Machine learning–assisted quantification of organelle abundance using Fiji
Speakers: Alexander James Long
Abstract
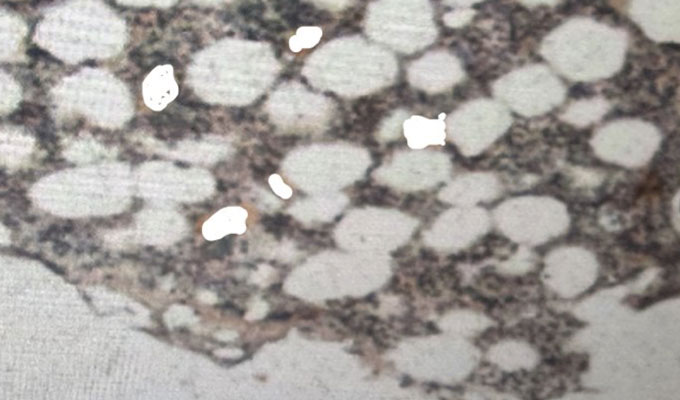
Organelle abundance is a key microscopic readout of organelle formation and, in many cases, function. Quantification of organelle abundance using confocal microscopy requires estimating their area based on the fluorescence intensity of compartment-specific markers. This analysis usually depends on a user-defined intensity threshold to distinguish organelle regions from the surrounding cytoplasm, which introduces potential bias and variability. To address this issue, we present a machine learning–assisted algorithm that allows for the quantification of organelle density using the open-source Fiji platform and WEKA segmentation. Our method enables the automated quantification of organelle number, area, and density by learning from training data. This standardizes threshold selection and minimizes user intervention. We demonstrate the utility of this approach for both membrane and non-membrane organelles, such as peroxisomes, lipid droplets, and stress granules, in human cells and whole fish samples.
Highlights:
• The organelle abundance algorithm is an automated, open-source, Fiji-based tool that extracts organelle number and area and calculates abundance based on a single marker.
• The macro measures the average intensity of all segmented areas and quantifies their area.
• The algorithm is applicable to cellular compartments, including membrane-bound and membrane-less organelles.
• Training is performed on a sample dataset, enabling the algorithm to be applied to all images obtained with the same imaging parameters.
Speaker

Alexander James Long, M.Sc.
Laboratory Technical Assistant, School of Biological Sciences at the University of Southampton, UK
Alexander James Long is currently a Laboratory Technical Assistant at the School of Biological Sciences at the University of Southampton, UK. He re...
View more
Keywords
Organelle, Peroxisome, Stress granule, Quantification, Abundance, Density, Machine learning, Fiji, WEKA segmentation, PeroxiSPY
References
Long, A. J., Candeias, D., Coveña, N. F., Reymond, L., Schuhmacher, M., Kemp, S., Hamilton, N. and Amen, T. (2026). Machine Learning-Assisted Quantification of Organelle Abundance. Bio-protocol 16(5): e5626. DOI: 10.21769/BioProtoc.5626.
Do you have a question about this webinar?
Post your question, and we'll invite the webinar speaker to respond. You're welcome to join the discussion by answering or commenting on questions ( Note: Not all questions, especially those not directly relevant to the webinar topic, may be answered by the speaker. ).
Write a clear, specific, and concise question. Don’t forget the question mark!
Tips for asking effective questions
+ Description
Write a detailed description. Include all information that will help others answer your question.
0 Q&A