- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Structural Based Strategy for Predicting Transcription Factor Binding Sites
Published: Vol 3, Iss 12, Jun 20, 2013 DOI: 10.21769/BioProtoc.794 Views: 9157
Reviewed by: Fanglian HeLin FangAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
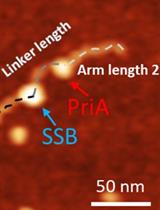
Characterize the Interaction of the DNA Helicase PriA with the Stalled DNA Replication Fork Using Atomic Force Microscopy
Yaqing Wang [...] Yuri L. Lyubchenko
Mar 5, 2021 4890 Views
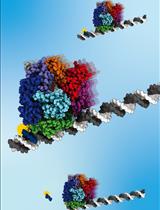
A Gel-Based Assay for Probing Protein Translocation on dsDNA
Christiane Brugger and Alexandra M. Deaconescu
Jul 20, 2021 4377 Views
Abstract
Scanning through genomes for potential transcription factor binding sites (TFBSs) is becoming increasingly important in this post-genomic era. The position weight matrix (PWM) is the standard representation of TFBSs utilized when scanning through sequences for potential binding sites. Many transcription factor (TF) motifs are short and highly degenerate, and methods utilizing PWMs to scan for sites are plagued by false positives. Furthermore, many important TFs do not have well-characterized PWMs, making identification of potential binding sites even more difficult. One approach to the identification of sites for these TFs has been to use the 3D structure of the TF to predict the DNA structure around the TF and then to generate a PWM from the predicted 3D complex structure. However, this approach is dependent on the similarity of the predicted structure to the native structure. We introduce here a novel approach to identify TFBSs utilizing structure information that can be applied to TFs without characterized PWMs, as long as a 3D complex structure (TF/DNA) exists. Our approach utilizes an energy function that is uniquely trained on each structure thus leads to increased prediction accuracy and robustness compared with those using a more general energy function. The software is freely available upon request. Please see reference supplementary material for details.
Keywords: ChIP-SeqData and Software
- TF/DNA structure
tFIRE takes standard PDB format for TF/DNA structures.- A good place to look for such data is PDB (Rose et al., 2011) (http://www.pdb.org).
- If the TF/DNA complex structure does not exist but the TF structure exists, you can generate the TF/DNA structure by docking DNA to the TF structure. Software that can be utilized for this includes HADDOCK (De Vries et al., 2007) (http://www.nmr.chem.uu.nl/haddock), FTDOCK (Jackson R.M. et al., 1998) (http://www.sbg.bio.ic.ac.uk/docking/ftdock.html), YASARA DOCK (http://www.yasara.org/dnadock.htm), ParaDock (Banitt and Wolfson, 2011) (http://www.paradocks.org). Our method indicates that such docking will not affect a result significantly but we have not tested any of these docking predictions ourselves for validation. Hence we urge their use with caution, and revalidate the results once the structures are available.
- If the TF structure does not exist in 3D structure databases, you can predict TF structure using homology modeling like SWISS-MODEL (Guex and Peitsch, 1997) (http://swissmodel.expasy.org), Rosetta (Bradley et al., 2003) (https://www.rosettacommons.org), Sybyl (Visegrády et al., 2001) (http://www.tripos.com/index.php?family=modules,SimplePage,,,&page=SYBYL-X). Please use with caution that each prediction step would reduce the accuracy.
- Predict TFBSs
tFIRE predicted motif(PWM) can be used to predict TFBSs.- Motif scanning programs can be used to scan the whole genome for motif matches. Such methods included MAST (Bailey et al., 2006) (http://meme.nbcr.net/meme/cgi-bin/mast.cgi) from MEME suite and STORM (Schones et al., 2007) (http://rulai.cshl.edu/storm) from Cold Spring Harbor Laboratory.
- TFBSs vary for different cells. Recently, a newly developed method, CENTIPEDE (Pique-Regi et al., 2010) (http://centipede.uchicago.edu) shows that with the result from a single DNase-seq experiment, one can accurately predict TFBSs for all TFs. Therefore, downloading DNase-seq data from ENCODE project can be very helpful.
- Recently developed FAIRE-seq technology allow similar predictions for detection of chromatin accessibility regions (Song et al., 2011). Such data can be substituted for DNase-seq, but needs be tested and validated before use.
- Epigenetic information can also be employed instead of DNase-seq (Cuellar-Partida et al., 2012). We propose to update our data and methods available for such predictions on a regular basis.
- Software
- C++ software environment, better with Linux system
- tFIRE, Feel Free to ask the author for a linux version
- WebLogo (Crooks et al., 2004) can be used for visualization the PWM we predicted (http://weblogo.berkeley.edu)
Procedure
- If you are confident of your TF/DNA complex 3D structure, then you can use tFIRE default function pre-trained by all available TF/DNA structures in the PDB database. You can also construct your own energy function with tFIRE by several non-homology structures. You can use the PISCES server (Wang and Dunbrack, 2003) (http://dunbrack.fccc.edu/PISCES.php) that this server will give you a subset of your input structure list (PDB id) that each protein in the subset has little homology to another.
- If you are not confident of your TF/DNA structure, you can train tFIRE with a single structure and subsequently predict PWMs using tFIRE.
Acknowledgments
This protocol have been adapted from: Xu et al. (2013). We thank the funding supported by the National Sciences Foundation of China (no. 31070641) and National 973 Program of China (no. 2012CB721000) and start-up funding from SKLMRD and DICP, CAS (Chinese Academy of Sciences). The funders offered most of the costs of study design, data collection and analysis, decision to publish, or preparation of the manuscript. We would like to thank Dr. Yan Cui, Dr. Yaoqi Zhou, Dr. Yuedong Yang, Dr. Chi Zhang, Dr. Song Liu, Dr. Jason Donald, Dr. Eugene Shakhnovich, Dr. Timothy Robertson, Dr. Gabriele Varani, Dr. Marc Jung, Dr. Amy Leung and Dr. Rongze Lu, Juan Du for their databases, programs and helpful discussions.
References
- Bailey, T. L., Williams, N., Misleh, C. and Li, W. W. (2006). MEME: discovering and analyzing DNA and protein sequence motifs. Nucleic Acids Res 34(Web Server issue): W369-373.
- Banitt, I. and Wolfson, H. J. (2011). ParaDock: a flexible non-specific DNA--rigid protein docking algorithm. Nucleic Acids Res 39(20): e135.
- Bradley, P., Chivian, D., Meiler, J., Misura, K. M., Rohl, C. A., Schief, W. R., Wedemeyer, W. J., Schueler-Furman, O., Murphy, P., Schonbrun, J., Strauss, C. E. and Baker, D. (2003). Rosetta predictions in CASP5: successes, failures, and prospects for complete automation. Proteins 53 Suppl 6: 457-468.
- Crooks, G. E., Hon, G., Chandonia, J. M. and Brenner, S. E. (2004). WebLogo: a sequence logo generator. Genome Res 14(6): 1188-1190.
- Cuellar-Partida, G., Buske, F. A., McLeay, R. C., Whitington, T., Noble, W. S. and Bailey, T. L. (2012). Epigenetic priors for identifying active transcription factor binding sites. Bioinformatics 28(1): 56-62.
- de Vries, S. J., van Dijk, A. D., Krzeminski, M., van Dijk, M., Thureau, A., Hsu, V., Wassenaar, T. and Bonvin, A. M. (2007). HADDOCK versus HADDOCK: new features and performance of HADDOCK2.0 on the CAPRI targets. Proteins 69(4): 726-733.
- Guex, N. and Peitsch, M. C. (1997). SWISS-MODEL and the Swiss-PdbViewer: an environment for comparative protein modeling. Electrophoresis 18(15): 2714-2723.
- Jackson, R. M., Gabb, H. A. and Sternberg, M. J. (1998). Rapid refinement of protein interfaces incorporating solvation: application to the docking problem. J Mol Biol 276(1): 265-285.
- Pique-Regi, R., Degner, J. F., Pai, A. A., Gaffney, D. J., Gilad, Y. and Pritchard, J. K. (2011). Accurate inference of transcription factor binding from DNA sequence and chromatin accessibility data. Genome Res 21(3): 447-455.
- Rose, P. W., Beran, B., Bi, C., Bluhm, W. F., Dimitropoulos, D., Goodsell, D. S., Prlic, A., Quesada, M., Quinn, G. B., Westbrook, J. D., Young, J., Yukich, B., Zardecki, C., Berman, H. M. and Bourne, P. E. (2011). The RCSB Protein Data Bank: redesigned web site and web services. Nucleic Acids Res 39(Database issue): D392-401.
- Schones, D. E., Smith, A. D. and Zhang, M. Q. (2007). Statistical significance of cis-regulatory modules. BMC Bioinformatics 8: 19.
- Song, L., Zhang, Z., Grasfeder, L. L., Boyle, A. P., Giresi, P. G., Lee, B. K., Sheffield, N. C., Graf, S., Huss, M., Keefe, D., Liu, Z., London, D., McDaniell, R. M., Shibata, Y., Showers, K. A., Simon, J. M., Vales, T., Wang, T., Winter, D., Clarke, N. D., Birney, E., Iyer, V. R., Crawford, G. E., Lieb, J. D. and Furey, T. S. (2011). Open chromatin defined by DNaseI and FAIRE identifies regulatory elements that shape cell-type identity. Genome Res 21(10): 1757-1767.
- Visegrady, B., Than, N. G., Kilar, F., Sumegi, B., Than, G. N. and Bohn, H. (2001). Homology modelling and molecular dynamics studies of human placental tissue protein 13 (galectin-13). Protein Eng 14(11): 875-880.
- Wang, G. and Dunbrack, R. L., Jr. (2003). PISCES: a protein sequence culling server. Bioinformatics 19(12): 1589-1591.
- Xu, B., Yang, Y., Liang, H. and Zhou, Y. (2009). An all-atom knowledge-based energy function for protein-DNA threading, docking decoy discrimination, and prediction of transcription-factor binding profiles. Proteins 76(3): 718-730.
- Xu, B., Schones, D. E., Wang, Y., Liang, H. and Li, G. (2013). A structural-based strategy for recognition of transcription factor binding sites. PLoS One 8(1): e52460.
Article Information
Copyright
© 2013 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Xu, B., Wang, Y., Liang, H. and Li, G. (2013). Structural Based Strategy for Predicting Transcription Factor Binding Sites . Bio-protocol 3(12): e794. DOI: 10.21769/BioProtoc.794.
Category
Molecular Biology > DNA > DNA-protein interaction
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link