- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
IEF-2DE Analysis and Protein Identification
Published: Vol 3, Iss 19, Oct 5, 2013 DOI: 10.21769/BioProtoc.924 Views: 16297
Reviewed by: Tie Liu

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
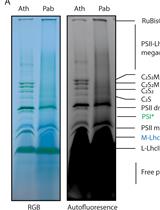
Solubilization Method for Isolation of Photosynthetic Mega- and Super-complexes from Conifer Thylakoids
Pushan Bag [...] Domenica Farci
Sep 5, 2021 4140 Views
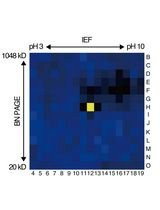

An Activity-Based Proteomics with Two-Dimensional Polyacrylamide Gel Electrophoresis (2D-PAGE) for Identifying Target Proteases in Arabidopsis Apoplastic Fluid
Sayaka Matsui and Yoshikatsu Matsubayashi
Mar 5, 2025 2035 Views
Advancing 2-DE Techniques: High-Efficiency Protein Extraction From Lupine Roots
Sebastian Burchardt [...] Emilia Wilmowicz
Oct 5, 2025 1801 Views
Abstract
Isoelectric focusing followed by SDS-PAGE (IEF-2DE) separates proteins in a two-dimensional matrix of protein pI (Protein Isoelectric Point) and molecular weight (MW). The technique is particularly useful to distinguish protein isoforms (Radwan et al., 2012) and proteins that contain post-translational modifications such as phosphorylation (Oh et al., 2012) and lysine acetylation (Wu et al., 2011). Proteins that are separated by IEF-2DE can be identified by immunoblot analysis using sequence-specific antibodies or by mass spectrometry.
Materials and Reagents
- Protein extracts
- Phenol (TE-saturated pH 7.9) (Sigma-Aldrich, catalog number: P4557 )
- 0.1 M ammonium acetate (dissolved in methanol)
- Tris-base (General Electric Company, catalog number: 17-1321-01 )
- Glycine
- Urea (Life Technologies, Invitrogen™, catalog number: ZU10001 )
- Thiourea (Life Technologies, Invitrogen™, catalog number: ZT10002 )
- CHAPS (Life Technologies, Invitrogen™, catalog number: ZC10003 )
- DTT (General Electric Company, catalog number: 17-1318-01 )
- Bromophenol blue (General Electric Company, catalog number: 17-1329-01 )
- Glycerol (General Electric Company, catalog number: 17-1325-01 )
- SDS (General Electric Company, catalog number: 17-1313-01 )
- Iodoacetamide (General Electric Company)
- 100% Ethanol (HPLC grade)
- Water (HPLC grade)
- IPG buffer (General Electric Company, catalog number: 17-6000-87 )
- IPG strips (General Electric Company, catalog number: 17-6000-14 )
- Immobiline DryStrip Cover Fluid (General Electric Company, catalog number: 17-1335-01 )
- Bradford assay dye reagent (Bio-Rad Laboratories, catalog number: 500-0006 )
- SimplyBlueTM SafeStain protein stain (Life Technologies, Invitrogen™, catalog number: LC-6065 )
- Precision PlusTM All Blue Protein Standards (Bio-Rad Laboratories, catalog number: 161-0373 )
- IEF sample buffer (see Recipes)
- SDS equilibration buffer (see Recipes)
- 10x Tank buffer (see Recipes)
- Agarose overlay solution (General Electric Company, catalog number: 17-0554-01 ) (see Recipes)
Equipment
- 1.5 ml microfuge tubes
- 15 ml tubes
- EttanTM IPGphoreTM Isoelectric Focusing System (General Electric Company, Model: 80-6414-02 )
- Large Format 1-D Electrophoresis Systems (Bio-Rad Laboratories)
- Semi-dry transfer system (Bio-Rad Laboratories)
- Mass spectrometers (such as MALDI-TOF or tandem mass spectrometer)
Procedure
- Sample preparation
To maximize the separation of proteins by IEF, the protein extracts should contain minimal contaminants of salts and detergents. Here is the method we used to prepare protein extracts (protein extraction buffer: 100 mM Tris-HCl, pH 8.0) for IEF analysis.
- Mix crude protein extract and ice-cold phenol (TE-saturated, pH 7.9) in a 1:1 (v:v) ratio. Vortex for 1 min.
- Centrifuge the mixture at 24,000 x g for 15 min at 4 °C.
- Carefully remove upper aqueous phase and discard (do not disturb interphase).
- Add equal volume of ice-cold 50 mM Tris-HCl (pH 8.0) to the protein mixture. Vortex for 1 min.
- Centrifuge at 24,000 x g for 15 min at 4 °C.
- Remove upper aqueous phase and discard (do not disturb interphase).
- Repeat steps 4 to 6 once more.
- Transfer the lower phenol phase (including interphase) to a new ice-cold 15 ml tube and add 5 volumes of ice-cold 0.1 M ammonium acetate (in methanol). Vortex for 1 min.
- Precipitate proteins overnight at -20 °C.
- Gently and briefly vortex to resuspend the protein precipitate. Split the samples between 1.5 ml microfuge tubes as necessary.
- Centrifuge at 24,000 x g for 20 min at 4 °C and carefully remove the supernatant.
- Add 1 ml ice-cold 0.1 M ammonium acetate (in methanol) to each microfuge tube.
- Let pellet “soak” in the buffer for 10 min at -20 °C.
- Centrifuge at 24,000 x g for 20 min at 4 °C.
- Pipette out the supernatant and take care to not disturb the pellet.
- Repeat steps 12 to 15 twice.
- Wash each pellet further with 1 ml ice-cold 100% ethanol.
- Centrifuge at 24,000 x g for 20 min at 4 °C.
- Pipette out the supernatant (as much as possible).
- Resuspend each protein pellet with the volume of about 50-100 μl IEF sample buffer. The total resuspension volume per sample is about 500 μl.
- Vortex the pellet and let it dissolve in IEF sample buffer at room temperature for 1 h. Do not heat the pellet in IEF sample buffer.
- Centrifuge at 24,000 x g for 20 min at room temperature. The resulting supernatant is suitable for IEF analysis.
- Quantify the protein concentration with the Bradford assay. Store protein extract at -80 °C until use.
- Mix crude protein extract and ice-cold phenol (TE-saturated, pH 7.9) in a 1:1 (v:v) ratio. Vortex for 1 min.
- IEF Protein separation
- Thaw the protein extract at room temperature.
- Centrifuge the protein extract at 24,000 x g for 20 min at room temperature to remove any precipitate.
- Take an aliquot containing 200 μg of total protein and adjust the volume to 250 μl with IEF sample buffer.
Note: A good range of sample loading for a 13 cm IPG strip is about 100 – 250 μg total proteins. Generally, we used 13 cm IPG strip (pH 3-10), but we can choose the strip depend on proteins analyzing.
- Load 250 μl of each sample uniformly into the ceramic strip holder (13 cm size).
- Take out a 13 cm IPG strip and remove the thin plastic seal from the gel side of the strip.
Notes:
- The pH range of the strip should match the pH range of the IPG buffer in the IEF sample buffer.
- Use a tweezer to handle IPG strip. Avoid touching the gel area of the strip.
- The pH range of the strip should match the pH range of the IPG buffer in the IEF sample buffer.
- Carefully place the IPG strip onto the surface of the IEF sample in the ceramic strip holder, with gel-side down and the acidic end of the strip (marked with a “+”) towards the pointed (+) end of strip holder (Figure 1A).
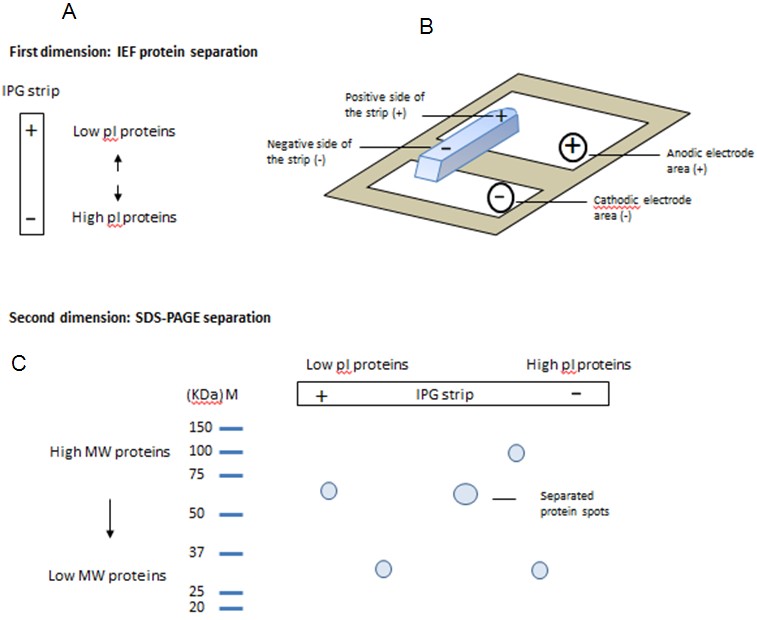
Figure 1. Illustration of IEF-2DE protein separation by protein pI and MW. A. In the first dimension, proteins are separated by protein pI in IEF strip. B. IEF setup to show the positive side the IEF gel should match the arrow direction of ceramic strip holder and the IPGphor positive electrode plate. C. In the second dimension, proteins are further separated by protein MW in SDS-PAGE.
- Carefully raise and lower the strip to assure the gel side of IPG strip is in good contact with the IEF sample solution.
- Overlay strip with about 1.5 ml Immobiline DryStrip Cover Fluid.
- Cover the strip holder with the plastic lid.
- Place the strip unit onto the IPGphor Manifold with pointed (+) end on the large “+” electrode plate.
- Begin IEF protocol #1 (Voltage-assisted rehydration sample loading) 20 °C, 50 μA per strip, 0 h rehydration (under non-voltage).
Note: Ensure that the correct number of IPG strips to be run is entered.
Step 1 = 30 V for 360 voltage-hour (Vh)
Step 2 = 500 V for 500 Vh
Step 3 = 1,000 V for 1,000 Vh
Step 4 = 8,000 V for 32,000 Vh
Step 5, 10 h hold at 8,000 V
Total running time = about 20 h (This range of voltage-hour recommended for all pH ranges of IPG strip)
- The IEF separation is completed. A good range to stop is at the 32,000 Vh to 42,000 Vh.
- Proceed directly to IPG strip equilibration (described below) or store the strip in a closed container before processing further at -80 °C.
- Thaw the protein extract at room temperature.
- SDS-PAGE Protein separation
- Pour a 1 mm x 20 cm sized 10% acrylamide gel (pH 8.8 or 9.2) using a flat comb with only a single protein marker well.
- Allow to polymerize for 45 to 60 min at room temperature.
- Rinse upper surface of running gel with de-ionized water.
- Prepare 2 L of 1x solution of Tank Buffer.
- Prepare fresh SDS equilibration buffer (± DTT or Iodoacetamide). 20 ml of buffer is needed for each IPG strip.
- Directly after IEF, remove the IPG strip and place it into equilibration tube with plastic backing against the tube wall.
Note: Handle each strip individually. Don’t mix up the strips.
- Add 10 ml of SDS equilibration buffer (+ DTT) to each tube and gently shake on the rocker platform for 10 min at room temperature.
- Decant liquid from tube and add 10 ml SDS equilibration buffer (+ DTT) and gently shake on the rocker platform for an additional 15 min at room temperature.
- Decant liquid from tube and replace with 10 ml SDS equilibration buffer (+ Iodoacetamide) and gently shake on the rocker platform for 10 min at room temperature.
- Decant liquid from tube and add 10 ml SDS equilibration buffer (+ Iodoacetamide) and gently shake on the rocker platform for additional 15 min at room temperature.
- Use a tweezer to place IPG strip in-between the glass plates of the gel with gel-side up and acidic end strip (+) nearest the reference well.
- Use a 0.75 mm spacer to gently push the IPG strip onto the top surface of the second dimension gel. Avoid bubbles between strip and gel, and ensure good contact.
- Seal strip in place with about 500 μl of “cooled” Agarose overlay solution. Let solidify for about 3 min.
- Assemble gels into Bio-Rad Protean II.
- Fill upper buffer tank with 1x Tank buffer until full. Place remainder in bottom buffer tank with stir bar.
- Load 20 μl Bio-Rad protein standards into the reference well.
- Electrophorese at 10 mA per gel (constant current) (~ 40-125 V) for about 15 h, or at 20 mA per gel (constant current) (~ 80-250 V) for about 7.5 h. (This is the approximate time required for bromophenol blue dye to migrate within 1 cm of the end of a 10% gel.)
- Pour a 1 mm x 20 cm sized 10% acrylamide gel (pH 8.8 or 9.2) using a flat comb with only a single protein marker well.
- Protein identification from IEF-2DE
- Following IEF-2DE protein separation, proteins can be transferred onto PVDF membrane for immunoblot analysis. We used the Bio-Rad semi-dry transfer system and found efficient transfer.
- Alternatively, the 2-DE gel can be stained with Coomassie blue to visualize protein spots. Protein spots of interests can be excised from the gel for mass spectrometry analysis following removal of the dye. (Figure 2)
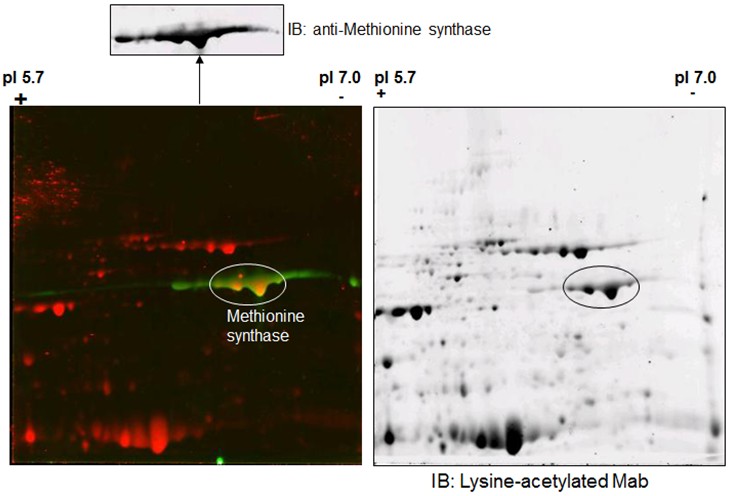
Figure 2. Total soluble proteins (250 μg) of Arabidopsis thaliana are separated by 2-D gel electrophoresis and identified specific proteins with methionine synthase and lysine-acetylated Abs
- Following IEF-2DE protein separation, proteins can be transferred onto PVDF membrane for immunoblot analysis. We used the Bio-Rad semi-dry transfer system and found efficient transfer.
Recipes
- IEF sample loading buffer Final concentration
Add 0.42 g Urea 7 M
0.15 g Thiourea 2 M
400 μl of 10% CHAPS 4%
20 μl of IPG buffer (appropriate pH range for IPG strip) 2%
0.01 g DTT 1%
20 μl of 0.1% bromophenol blue 0.002%
Check that final volume is 1 ml
- SDS equilibration buffer (± DTT or Iodoacetamide), make freshly Final concentration
Add 1.333 ml of 1.5 M Tris Base (pH 8.8) into a 50 ml tube 50 mM
12 ml of 100% glycerol 30%
8 ml of 10% SDS 2%
800 μl of 0.1% bromophenol blue 0.002%
14.41 g Urea 6 M
Add water to 40 ml total volume
After the solution is completely dissolved. Divide the solution in half
Add 0.2 g DTT to 20 ml SDS equilibration buffer(and no iodoacetamide) 2%
Add 0.5 g iodoacetamide to 20 ml SDS equilibration buffer (and no DTT) 5%
- 10x Tank buffer Final concentration
Dissolve 60.4 g Tris Base 25 mM
288 g glycine 192 mM
20 g SDS 0.1%
In 2 L of water
Do not adjust pH, 10x pH ~ 8.6, 1x pH ~ 8.1
- Agarose overlay solution Final Concentration
980 μl of 1x Tank buffer 1x
20 μl of 0.1% bromophenol blue 0.002%
0.005 g Agarose powder 0.5%
Total volume is 1 ml ( ~ 250 μl needed per strip)
Heat at 95 °C for ~10 min to melt
Let cool to touch (~70 °C) before use
Acknowledgments
None.
References
- Oh, M. H., Clouse, S. D. and Huber, S. C. (2012). Tyrosine phosphorylation of the BRI1 receptor kinase occurs via a post-translational modification and is activated by the juxtamembrane domain. Front Plant Sci 3: 175
- Radwan, O., Wu, X., Govindarajulu, M., Libault, M., Neece, D. J., Oh, M. H., Berg, R. H., Stacey, G., Taylor, C. G., Huber, S. C. and Clough, S. J. (2012). 14-3-3 proteins SGF14c and SGF14l play critical roles during soybean nodulation. Plant Physiol 160(4): 2125-2136.
- Wu, X., Oh, M. H., Schwarz, E. M., Larue, C. T., Sivaguru, M., Imai, B. S., Yau, P. M., Ort, D. R. and Huber, S. C. (2011). Lysine acetylation is a widespread protein modification for diverse proteins in Arabidopsis. Plant Physiol 155(4): 1769-1778.
Article Information
Copyright
© 2013 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Readers should cite both the Bio-protocol article and the original research article where this protocol was used:
- Wu, X., Clough, S. J., Huber, S. C. and Oh, M. (2013). IEF-2DE Analysis and Protein Identification. Bio-protocol 3(19): e924. DOI: 10.21769/BioProtoc.924.
- Radwan, O., Wu, X., Govindarajulu, M., Libault, M., Neece, D. J., Oh, M. H., Berg, R. H., Stacey, G., Taylor, C. G., Huber, S. C. and Clough, S. J. (2012). 14-3-3 proteins SGF14c and SGF14l play critical roles during soybean nodulation. Plant Physiol 160(4): 2125-2136.
Category
Biochemistry > Protein > Electrophoresis
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link











