- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
H2O2 Kill Assays of Biofilm Bacteria
Published: Vol 3, Iss 21, Nov 5, 2013 DOI: 10.21769/BioProtoc.952 Views: 11701
Reviewed by: Anonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
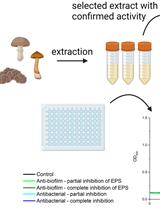
SIMBA Method—Simultaneous Detection of Antimicrobial and Anti-biofilm Activity of New Compounds Using Salmonella Infantis
Meta Sterniša [...] Anja Klančnik
Aug 5, 2023 2060 Views
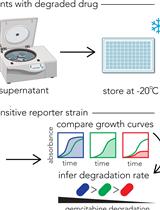
Functional Assay for Measuring Bacterial Degradation of Gemcitabine Chemotherapy
Serkan Sayin and Amir Mitchell
Sep 5, 2023 1766 Views
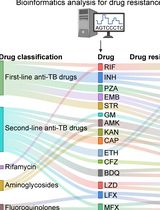
Identification of Mycobacterium tuberculosis and its Drug Resistance by Targeted Nanopore Sequencing Technology
Chen Tang [...] Guangxin Xiang
Feb 5, 2025 2046 Views
Abstract
Ubiquitous in nature and often surface associated, biofilms cause numerous chronic human infections. Biofilms are structured multicellular bacterial communities where cells are entrapped in a polymer matrix. Bacteria growing as biofilms are characterized by marked tolerance to many biocides, including oxidants such as hydrogen peroxide. Hydrogen peroxide is both produced by host phagocytic cells, and used as an antimicrobial compound. Understanding biofilm tolerance to hydrogen peroxide is therefore relevant to the persistence of Pseudomonas aeruginosa in human infections (such as chronic Pseudomonas aeruginosa infections in cystic fibrosis airways) as well as in environmental settings (such as water pipes).
This protocol was developed to determine the tolerance of Pseudomonas aeruginosa biofilms to hydrogen peroxide (H2O2) killing. The bacteria are grown as colony biofilms on polycarbonate membranes, as previously described in Walters et al. 2003. The protocol may be adapted for other bacterial, with appropriate changes in H2O2 concentrations, since different bacterial species may be more or less susceptible to H2O2 than Pseudomonas aeruginosa.
Materials and Reagents
- Phosphate buffered saline (PBS) solution (Sigma-Aldrich, catalog number: P4417-100TAB )
- 30% w/w Hydrogen peroxide solution (undiluted, as sold commercially) (RICCA Chemical, catalog number: 3821.7-32 )
- Sodium thiosulfate solution (dissolved in ddH2O) (Sigma-Aldrich, catalog number: S8503 )
- 0.2 μM Polycarbonate 25 mm membranes (General Electric Company, catalog number: K02BP02500 )
- P.aeruginosa strains in freezer stock
- 25% Lennox broth (LB) medium (Becton Dickinson and Company, DifcoTM, catalog number: 240230 ) (see Recipes)
- 25% LB agar plates (see Recipes)
Equipment
- 6-well and 96-well plates
- Standard petri plates
- Spectrophotometer (cuvette) (Thermo Fisher Scientific, model: GENESYS 10S UV-Vis )
- Spectrophotometer (96-well plate) (Bio-Rad, model: 680 )
- Cuvettes for OD600 reading
- Shaking incubator at 37 °C, 250 rpm
- Static incubator at 37 °C
- Sterile glassware: 150 ml Erlenmeyer flasks, capped or foiled
- 1.5 ml and 2 ml microcentrifuge tubes
- Source of UV irradiation
- Sterile wire-loops (sterilized with 70% ethanol and flame)
- Stainless steel forceps (sterilized with 70% ethanol and flame)
Procedure
- Day 0. Streak P.aeruginosa cells from the freezer stock onto a LB agar place and incubate statically overnight at 37 °C.
- Day 1. Pick 4-5 single colonies from the P.aeruginosa agar plate with a sterile wired-loop and inoculate 15 ml liquid LB medium in a 150 ml Erlenmeyer flask. Grow liquid bacterial cultures overnight for 16-18 hours at 37 °C, with shaking at 250 rpm.
- Day 2. Gently place polycarbonate membranes on agar surface of fresh sterile 25% LB agar plates and sterilize the membranes by placing them under UV irradiation for 1 hour. Handle membranes carefully with sterile forceps and use the membranes immediately after sterilization. Use eyes and skin UV protective equipment. Use at least 3 membranes per strain per condition for adequate biological replicates, and up to 6 membranes may be placed on each agar plate.
- Measure the OD600 of the overnight bacterial culture and dilute the bacterial suspension in LB medium to a starting concentration of 108 cells/ml. Depending on the bacterial strain used, the OD600 to CFU ratio will differ and needs to be determined for each strain: for example, for the PAO1 wild type strain, 108 cells/ml = ~OD600 0.1.
- Spot 5 μl (5 x 105 cells) onto the sterile membranes and allow the liquid to be absorbed (10-20 minutes).
- Incubate the colony biofilm on agar plates for 24 h at 37 °C.
- Day 3. Using sterile forceps, gently lift the membranes off the agar surface and transfer them (cells side down) into 6-well plates filled with 2 ml of 25% LB liquid medium in each well. Make sure the membranes are spread flat (i.e. not rolled up) and biofilm cells are remain on the membrane.
- For the H2O2 treated biofilms, add 30 μl H2O2 (150 mM) to each well in pulses every 10 minutes for 30 minutes (for a final concentration of 450 mM H2O2 per challenge). The pulsing is done to mimic a continuous exposure of cells to H2O2. In between H2O2 pulses, incubate cells at 37 °C without shaking. Include untreated controls that are challenged with PBS. Each condition should be done at least in triplicates.
- After H2O2 or PBS challenge, add 0.2% sodium thiosulfate to all samples to neutralize any remaining H2O2. Add even when samples are only challenges with PBS as a control.
- To determine the viable cell count in H2O2 or PBS treated biofilms, collect biofilm cells by transferring the membranes and 2 ml of liquid from each well into 2 ml microcentrifuge tubes. Membranes are moved by gently lifting and rolling them using sterile forceps, with biofilm cells facing inward. The entire membrane should be submerged in liquid. Ensure to sterilize forceps between membrane transfers. Vortex biofilms in microcentrifuge tubes at maximal speed for at least 1 minute to detach and resuspend cells. Additional pipetting up and down and vortexing may also necessary to make sure there are no cell clumps visible in the bacterial suspension.
- Aliquot 100 μl of bacterial suspension into 96-well plate, serially dilute cells 1:10, then plate 100 μl of each dilution on LB agar plates for CFU count. Incubate CFU count plates at 37 °C overnight.
- Day 4. Count CFU on LB agar plates and calculate the viable CFU per biofilm based on the dilution factors applied.
- Determine hydrogen peroxide killing by comparing the viable CFU count in the PBS treated and the H2O2 treated conditions.
Recipes
- 25% LB medium
5 g LB powder medium per L
Dissolved in ddH2O and autoclaved
- 25% LB agar plates
25% LB medium with 1.5% agar
Dissolved in ddH2O and autoclave
Acknowledgments
We would like to acknowledge CIHR (MOP-102727 to DN) and the Burroughs Wellcome Fund (1006827.01 to DN) for funding. This protocol was adapted from the previously published paper Khakimova et al. (2013).
References
- Khakimova, M., Ahlgren, H. G., Harrison, J. J., English, A. M. and Nguyen, D. (2013). The stringent response controls catalases in Pseudomonas aeruginosa and is required for hydrogen peroxide and antibiotic tolerance. J Bacteriol 195(9): 2011-2020.
- Walters, M. C., 3rd, Roe, F., Bugnicourt, A., Franklin, M. J. and Stewart, P. S. (2003). Contributions of antibiotic penetration, oxygen limitation, and low metabolic activity to tolerance of Pseudomonas aeruginosa biofilms to ciprofloxacin and tobramycin. Antimicrob Agents Chemother 47(1): 317-323.
Article Information
Copyright
© 2013 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Khakimova, M. and Nguyen, D. (2013). H2O2 Kill Assays of Biofilm Bacteria. Bio-protocol 3(21): e952. DOI: 10.21769/BioProtoc.952.
Category
Microbiology > Microbial biofilm > Killing assay
Biochemistry > Other compound > Reactive oxygen species
Microbiology > Antimicrobial assay > Antibacterial assay
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










