Advanced Search
Cultivation and Lugol staining of Chlamydia trachomatis
Last updated date: Apr 27, 2026 DOI: 10.21769/p2938 Views: 87 Forks: 0
Key features
Provides an efficient protocol for CT infection in McCoy B cells, followed by lugol stain.
Offers a low-cost option, in comparison to fluorophore-conjugated secondary antibodies.
Keywords
Chlamydia trachomatis; McCoy B cells, Cell culture; Lugol iodine staining
Abstract
Chlamydia trachomatis (CT) is an obligate intracellular bacterium that requires growth inside a mammalian host cell for propagation and survival. CT is the most common etiological agent of bacterial sexually transmitted infections (STIs) and can cause urethritis, cervicitis, pelvic inflammatory disease (PID), and infertility. Diagnosis is currently based on nucleic acid amplification tests (NAATs), with the use of cell culture reserved almost exclusively for research. In most laboratories, culture is no longer suitable as a diagnostic test because it has lower sensitivity than NAATs, involves a significantly long turnaround time, is technically complex, and lacks an internationally standardized and quality-assured methodology. However, Chlamydia culture capability should be maintained in some laboratories to monitor changes in antibiotic susceptibility and to support surveillance and research activities, as well as to enable research into other areas regarding the bacterium's molecular mechanisms. The current protocol provides a standardization for CT infection of eukaryotic cells and the use of Lugol's iodine staining.
Graphical overview
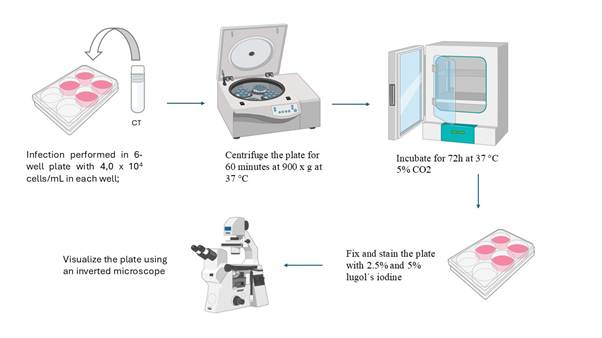
Background
Characterized by a biphasic life cycle, the bacterium differentiates into elementary body (EB) and reticulate body (RB). The host cell is infected when an EB adheres to the cytoplasmic membrane via multiple specific bindings (3, 4). The EB enters the cell through endocytosis, remaining within a parasitophorous vacuole - or inclusion. Inside this vacuole, CT differentiates from EB into RB and multiplies by fission 24 to 36 hours post-infection (3). During the multiplication process, the interior of the inclusion may contain RBs, EBs, various intermediate forms, and glycogen granules (3). When the RB-filled inclusion reaches a critical volume, coinciding with a depletion of nutrients and ATP, RBs convert back into EBs (4). EBs can be released from the host cell via two mechanisms: cell lysis or inclusion extrusion, a process resembling exocytosis (5).
To confirm CT infection in host cells in vitro, staining techniques are required to visualize the inclusion. The most widely used method involves fluorescein-conjugated monoclonal antibodies, causing the CT bodies within the inclusion to appear as bright green fluorescent cytoplasmic masses (6). While this staining method is more sensitive, it is also more expensive and labor-intensive. Another option is Lugol's iodine staining, in which the glycogen granules within the inclusion stain dark brown. Although this method has lower sensitivity compared to fluorescein-labeled monoclonal antibodies, it is easier to perform and it can be visualized with a conventional inverted microscope, without the need for fluorescence. Futhermore, it is also less expensive compared to fluorescein-labeled monoclonal antibodies (6).
The current protocol enables CT infection in McCoy B cells and standardizes Lugol's iodine staining technique.
Materials and reagents
Biological materials
Chlamydia trachomatis strain
McCoy B cells
Reagents
DMEM (Cultilab, catalog number: D1641)
Fetal bovine serum (FBS) (Cultilab, catalog number: F063)
Phosphate-Buffered Saline (PBS) (Laborclin, catalog number: LB 172373)
Methanol 100% (Synth, catalog number: A1085.01.BJ)
Ethanol 50% (Neon, catalog number: 03467)
Iodine crystals (I₂)
Potassium iodide (KI)
Solutions
Fixation solution (see recipes);
5% lugol solution (see recipes);
2,5% lugol solution (see recipes);
Recipes
Fixation solution
Add equal parts of 100% methanol and PBS (pH 7,0) (1:1) into a conical tube or another type of container;
Homogenize using a pipette.
5% lugol solution
Weigh 5 g of potassium iodide and 5 g of Iodine crystals in a beaker;
Add to 100 mL of 50% ethanol;
Filter slowly through standard grade cellulose paper;
Store in an amber container or wrapped in aluminum foil.
2,5% lugol solution
Weigh 2,5 g of potassium iodide and 2,5 g of Iodine crystals in a beaker;
Add to 100 mL of 50% ethanol;
Filter slowly through standard grade cellulose paper;
Store in an amber container or wrapped in aluminum foil.
Laboratory supplies
20 µL pipette tips (Olen, catalog number: K62-20-1)
50 µL pipette tips (Kasvi, catalog number: K66-100F-1)
200 µL pipette tips (Axygen, catalog number:TF-100-L-R-S-RCK)
1,5 mL Microcentrifuge tube (Eppendorf, catalog number: 0030120086)
5 mL serological pipettes (Kasvi, catalog number: K17-005)
25 mL serological pipettes (Kasvi, catalog number: K17-025)
15 mL conical tubes (Axygen, catalog number: AXYSCT15ML25S)
50 mL conical tubes (Axygen, catalog number: AXYSCT50ML25S)
Cell culture multi-well plates (6-well plate) (Kasvi, catalog number: K12-006)
Filter paper (BioCentrix, catalog number: FI193 )
Equipament
Forma™ Direct Heat CO2 Incubator (ThermoFisher, catalog number: 311)
Class II biological safety cabinet (Eletrospitalar, catalog number: EL0450/IIA2)
Laboratory centrifuge with rotors for plate (Eppendorf, catalog number: 5430)
Vortex mixer (Kasvi, catalog number: K45-2820)
Refrigerator (2 - 8 ºC)
Racks
Water bath, 37 oC (Solab, catalog number:SL-15/10)
XDS-1B inverted biological microscope (objectives: 4×, 10×, 20×) (Chongqing MIC Technology Co., Ltd)
Procedure
Plate Preparation
Clean the biosafety cabinet with 70% alcohol, turn on the UV light, and wait for 15 minutes;
Infection is performed in 6-well plate with 4,0 x 104 McCoy B cells/mL in each well;
The plate must contain a negative control, i.e., a well with cells that will not be infected;
Plate Infection
Homogenize the sample using a vortex or sonicator (stool or tissue samples);
Aspirate the culture medium from the plate wells and inoculate 200 μL; Add 50 μL of PBS 200 μL to the negative control;
Centrifuge the plate for 60 minutes at 900 x g at 37 °C;
Add 2 mL of culture medium supplemented with 10% FBS to each well and incubate the cells at 37°C (+/- 1 °C) for 72 h;
Fix and stain
After 72h of infection, aspirate the culture medium from the wells;
Gently wash the wells with 1 mL of PBS (pH 7,0);
Aspirate the PBS (pH 7,0);
Add 1 mL of fixation solution to the desired wells and leave for 10 minutes in the refrigerator;
Return to the biosafety cabinet and aspirate the solution from the wells;
Add 1 mL of 5% Lugol's iodine to the desired wells and leave for 10 minutes with the biosafety cabinet light turned off;
Aspirate the 5% Lugol's iodine from the wells;
Add 1 mL of 2.5% Lugol's iodine to the desired wells and leave for 10 minutes with the biosafety cabinet light turned off;
Aspirate the 2.5% Lugol's iodine from the wells, then wash the wells with distilled water and aspirate again;
Visualize the plate using an inverted microscope.
General notes and troubleshooting
During this protocol, wear protective clothing, including lab coat, gloves, safety glasses, and mask;
When preparing a solution, always pay attention to the mixing order: add the powders to the liquid;
If no inclusion formation is visible under the inverted microscope, blind passage sample two to three times before checking for inclusion development;
Check for Mycoplasma spp. contamination every couple of months;
In case of contamination with another bacteria or fungus, discard the plate into a biohazard container and prepare fresh reagents and properly sterilize by autoclaving or filtering.
Acknowledgments
The authors would like to thank the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES), Fundação de Amparo à Pesquisa e Extensão Universitária (FAPEU) and the Laboratório de Virologia Aplicada (LVA/UFSC).
Competing interests
The authors declare no conflict of interest.
References
Scidmore, M.A. (2005). Cultivation and Laboratory Maintenance of Chlamydia trachomatis. Curr Protoc Microbiol. Chapter 11:Unit 11A.1. doi: 10.1002/9780471729259.mc11a01s00
Unemo, M., Seth-Smith, H.M.G; Cutcliffe, L.T.; Skilton, R.J.; Barlow, D.; Goulding, D.; Persson, K.; Harris, S. R.; Kelly, A.; Bjartling, C.; Fredlund, H.; Olcén, P.; Thomson, N. R.; Clarke, I.N. (2010). The Swedish new variant of Chlamydia trachomatis: genome sequence, morphology, cell tropism and phenotypic characterization. Microbiology (Reading). 156(Pt 5):1394–1404.
Cevenini, R., Donati, M. and Sambri, V. (2002). Interactions between Chlamydia trachomatis and the host cell. Ann N Y Acad Sci. 975: 266–271. https://doi.org/10.1111/j.1749-6632.2002.tb05963.x
Witkin, S.S., Minis, E., Athanasiou, A., Leong, J. and Linhares, I.M. (2017). Chlamydia trachomatis: the persistent pathogen. Clin Vaccine Immunol. 24(10): e00203-17. https://doi.org/10.1128/CVI.00203-17
Elwell, C., Mirrashidi, K. and Engel, J. (2016). Chlamydia cell biology and pathogenesis. Nat Rev Microbiol. 14(6): 385–400. https://doi.org/10.1038/nrmicro.2016.30
Brasil. Ministério da Saúde. (2015). Diagnóstico laboratorial das doenças sexualmente transmissíveis. Brasília: Ministério da Saúde.
- Wachter, J K, Venturi, C M, Rocha, A V, Barazzetti, F H, Martinez, R E and Bazzo, M(2026). Cultivation and Lugol staining of Chlamydia trachomatis. Bio-protocol Preprint. 10.21769/p2938.
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link
