- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
In vitro and in vivo Assessment of Protein Acetylation Status in Mycobacteria
Published: Vol 9, Iss 13, Jul 5, 2019 DOI: 10.21769/BioProtoc.3291 Views: 5679
Reviewed by: Kristin L. ShinglerDarrell CockburnGunjan Mehta

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
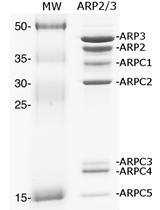
ARP2/3 Phosphorylation Assay in the Presence of Recombinant Bacterial Effectors
Céline Michard [...] Patricia Doublet
Apr 5, 2017 9184 Views
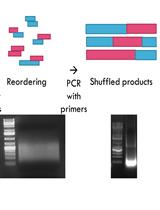
Characterization of Protein Domain Function via in vitro DNA Shuffling
Kathy Hiu Laam Po [...] Sheng Chen
Jun 5, 2018 6643 Views
In vitro Glutamylation Inhibition of Ubiquitin Modification and Phosphoribosyl-Ubiquitin Ligation Mediated by Legionella pneumophila Effectors
Alan G. Sulpizio [...] Yuxin Mao
Nov 5, 2020 3944 Views
Abstract
Protein acetylation is one of the standard post-translational modifications found in proteins across all organisms, along with phosphorylation which regulates diverse cellular processes. Acetylation of proteins can be enzymatically catalyzed through acetyltransferases, acetyl CoA synthetases or non-enzymatically through acyl carrier metabolic intermediates. In this protocol, using response regulator proteins as targets we describe the experimental strategy for probing the occurrence of acetylation using purified recombinant proteins in an in vitro setup. Further using M. smegmatis strains overexpressing the wild type or mutant response regulator protein, we also describe how in vivo acetylation can be validated in Mycobacterial proteins. The described approach can be used for analyzing acetylation of any mycobacterial protein under both in vitro and in vivo conditions.
Keywords: TcrXBackground
Lysine acetylation is a typical post-translational modification (PTMs) found to be present in proteins across all living organisms. It involves covalent attachment of an acetyl group from acetyl donor, e.g., acetyl phosphate, acetyl CoA, acetate, acetyladenylate, etc. on to the ε-NH2 group of the amino acid lysine in an acceptor protein. Protein acetylation has been extensively studied in context to the histone modification, and for many transcription factors and is associated with regulating chromatin remodeling to cell signaling, antibiotic resistance, environmental stress survival, and metabolism.
While many recent studies have revealed the presence of acetylated proteins in prokaryotes using global proteome analysis approaches, there have been limited attempts to study the impact of acetylation on the function of identified protein due to multistep validation needed for acetylated proteins. For Mycobacterium tuberculosis, which is a slow-growing bacterium, development of highly sensitive mass spectrometry approaches facilitated total acetylome analysis (Liu et al., 2014; Xie et al., 2015), which included many signaling proteins, thus warranting detailed mechanistic analysis of the impact of the identified modification. Recently, the effect of acetylation on the activity of two-component signaling protein DosR in the hypoxic response of Mycobacterium tuberculosis was reported (Bi et al., 2018; Yang et al., 2018). And more recently we demonstrated that acetylation of the response regulator TcrX alters the crosstalk known to be present in two-component signaling systems of M. tuberculosis as well as phosphatase activity of the sensor kinase and DNA binding activity of the response regulator (Singh et al., 2019). During the course of this study, we developed an optimized protocol for in vitro as well as in vivo acetylation analysis of various target proteins which is described here, using the response regulator TcrX as a template. This protocol can be used for testing acetylation status of any mycobacterial protein and involves two steps, first, where the presence of acetylation is probed in vitro in an enzymatically catalyzed reaction and second, the protein is probed for acetylation presence in vivo using M. smegmatis mc2155 as a surrogate host. Utilization of a protein carrying a Lys substitution mutation in vivo helps confirm that the identified Lys is indeed the target acetylation site.
Materials and Reagents
- Pipette tips
- 1 ml tip (Tarsons, catalog number: 521020)
- 2-200 µl (Tarsons, catalog number: 521010)
- 0.2-10 µl (Tarsons, catalog number: 521000)
- 1.5 ml microcentrifuge tubes (Tarsons, catalog number: 500010)
- 15 ml polypropylene centrifuge tubes (Tarsons, catalog number: 546021)
- 50 ml polypropylene centrifuge tubes (Tarsons, catalog number: 546041)
- 0.1 mm diameter zirconia/silica beads (Thomas Scientific, catalog number: 3411F09)
- 2 ml polypropylene bead beating vials (Bio Spec Products Inc., catalog number: 10831)
- PVDF membrane (Bio-Rad, catalog numbers: 1620177 or 1620238)
- Whatman filter paper 1.5 MM (GE Healthcare, catalog number: 10426981)
- 0.45 µm filters (Sartorius Minisart syringe filter, catalog number:16555-K)
- 96-well Micro test plate (Tarsons, catalog number: 941196)
- Expression vector with affinity tag for recombinant protein overexpression and purification (pProEx-Ht vector contains 6x His-tag at the N-terminal has been used for overexpression and purification of TcrX response regulator protein (Agrawal et al., 2015) as well as KATms (acetyl transferase) was a kind gift from Prof. Sandhya Visweswariah, IISc (Nambi et al., 2010)
- Expression strain E. coli BL21 Arctic ExpressTM [B F– ompT hsdS(rB– mB–) dcm+ Tetr gal λ(DE3) endA Hte [cpn10 cpn60 Gentr] (Ferrer et al., 2003) (Agilent Technologies, USA, catalog number: 230191) used for overexpression of TcrX and SP850cyc- strain for KATms as reported (Nambi et al., 2010)
- Mycobacterium smegmatis mc2155 (ATCC, catalog number: 700084)
- pMV261 vector (Received as a kind gift from Prof. Dipankar Chatterji, IISc)
- pMV261 containing a gene coding for TcrX (Singh et al., 2019) or your gene of interest
- pMV261 containing a gene coding for TcrX K231R (Singh et al., 2019) or your gene of interest
- Middlebrook 7H9 medium (BD Biosciences, catalog number: 271310)
- Glycerol (Sisco Research Ltd., catalog number: 62417)
- Tween 80 (Sigma-Aldrich, catalog number: P4780)
- Tween 20 (Sigma-Aldrich, catalog number: P9416)
- Tyloxapol (Sigma-Aldrich, catalog number: T8761)
- Kanamycin (Goldbio, catalog number: K-120)
- Oleic acid (Sigma-Aldrich, catalog number: 75090)
- Albumin (Amersco, catalog number: 0332)
- Dextrose (Sisco Research Ltd., catalog number: 51758)
- Catalase (Sigma-Aldrich, catalog number: C9322)
- Coomassie Brilliant Blue G-250 (Sisco Research Ltd., catalog number: 64222)
- Methanol (Sisco Research Ltd., catalog number: 96446)
- Phosphoric Acid (Alfa Aesar, catalog number: A18067)
- Hydrochloric acid (HCl) (Fisher Scientific, catalog number: A144SI-212)
- Sodium chloride (NaCl) (Merck Millipore Corporation, catalog number: 1064040500)
- Potassium chloride (KCl) (Merck Millipore Corporation, catalog number: 1049360500)
- Di-sodium hydrogen phosphate (Merck Millipore Corporation, catalog number: 1065860500)
- Potassium dihydrogen phosphate (KH2PO4) (Merck Millipore Corporation, catalog number: 1048730250)
- Trizma base (Sigma-Aldrich, catalog number: T6066)
- 2-Mercaptoethanol (Sigma-Aldrich, catalog number: M6250)
- Phenylmethylsulfonyl fluoride (Sigma-Aldrich, catalog number: P7626)
- Benzamidine hydrochloride (Sigma-Aldrich, catalog number: B6506)
- 1% Nonidet-P40 (Sigma-Aldrich, CAS: 9016-45-9)
- Anti-acetyl antibody (Cell Signaling Technology Inc., catalog number: 9441, Dilution 1:7,500)
- HRPO-conjugated secondary antibody (Cell Signaling Technology Inc., catalog number: 7074, Dilution 1:5,000)
- Anti-His antibody (Cell Signaling Technology Inc., catalog number: 2365, Dilution 1:5,000)
- Anti-TcrX polyclonal antibody (Dilution 1:1,000) (generated in the laboratory [Singh et al., 2019])
- Western Lightning PLUS ECL containing enhanced luminol reagent and oxidizing reagent (PerkinElmer, catalog number: NEL100001EA)
- SDS (Affymetrix USB, catalog number: 18220)
- EDTA (Sisco Research Ltd., catalog number: 40648)
- Bromophenol blue (MERCK, catalog number: 1081220005)
- cAMP (Sigma-Aldrich, catalog number: A6885)
- Acetyl CoA (Sigma-Aldrich, catalog number: A2056)
- ADC solution (Albumin, Dextrose, Catalase) (see Recipes)
- 10x acetylation Buffer (see Recipes)
- cAMP, 5 mM stock (see Recipes) (Sigma-Aldrich, catalog number: A6885)
- Acetyl CoA, 5 mM stock (see Recipes) (Sigma-Aldrich, catalog number: A2056)
- 7H9-Glycerol-Tween 80 medium (see Recipes)
- Recovery Media (see Recipes)
- Phosphate buffered saline (see Recipes)
- Lysis buffer (see Recipes)
- 5x Bradford Reagent (see Recipes)
- Storage buffer (see Recipes)
- SDS-PAGE running buffer (see Recipes)
- Transfer buffer (see Recipes)
- Tris Buffer Saline Tween 20 (TBS-T) (see Recipes)
- Stripping Buffer (see Recipes)
- 5x Protein sample buffer (see Recipes)
Equipment
- Pipettes
a.100-1,000 µl pipette (Rainin pipet-lite, catalog number: SL-1000)
b.20-200 µl pipette (Rainin pipet-lite, catalog number: SL-200)
c.2-20 µl pipette (Rainin pipet-lite, catalog number: SL-20)
d.0.1-2 µl pipette (Rainin pipet-lite, catalog number: SL-2) - 2 mm electroporation cuvette (Bio-Rad Laboratories, catalog number: 165-2086)
- Mini-Protean gel electrophoresis system (Bio-Rad Laboratories, model: 165-8000)
- Mini Trans-Blot cell (Bio-Rad Laboratories, model: Trans-Blot® SD cell)
- Gel Doc Imaging System (Bio-Rad Laboratories, model: Universal Hood III)
- 37 °C incubator/shaker (N-BIOTEK, model: NB-205L)
- Refrigerated centrifuge (Thermo Fisher Scientific, model: Sorvall Legend X1R; Hettich Zentrifugen, model: Rotina 420R)
- Spectrophotometer (Eppendorf, BioPhotometer®, model: D30)
- Bead beater (Bio Spec Products, model: Mini-Bead beater)
- Microcentrifuge (Tomy Kogyo Co. Ltd., model: Kitman-T24)
- Vortexes (Shalom Instruments, model: SLM-VM-3000)
- Heating block (heats to 95 °C) (Biobee Lab)
- Ultra-low temperature freezer (Sanyo, model: MDF-U700VX)
- Electroporator (Eppendorf Eporator, catalog number: 4309000019)
- Multimode microplate reader (Tecan, model: Infinite M1000 PRO)
- Autoclave
- -80 °C freezer
Software
- ImageJ https://imagej.nih.gov/ij/download.html
- Image lab version 5.2.1 software from BIO-RAD (used in ChemiDocTM Gel Documentation system)
Procedure
- In vitro acetylation analysis
The acetylation status of TcrX protein (as a representative) is assessed by probing the protein using an anti-acetyl antibody by Western blot analysis, which is discussed in a stepwise manner in this protocol. Response regulator TcrX (6x his-tagged) and mutant TcrX (K231R) proteins are overexpressed and purified from E. coli BL21 Arctic ExpressTM and KATms (acetyltransferase/MSMEG_5458) from SP850cyc-strain, as previously reported in Singh et al. (2019). The purified proteins are quantitated through Bradford’s protein estimation assay (Stoscheck, 1990; He, 2011a) and the proteins are stored in storage buffer (-20 °C; see Recipes) until use. The protocol below describes the experimental procedure subsequent to protein purification.
Setting up in vitro acetylation reaction followed by anti-acetyl Western blotting:- Mix the following ingredients in the following amounts:
TcrX or TcrX K231R proteins, 3 µg
10x acetylation buffer, 2 μl (see Recipes)
cAMP, 0.4 μl (see Recipes for stock concentration)
Acetyl CoA, 0.4 µl (see Recipes for stock concentration)
KATms, 0.2 µg
Add autoclaved MQ water to 20 µl - Incubate at 37 °C for 4 h (the incubation duration might vary between substrate proteins linked to extent of acetylation).
- Terminate reactions by diluting the reaction mixture with 5 µl of sample buffer (see Recipes). Heat the sample at 95 °C for 10 min.
- Resolve samples using 15% SDS-PAGE (He, 2011b) as per standard protocol at 100 V for 3-4 h.
- Transfer the protein to PVDF membrane by electroblotting using the Bio-Rad Mini Trans-Blot cell, as per the manufacturer’s protocol at 10 V for 1 h.
Notes:- Cut the PVDF membrane from the PVDF roll according to someone’s SDS-PAGE gel size and lanes loaded or use precut preassembled blotting membrane/filter paper sandwich from Bio-Rad.
- Transfer of proteins onto PVDF membrane takes 10 V for 1 h for this particular low molecular protein (~30 kDa) in a semidry transfer apparatus (Trans-Blot® SD cell) of Bio-Rad. It should be optimized according to someone’s protein of interest (high or low molecular weight) for voltage, current, timing and type of apparatus (Dry/Semi dry/Wet) to get an efficient transfer.
- Block the PVDF membrane with 5% BSA in 1x TBST (30 ml) for 90 min with gentle rocking at room temperature.
- Incubate with primary anti-acetyl lysine antibody diluted at 1:7,500 for overnight at 4 °C with gentle rocking.
- Wash the membrane 3 times with 1x TBST (50 ml) for 10 min each with rapid rocking.
- Incubate with 1:5,000 diluted secondary antibody for 1 h at room temperature with gentle rocking.
- Wash the membrane again 3 times with 1x TBST (60-70 ml) for 10 min each with rapid rocking.
- Develop the blot using enhanced chemi-luminescence (ECL) as per the manufacturer’s protocol and image the blot in a Chemidoc gel documentation system (Figure 1).
- After developing of PVDF membrane wash the membrane with 1x TBST (50 ml) 3 times for 10 min each.
- Incubate the PVDF membrane at 55 °C with stripping buffer (30 ml; see Recipes) for 25-30 min (optimize accordingly).
- Pour off the stripping buffer and wash the PVDF membrane 6-7 times with 1x TBST (100 ml) with rapid rocking for 10 min each or β-mercaptoethanol is completely removed.
Note: Do check for the typical smell of β-mercaptoethanol from PVDF membrane to ensure that it has been completely removed after washing with 1x TBST buffer. Repeat washing until it goes off completely. - Block the PVDF membrane with 5% BSA in 1x TBST (30 ml) for 90 min at room temperature.
- Repeat the steps from Step A7 (use anti-his 6x his antibody at a dilution of 1:5,000) until Step A10 for developing the blot and acquire the image as specified in Step A11 (Figure 1). This step provides a reference image for the amount of protein loaded.
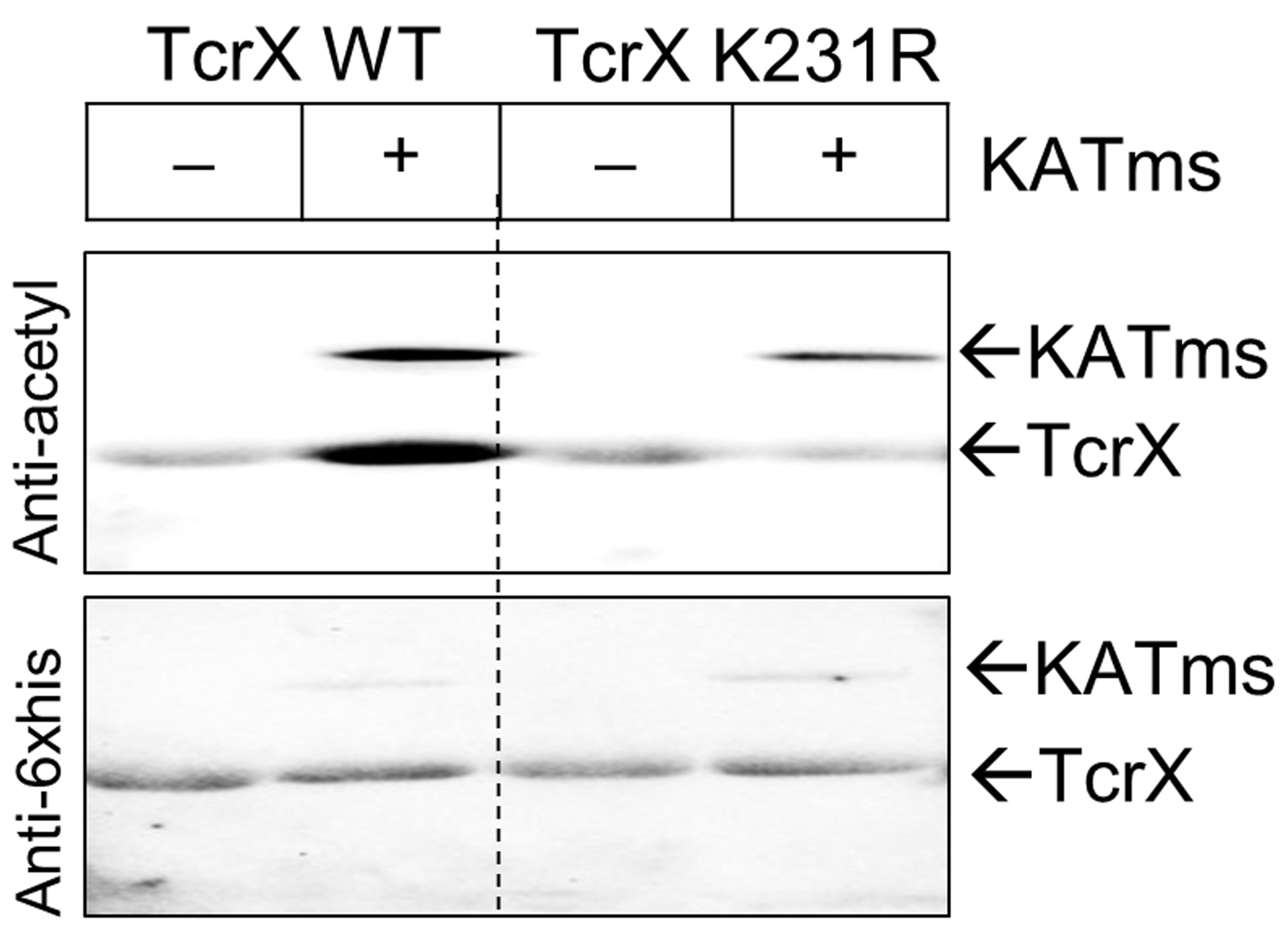
Figure 1. In vitro acetylation assessment. RR proteins TcrX and TcrXK231R were incubated with KATms (acetyltransferase, MSMEG_5458enzyme from M. smegmatis) as per the protocol. Acetylation site defective TcrX protein was generated after site identification by mass spectrometry, as reported in Singh et al. (2019). Top, an image of blot probed with anti-acetyl lysine antibody and bottom, anti-His antibody.
- Mix the following ingredients in the following amounts:
- In vivo acetylation analysis of TcrX protein
In vivo assessment of acetylation of a specific protein requires it to be overexpressed in Mycobacterial cells. Towards this, the typical approach involves cloning and expression of the ORF under a strong mycobacterial promoter like hsp60 or msp12 in a mycobacterial shuttle vector. Introduce the plasmid in Mycobacterial strains such as M. smegmatis and then analyze the expression or modification status of the overexpressed protein. The following steps describe the entire workflow towards that.- Preparation for electrocompetent Mycobacterium smegmatis mc2155
Mycobacterium smegmatis mc2155 strain is made electrocompetent to introduce the recombinant pMV261 mycobacterial plasmids carrying a gene of interest.- Prepare 1% primary culture by inoculating 50 µl of Mycobacterium smegmatis mc2155 stock in 5 ml of Middlebrook 7H9 media containing 0.2% glycerol, 0.05% Tween-80 and 10% ADC.
- Incubate the culture at 37 °C in the shaker at 180 rpm until OD600 ~2 is obtained (approximately 2 days)
- Inoculate 1 ml of the primary culture to 100 ml of 7H9 broth supplemented with 0.2% glycerol, 0.05% Tween-80 and 10% ADC and grow at 37 °C in the shaker at 180 rpm until the OD600 of 0.8-1.0 (~24-32 h).
- Pellet the cells at 4,500 x g, 4 °C for 15 min, followed by 3-4 washes in 20 ml of autoclaved MilliQ water.
- This is followed by 3 washes with 20 ml of sterile 10% glycerol.
- Pellet the cells and resuspend in 2 ml of sterile 10% glycerol.
- Prepare aliquots of 200 μl each for electroporation in the 1.5 ml centrifugation tubes.
- Spot freeze the remaining aliquots of electrocompetent cells in 1.5 ml centrifugation tubes using pre-chilled 100% ethanol and store them at -80 °C until further use.
Note: Keep a container filled with 100% ethanol at -80 °C for 6-8 h prior to start the preparation of electrocompetent cells. When the electrocompetent cells aliquots are ready in microcentrifuge tubes then dip the tubes in the container containing the prechilled ethanol before storage at -80 °C.
- Electroporation of plasmid DNA in M. smegmatis mc2155
- To an aliquot of electrocompetent cells add 1 µg of the recombinant plasmids and incubate on ice for 2 min.
- Prechill 2 mm electroporation cuvette on ice, add electrocompetent cells and the plasmid mixture into the cuvette and incubate on ice for 10 min. Wipe the outer walls of the cuvette and pulse at 2,500 V using an electroporator and immediately keep on ice.
Note: At the end of electroporation, ~4-5 millisecond time can be observed on display of electroporator which good indication of correct current flow between the two parallel electrodes and efficient electroporation. If arcing takes place, timing is in seconds on display and set voltage differs from applied voltage on the cells during the electroporation. This is possibly due to high ionic strength of buffer, high salt or insufficient washing of cells which ultimately results in ineffective or no electroporation. - Add 200 µl recovery media (see Recipes) to the cuvette and transfer cells to a 15 ml conical tube. Add more media to bring the volume to 3 ml and shake the tube for 4 h at 37 °C at 180 rpm in shaker incubator for recovery of cells.
Note: Keep the recovery media (see Recipes) in excess (20 ml) ready before electroporation. - Harvest the cells at 4,500 x g for 10 min at room temperature. Discard the supernatant and resuspend pellet in 100 µl of recovery media.
- Plate cells on 7H11 media (agar base) supplemented with 0.2% glycerol, 0.05% Tween 80, 10% ADC containing appropriate antibiotics (here 25 µg/ml kanamycin).
- Incubate the plate at 37 °C for 2 days to obtain transformants.
- Lysate preparation and in-vivo expression and acetylation status analysis
- For starter culture, inoculate single colony each of the recombinant strains in 5 ml of 7H9 media containing 0.2% glycerol, 0.05% Tween 80 and 10% ADC along with 25 µg/ml kanamycin and incubate it at 37 °C at 180 rpm in a shaker for 2 days.
- Subculture 0.5-1 ml of the primary culture in 50 ml of 7H9 broth supplemented with 0.2% glycerol, 0.05% tyloxapol and 10% ADC along with 25 µg/ml kanamycin and grow at 37 °C in the shaker at 180 rpm.
Note: When a large volume of secondary culture (e.g., 1/2 liter) is needed, primarily to get good amount of protein, primary culture should also be of a larger volume (10-50 ml). - Once OD600 reaches ~2-3 (approximately 2-3 days), pellet the cells at 4 °C at 4,500 x g, wash 3 times each with 10 ml of sterile PBS and resuspend pellet in 500 µl of Tris lysis buffer (pH ~7.5).
- Transfer the suspension in 2 ml bead beating vial containing 0.1 mm diameter zirconia/silica beads (~250 µl volume of buffer) and bead beat for 30 s, 8-10 times with intermittent incubation on ice (optimize according to the instructions from the bead homogenizer manufacturer).
- Centrifuge the lysate for 10 min at 4 °C at 10,000 x g and collect ~400 µl of supernatant. Care should be taken to avoid collecting the unlysed cells and debris.
- Quantify the lysate by Bradford’s protein estimation assay.
- Resolve 100 µg of protein lysate by 15% SDS-PAGE as per standard protocol.
- Transfer the resolved protein to PVDF membrane by electroblotting as per the manufacturer’s protocol and proceed for Western blotting with anti-acetyl lysine antibody as described in the section above, followed by stripping and probing with protein-specific antibody (here anti-TcrX).
- Figure 2 depicts a representative developed blot reporting the overexpression of candidate response regulator protein TcrX and its acetylation status through specific antibodies. The blot also shows an absence of acetylation when a lysine mutant protein is used, confirming the site which undergoes acetylation in vivo.
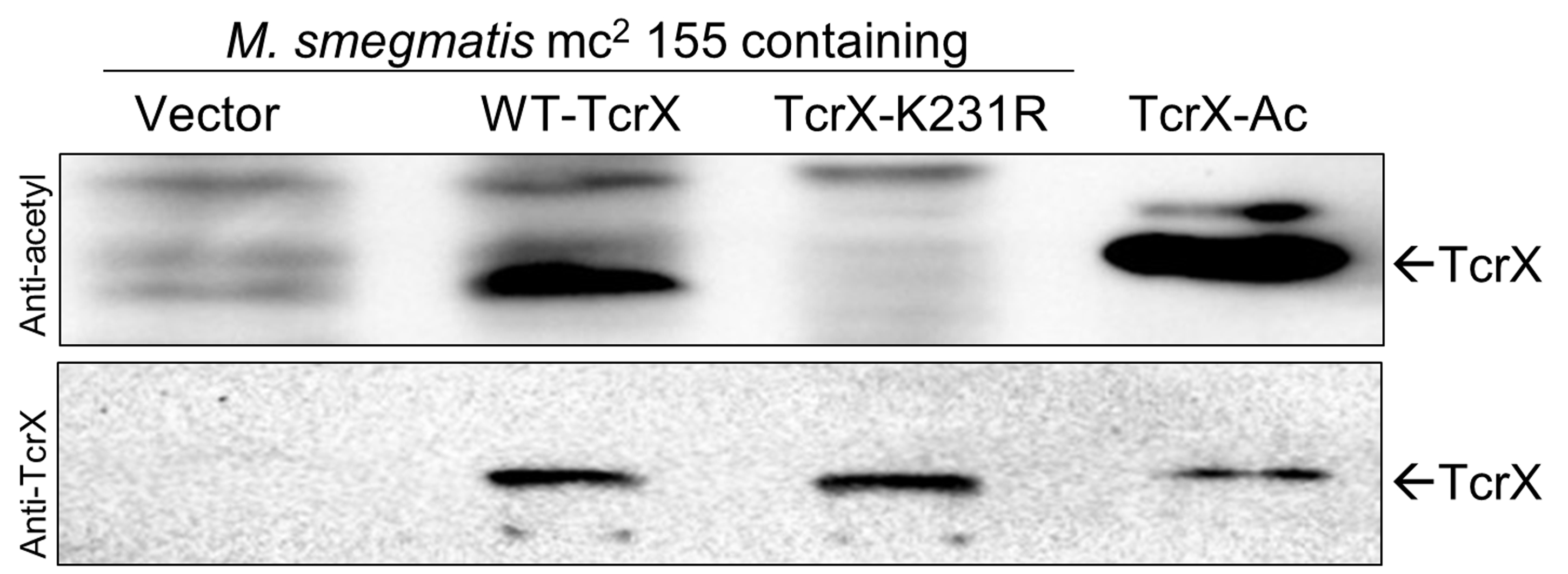
Figure 2. In vivo acetylation analysis. M. smegmatis mc2155 strains carrying either vector pMV261 alone or expressing wild type TcrX or its mutant protein were cultured in Middlebrook 7H9 medium (7H9). A total of 100 µg of total protein was analyzed by Western blotting using anti-acetyl lysine (top) and anti-TcrX (bottom) antibodies. Arrow represents the TcrX protein.
- Preparation for electrocompetent Mycobacterium smegmatis mc2155
Notes
- Care should be taken while preparing M. smegmatis cell lysates. The lysis step involving the bead-beating have to be optimized based on the homogenizer which might vary in intensity and beating frequency.
- Dilution of anti-lysine antibody should be optimized as it might vary between batch and batch.
Recipes
- ADC solution (Albumin Dextrose Catalase) (100 ml)
Dissolve albumin, dextrose and catalase in ~50 ml MilliQ water (~50 ml) and then make up the volume to 100 ml. Filter sterilize with 0.45-micron filters.Albumin 5 g Dextrose 2 g Catalase 3 mg - 10x acetylation buffer (1 ml)
Tris (pH ~7.4) 250 mM NaCl 1,000 mM - cAMP, 5 mM stock (1 ml)
Dissolve 1.84 mg of powder in ~0.5 ml MilliQ water till it completely dissolves and then make up the volume to 1 ml - Acetyl CoA, 5 mM stock (1 ml)
Dissolve 4.046 mg of Acetyl CoA powder in ~0.5 ml MilliQ water till it completely dissolves and then make up the volume to 1 ml - 7H9-Glycerol-Tween 80 medium (90 ml)
7H9 powder 0.47 g Glycerol (50%) 400 µl Tween 80 (20%) 250 µl - Dissolve 0.47 g of 7H9 powder in ~50 ml MilliQ water
- Add glycerol and Tween 80 as mentioned above and make up the total volume to 90 ml with MilliQ water
- Sterilize by autoclaving and store at RT
- Dissolve 0.47 g of 7H9 powder in ~50 ml MilliQ water
- Recovery Media (7H9-Glycerol-Tween 80-ADC) (20 ml)
7H9 17.87 ml ADC 2 ml Glycerol (50%) 80 µl Tween 80 (20%) 50 µl - Phosphate Buffer Saline (1,000 ml)
Sterilize by autoclaving and store at RTNaCl 8 g KCl 0.2 g Na2HPO4 1.44 g KH2PO4 0.24 g - Lysis Buffer (5 ml)
Tris-Cl (pH ~7.4) 20 mM NaCl 100 mM Glycerol 10% PMSF 1 mM Benzamidine Hydrochloride 1 mM - 5x Bradford Reagent (100 ml)
Make up the volume to 100 ml by adding MilliQ waterCoomassie Brilliant Blue G-250 50 mg Methanol (100%) 23.5 ml Phosphoric Acid (85%) 50 ml - Storage buffer (500 ml)
Tris-HCl, pH 8.0 50 mM Glycerol 50% NaCl 50 mM DTT 1 mM - SDS-PAGE Running Buffer (1,000 ml)
Make up the volume by adding MilliQ waterTris 3 g Glycine 14.4 g SDS 1 g
Note: SDS-PAGE running buffer will be of pH ~8.2-8.5. - Transfer Buffer (1 L)
Make up the volume to 1 L by adding MilliQ waterTris 3 g Glycine 14.4 g Methanol 200 ml
Note: Transfer buffer also will be of pH ~8.2-8.5. - Tris Buffer Saline Tween 20 (TBS-T) (1,000 ml)
Tris (pH ~7.5) 10 mM NaCl 153.8 mM Tween 20 0.1% - Stripping Buffer (20 ml)
Tris-Cl (pH 6.8) 62.5 mM SDS 2% β-mercaptoethanol 0.7% (140 μl/20 ml) - 5x Protein Sample buffer (10 ml)
Tris (pH-6.8) 0.25 M Glycerol 40% (4 ml/10 ml) SDS 10% Bromophenol blue 0.25% β-mercaptoethanol 5% (0.5 ml/10 ml)
Acknowledgments
This work was supported by Department of Biotechnology, India (Grant Nos. BT/PR17357/MED/29/1019/2016). The study was also supported in part by the DBT partnership program to Indian Institute of Science (DBT/BF/PRIns/2011-12) and Infosys Foundation; Equipment support by DST–Funds for Infrastructure in Science and Technology program (SR/FST/LSII-036/2016).
Competing interests
Authors declare no competing interests.
References
- Agrawal, R., Pandey, A., Rajankar, M. P., Dixit, N. M. and Saini, D. K. (2015). The two-component signalling networks of Mycobacterium tuberculosis display extensive cross-talk in vitro. Biochem J 469(1): 121-134.
- Bi, J., Gou, Z., Zhou, F., Chen, Y., Gan, J., Liu, J., Wang, H. and Zhang, X. (2018). Acetylation of lysine 182 inhibits the ability of Mycobacterium tuberculosis DosR to bind DNA and regulate gene expression during hypoxia. Emerg Microbes Infect 7(1): 108.
- Ferrer, M., Chernikova, T. N., Yakimov, M. M., Golyshin, P. N. and Timmis, K. N. (2003). Chaperonins govern growth of Escherichia coli at low temperatures. Nat Biotechnol 21(11): 1266-1267.
- He, F. (2011a). Bradford protein assay. Bio-protocol 1(6): e45.
- He, F. (2011b). Laemmli-SDS-PAGE. Bio-protocol 1(11): e80.
- Liu, F., Yang, M., Wang, X., Yang, S., Gu, J., Zhou, J., Zhang, X. E., Deng, J. and Ge, F. (2014). Acetylome analysis reveals diverse functions of lysine acetylation in Mycobacterium tuberculosis. Mol Cell Proteomics 13(12): 3352-3366.
- Nambi, S., Basu, N. and Visweswariah, S. S. (2010). cAMP-regulated protein lysine acetylases in mycobacteria. J Biol Chem 285(32): 24313-24323.
- Singh, K. K., Bhardwaj, N., Sankhe, G. D., Udaykumar, N., Singh, R., Malhotra, V. and Saini, D. K. (2019). Acetylation of response regulator proteins, TcrX and MtrA in M. tuberculosis tunes their phosphotransfer ability and modulates two-component signaling crosstalk. J Mol Biol 431(4): 777-793.
- Stoscheck, C. M. (1990). Quantitation of protein. Methods Enzymol 182: 50-68.
- Xie, L., Wang, X., Zeng, J., Zhou, M., Duan, X., Li, Q., Zhang, Z., Luo, H., Pang, L., Li, W., Liao, G., Yu, X., Li, Y., Huang, H. and Xie, J. (2015). Proteome-wide lysine acetylation profiling of the human pathogen Mycobacterium tuberculosis. Int J Biochem Cell Biol 59: 193-202.
- Yang, H., Sha, W., Liu, Z., Tang, T., Liu, H., Qin, L., Cui, Z., Chen, J., Liu, F., Zheng, R., Huang, X., Wang, J., Feng, Y. and Ge, B. (2018). Lysine acetylation of DosR regulates the hypoxia response of Mycobacterium tuberculosis. Emerg Microbes Infect 7(1): 34.
Article Information
Copyright
© 2019 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Singh, K. K., Singh, D. P., Singh, R. and Saini, D. (2019). In vitro and in vivo Assessment of Protein Acetylation Status in Mycobacteria. Bio-protocol 9(13): e3291. DOI: 10.21769/BioProtoc.3291.
Category
Microbiology > Microbial biochemistry > Protein > Modification
Molecular Biology > Protein > Activity
Biochemistry > Protein > Posttranslational modification
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link








