Recipes
- DMMB reagent
Dissolve 16 mg DMMB, 3.04 g glycine, 1.6 g NaCl and 95 ml of 0.1 M acetic acid and complete the volume to 1 L
Filter (0.45 μm)
Protect from light
Do not use if precipitate is present in the solution
Acknowledgments
This protocol was adapted or modified from Farndale et al. (1982) and Whitley et al. (1989).
References
- Anwari, K., Webb, C. T., Poggio, S., Perry, A. J., Belousoff, M., Celik, N., Ramm, G., Lovering, A., Sockett, R. E., Smit, J., Jacobs-Wagner, C. and Lithgow, T. (2012). The evolution of new lipoprotein subunits of the bacterial outer membrane BAM complex. Mol Microbiol 84(5): 832-844.
- Burton-Wurster, N., Liu, W., Matthews, G. L., Lust, G., Roughley, P. J., Glant, T. T. and Cs-Szabo, G. (2003). TGF beta 1 and biglycan, decorin, and fibromodulin metabolism in canine cartilage. Osteoarthritis Cartilage 11(3): 167-176.
- Chandrasekhar, S., Esterman, M. A. and Hoffman, H. A. (1987). Microdetermination of proteoglycans and glycosaminoglycans in the presence of guanidine hydrochloride. Anal Biochem 161(1): 103-108.
- Farndale, R. W., Sayers, C. A. and Barrett, A. J. (1982). A direct spectrophotometric microassay for sulfated glycosaminoglycans in cartilage cultures. Connect Tissue Res 9(4): 247-248.
- Mort, J. S. and Roughley, P. J. (2007). Measurement of glycosaminoglycan release from cartilage explants. Methods Mol Med 135: 201-209.
- Warren, S. (2000). A critical analysis of the 1, 9-dimethylmethylene blue assay for sulfated glycosaminoglycans in Synovial fluid. University of Guelph.
- Whitley, C., Ridnour, M., Draper, K., Dutton, C. and Neglia, J. (1989). Diagnostic test for mucopolysaccharidosis. I. direct method for quantifying excessive urinary glycosaminoglycan excretion. Clin Chem 35(3): 374-379.
Article Information
Copyright
© 2014 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Coulson-Thomas, V. J. and Gesteira, T. F. (2014). Dimethylmethylene Blue Assay (DMMB). Bio-protocol 4(18): e1236. DOI: 10.21769/BioProtoc.1236.
Category
Biochemistry > Carbohydrate > Glycoprotein
Biochemistry > Carbohydrate > Polysaccharide
Cell Biology > Cell staining > Carbohydrate
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










