- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Quantification of Bacterial Fatty Acids by Extraction and Methylation
Published: Vol 3, Iss 21, Nov 5, 2013 DOI: 10.21769/BioProtoc.950 Views: 15701
Reviewed by: Fanglian HeLin Fang

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

Crude Preparation of Lipopolysaccharide from Helicobacter pylori for Silver Staining and Western Blot
Hong Li and Mohammed Benghezal
Oct 20, 2017 10720 Views
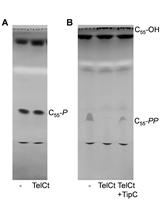

Bacterial Cell Wall Precursor Phosphatase Assays Using Thin-layer Chromatography (TLC) and High Pressure Liquid Chromatography (HPLC)
Manuel Pazos [...] Waldemar Vollmer
Mar 20, 2018 8353 Views
Analysis of Heterocyst and Akinete Specific Glycolipids in Cyanobacteria Using Thin-layer Chromatography
Ritu Garg [...] Iris Maldener
Mar 20, 2022 2487 Views
Abstract
This protocol describes two similar methods for the extraction and methylation of fatty acids from bacterial cultures. The acid derivatization protocol (Lennen et al., 2013; Bligh and Dyer, 1959) results in the extraction and methylation of all fatty acids, both free and bound, from a bacterial culture, while the base derivatization protocol (Lennen and Pfleger, 2013) captures only bound (phospholipid, acyl-thioester) species. After extraction into hexane, the lipids may be analyzed by gas chromatography.
Keywords: Free Fatty AcidMaterials and Reagents
- Cell culture for lipid analysis. When analyzing bacterial cultures expressing a thioesterase, it is advisable to wait until the late stationary phase (8-24 h for E. coli) to perform the extraction to allow sufficient time for product accumulation (Lennen et al., 2013)
- Methyl heptadecanoate (should be ≥ 99% purity) (e.g. Sigma-Aldrich, catalog number: 51633 )
- GC sample vials (VWR International, catalog number: 46610-722 ) (ensure you purchase vials compatible with any autosampler you are using)
- Compressed nitrogen gas/regulator/tubing
- Reverse osmosis water
- Methanol
- Absolute (100%) ethanol
- High resolution gas chromatography grade hexane (99.9% pure mixture of hexane isomers)
- Glacial acetic acid (Thermo Fisher Scientific, catalog number: A38-212 )
- Deionized water
- Anhydrous 1.25 M HCl in methanol (Sigma-Aldrich, catalog number: 17935 )
- Sodium bicarbonate
- 0.5 M Sodium methoxide in methanol (Sigma-Aldrich, catalog number: 403067 )
- Chloroform
- Appropriate internal standards (e.g. non-native, odd chain fatty acids). Some possibilities include:
- Heptadecanoic acid (C17:0) (99% + pure)
- Pentadecanoic acid (C15:0) (99% + pure)
- 1,2-dipentadecanoyl-sn-glycero-3-phosphoethanolamine in chloroform (99% + pure) (Avanti Polar Lipids, catalog number: 850704 )
- Heptadecanoic acid (C17:0) (99% + pure)
- External standards (the methyl esters of fatty acids you wish to quantify at a known concentration generally these may be obtained from a commercial source). A useful external standard mixture is sold by Supelco through Sigma-Aldrich (catalog number: 18918-1AMP )
- Antifoam 204 (Sigma-Aldrich, catalog number: A8311 )
Equipment
- 5 ml glass pipettes
- 10 ml Glass centrifuge tubes (Thermo Fisher Scientific, KimbleTM, catalog number: 73785-10 ) with fluroropolymer lined caps (Thermo Fisher Scientific, KimbleTM, catalog number: 73802-15415 )
- Gloves
- Goggles
- Appropriate personal protective equipment according to local regulations
- Chemical fume hood
- Centrifuge
- Vortex mixer
- Vacuum source/aspiration equipment
- Lyophilizer
- Water bath
- Mass spectrometer or flame ionization detector
- Agilent 7890 GC with a 30 m x 0.25 mm HP 5-ms capillary column
Procedure
- Collect 2.5 ml of cell culture in a glass centrifuge tube (this volume is sufficient for late exponential/stationary phase cultures of Escherichia coli).
Note: Most plastic tubes are not compatible with chloroform, which will be added in a subsequent steps.
Notes:- If you wish to normalize your measured fatty acid concentrations to cell density, record the OD600 at this point.
- If your culture is producing free fatty acids, a significant amount of foam may accumulate. It is necessary to collapse the foam, as a significant fraction of the free fatty acids present in the culture may be in the foam. This can be accomplished by the addition of antifoam followed by incubation in an 85 °C water bath for 5-10 min (Lennen et al., 2013). For example, 200 μl of a 1:10 dilution in ethanol of Antifoam 204 was added to collapse the foam of 50 ml shake flask cultures in Lennen et al. 2013.
- If you wish to normalize your measured fatty acid concentrations to cell density, record the OD600 at this point.
- To the cell culture, add ~50 μg of an appropriate internal standard. Over the course of this procedure, a fraction of this internal standard will be lost. From its known initial concentration and the final amount present, the measured concentration of each fatty acid methyl ester in the final extract can be used to determine the concentration of the corresponding acid in the culture. The choice of internal standard depends on which fatty acids are expected to be abundant in the samples to be analyzed. Often cells do not produce large amounts of odd-chain fatty acids. For example, E. coli grown on glucose does not produce large amounts of odd chain fatty acids, so heptadecanoic acid (C17:0) can be used as an internal standard.
- If measuring membrane fatty acids or long chain free fatty acids (C16-C18 species), 5 μl of 10 mg/ml heptadecanoic acid dissolved in ethanol is a useful internal standard.
- If the strain of interest is producing large quantities (0.5 g/L or higher) of medium chain free fatty acids (Lennen et al., 2013; Youngquist et al., 2012; Lenen et al., 2010), 50 μl of 10 mg/ml pentadecanoic acid (C15:0) dissolved in ethanol is a useful internal standard.
- If only analyzing phospholipid species, a phospholipid internal standard, such as 1,2-dipentadecanoyl-sn-glycero-3-phosphoethanolamine in chloroform, can be used instead of a free fatty acid standard (Lennen and Pfleger, 2013).
- Note that all values given for the volume/concentration of the internal standards may need to be adjusted for your particular system, and different standards may need to be used depending on the chain lengths of the fatty acids to be quantified. An internal standard should mimic the compounds to be quantified as closely as possible and should be added to the collected sample of culture in an amount which approximates (same order of magnitude) the concentration of the compounds it will be used to analyze.
- 5 μl of 10 mg/ml will correspond to a theoretical final concentration of 50 μg/ml methyl heptadecanoate if two 0.5 ml hexane extractions are performed in step 14. This is around the same order of magnitude as the C14:0, C16:0, and C16:1 fatty acids that are recovered from the membranes of wild-type E. coli.
- If measuring membrane fatty acids or long chain free fatty acids (C16-C18 species), 5 μl of 10 mg/ml heptadecanoic acid dissolved in ethanol is a useful internal standard.
- Add 100 μl of glacial acetic acid to acidify the culture. Carefully vortex the sample to mix. This should be performed under a chemical fume hood.
- Add 5 ml of 1:1 mixture by volume of chloroform and methanol with a glass pipette (chloroform will leach a variety of compounds out of a plastic pipette). Vortex thoroughly.
Notes:- This mixture may be stored at -80 °C and the extraction (detailed below) finished at a later date if desired.
- If samples are frozen, each should be thawed to room temperature before moving on to step 5.
- This mixture may be stored at -80 °C and the extraction (detailed below) finished at a later date if desired.
- Centrifuge each sample for 10 minutes at 1,000 x g.
- Using a vacuum line or aspirator, remove the upper aqueous layer and all cell debris at the interface.
Notes:- It is acceptable to aspirate a small amount of bottom chloroform layer if internal standards are present.
- Prior to aspiration, ensure that proper receptacles are available for the waste generated by this process.
- The chloroform layer can be stored at -80 °C at this point if necessary.
- This step should be performed under a chemical fume hood!
- It is acceptable to aspirate a small amount of bottom chloroform layer if internal standards are present.
- Evaporate the chloroform extract under a nitrogen stream, leaving a dried residue in the tubes.
Notes:- There is usually some residual water that is difficult to remove after this step.
- If tubes are being thawed out of the -80 °C freezer, you should re-aspirate any accumulated water droplets before proceeding to the evaporation step.
- This step should be performed under a chemical fume hood!
- There is usually some residual water that is difficult to remove after this step.
- Lyophilize the residue (30-60 min is sufficient) to remove any remaining water. All subsequent steps need to be performed with anhydrous samples.
For Total Fatty Acid Extraction (Free Fatty Acids and Bound Fatty Acid Species by Acid Catalysis): - To the dried extract, add 0.5 ml of anhydrous 1.25 M HCl in methanol. Cap tightly and heat at 50 °C overnight.
Notes:- Alternatively, the procedure can be performed at 75-80 °C for 1 h.
- The elevated temperature builds up pressure in the sealed vial, so ensure you have tubes capable of withstanding these conditions if you do this (those described in the equipment section are adequate).
- This step should be performed under a chemical fume hood!
- Alternatively, the procedure can be performed at 75-80 °C for 1 h.
- Cool tubes to room temperature.
- Add 0.5 ml of high resolution gas chromatography grade hexane. Take care to work rapidly to recap the tubes as hexane is highly volatile. This step should be performed under a chemical fume hood!
- Add 5 ml of a 100 mg/ml NaHCO3 aqueous solution (this concentration is close to the solubility limit–it will dissolve after stirring overnight or warming the solution gently). Addition of bicarbonate quenches the acid-catalyzed reaction. This step should be performed under a chemical fume hood!
For Bound (Phospholipid) Fatty Acid Extraction by Base Catalysis: - To the dried chloroform extract, add 0.5 ml of 0.5 M sodium methoxide in methanol. This step should be performed under a chemical fume hood!
- Incubate the reaction for 10 min at 50 °C, which is sufficient for trans-esterification of common phosphoplipids such as those found in E. coli.
Notes:- This reaction can also be used to trans-esterify other bound fatty acids, for example those found in triacylglycerols.
- For guidelines on the incubation time for these reactions, which vary from the time required for phospholipids, consult a resource such as Lipid Analysis by William Christie (Christie and Han, 2010). Do not allow the reaction to proceed longer than necessary as too long a reaction time, and the presence of water, can result in esterification of free fatty acids.
- The use of a free fatty acid internal standard as described previously can inform on whether only bound fatty acids are being methylated.
- This step should be performed under a chemical fume hood!
- This reaction can also be used to trans-esterify other bound fatty acids, for example those found in triacylglycerols.
- Cool the tubes to room temperature. Quench the reaction by adding 0.1 ml of glacial acetic acid followed by 5 ml of deionized water. This step should be performed under a chemical fume hood!
- Add 0.5 ml of high resolution gas chromatography grade hexane. This step should be performed under a chemical fume hood!
For Both Bound and Total Fatty Acid Extractions: - Vortex tubes thoroughly and centrifuge at room temperature for 10 minutes at 1,000 x g to create a stable interface. Note that the aqueous phase should be clear before proceeding.
- Collect 0.4 ml of the hexane layer from the first extraction in an appropriate gas chromatograph vial. Be sure to cap the vial to prevent sample evaporation. This step should be performed under a chemical fume hood!
- Add 0.5 ml of high resolution gas chromatography grade hexane to the methanol:water extract for a second extraction. Vortex thoroughly to mix and centrifuge at room temperature for 10 minutes at 1,000 x g to create a stable interface. This step should be performed under chemical fume hood!
- Collect 0.5 ml of the top hexane layer and add to the hexane collected in step 14. The collected hexane layers are ready for GC analysis.
Notes:- Samples may need to be diluted in hexane if fatty acid concentrations are outside the range of standard calibration curves.
- This step should be performed under chemical fume hood!
- Samples may need to be diluted in hexane if fatty acid concentrations are outside the range of standard calibration curves.
- Analyze by gas chromatography with either a mass spectrometer or flame ionization detector. It is wise to randomize the run order of your samples to prevent the introduction of bias into your analysis. The following is an example of a typical gas chromatography method for bacterial lipids using an Agilent 7890 GC with a 30 m x 0.25 mm HP 5-ms capillary column (Lenen et al., 2010):
- Inject 1 μl using a 1:10 split ratio of helium carrier gas (a lower split ratio, e.g. 1:100, can be used for more abundant species).
- Oven temperature of 100 °C for 2 minutes.
- Oven temperature of 150 °C for 4 minutes.
- Ramp to 250 °C at a rate of 4 °C/min.
- Inject 1 μl using a 1:10 split ratio of helium carrier gas (a lower split ratio, e.g. 1:100, can be used for more abundant species).
- To determine the concentrations of the fatty acid methyl esters of interest in your chromatography samples, you will need to run samples of these compounds at a variety of known concentrations (your external standards) spanning the full range of concentrations of each compound in the unknown samples. After running these samples, you will have a set of peak areas and corresponding concentrations for each compound. You may then construct a curve of best fit for each compound (including your internal standard), and calculate the concentration of each fatty acid methyl ester and the internal standard in your samples. These concentrations should be multiplied by the ratio of the theoretical to the actual internal standard concentrations to correct for sample lost during the extraction and methylation process. The concentrations in the bacterial culture may then be determined. Keep in mind that the concentration in units of mass for a fatty acid methyl ester and needs to be corrected to report the concentration of the corresponding fatty acid. It is wise to work in molar units during the analysis. As a rule of thumb, you should expect R2 values of about 0.98 to 0.99 or higher from your calibration curves.
- Lipids were extracted from a bacterial culture expressing a plant thioesterase specific for 12 carbon (C12) saturated acyl-ACPs, resulting in the overproduction of C12 free fatty acids and, to a lesser extent, C14 free fatty acids. After adding 510 μg of pentadecanoic acid (C15) to 2.5 ml of a late stationary phase culture and performing the total fatty acid extraction protocol by acid catalysis as described above, the resulting FAMEs were analyzed by GC-MS. A total of 1 ml of hexane was used in the extraction. The amount of C12, C14, and C15 FAMEs present in the chromatography sample was determined by constructing calibration curves relating the peak area of each species to its concentration (column 2 of Table 1). If the conversion of C15 acid to its methyl ester and its subsequent extraction were perfect, there would have been 539.5 μg/ml of C15 methyl ester in the chromatography sample (calculated from the listed molecular weights). This fact was used to adjust the concentratiosn of C12 and C14 FAMEs (the measured concentrations were multiplied by the ratio 539.5/484.3). These corrected concentrations were then used to determine the fatty acid titer in the culture by converting to the acid concentration using the molecular weights and then dividing by the volume of the culture sample (2.5 ml).
Table 1. The titers of C12 and C14 fatty acids in a thioesterase expressing culture were measured by GC-MS using pentadecanoic acid (C15:0) as an internal standard. The concentration refers to the concentration in the sample analyzed by chromatography while titer refers to the concentration in the culture. Molecular weights were obtained from the NIST Chemistry Webbook.
Notes
- Chloroform is toxic and should only be used under a fume hood. If chloroform comes into contact with latex or nitrile gloves, remove gloves immediately. Alternatively, use polyvinyl alcohol or SilverShield gloves when working with chloroform.
- This procedure uses corrosive (glacial acetic acid and anhydrous hydrochloric acid in methanol), water-sensitive as well as flammable (methanol and hexane) chemicals. Sodium methoxide in methanol is flammable and highly toxic by inhalation, ingestion, and contact with skin. All should be handled according to common chemical safety practices.
Acknowledgments
The protocol described here is based on the work of Bligh and Dyer (1959). The protocol was developed with funding from the US Department of Energy, Great Lakes Bioenergy Research Center.
References
- Bligh, E. G. and Dyer, W. J. (1959). A rapid method of total lipid extraction and purification. Can J Biochem Physiol 37(8): 911-917.
- Christie, W. W. and Han, X. (2010). Lipid analysis: Isolation, separation, identification and lipidomic analysis, Oily Press Bridgewater, UK.
- Lennen, R. M. and Pfleger, B. F. (2013). Modulating membrane composition alters free fatty acid tolerance in Escherichia coli. PLoS One 8(1): e54031.
- Lennen, R. M., Politz, M. G., Kruziki, M. A. and Pfleger, B. F. (2013). Identification of transport proteins involved in free fatty acid efflux in Escherichia coli. J Bacteriol 195(1): 135-144.
- Lenen, R. M., Braden, D. J., West, R. A., Dumesic, J. A. and Pfleger, B. F. (2010). A process for microbial hydrocarbon synthesis: Overproduction of fatty acids in Escherichia coli and catalytic conversion to alkanes. Biotechnol Bioeng 106(2): 193-202.
- NIST Chemistry WebBook, NIST Standard Reference Database Number 69, Linstrom, P. J. and Mallard, W.G. (eds.) http://webbook.nist.gov/chemistry.
- Youngquist, J. T., Lennen, R. M., Ranatunga, D. R., Bothfeld, W. H., Marner, W. D., 2nd and Pfleger, B. F. (2012). Kinetic modeling of free fatty acid production in Escherichia coli based on continuous cultivation of a plasmid free strain. Biotechnol Bioeng 109(6): 1518-1527.
Article Information
Copyright
© 2013 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Politz, M., Lennen, R. and Pfleger, B. (2013). Quantification of Bacterial Fatty Acids by Extraction and Methylation. Bio-protocol 3(21): e950. DOI: 10.21769/BioProtoc.950.
Category
Biochemistry > Lipid > Lipid isolation
Microbiology > Microbial biochemistry > Lipid
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link