- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Immunofluorescence (Especially for Cells Growing on a Coverglass)
Published: Vol 2, Iss 4, Feb 20, 2012 DOI: 10.21769/BioProtoc.79 Views: 25210

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
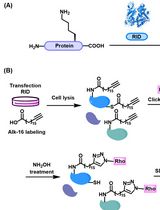
Preparation of Protein Lysates Using Biorthogonal Chemical Reporters for Click Reaction and in-Gel Fluorescence Analysis
Yaxin Xu and Tao Peng
Nov 20, 2024 2062 Views
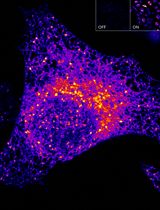
Sensitive and Adaptable Turn-On Maturation (ATOM) Fluorescent Biosensors for Detecting Subcellular Localization of Protein Targets in Cells
Harsimranjit Sekhon [...] Stewart N. Loh
Mar 20, 2025 2304 Views
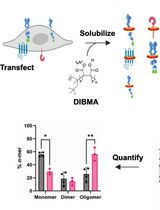
SiMPull-POP: Quantification of Membrane Protein Assembly via Single Molecule Photobleaching
Ryan J. Schuck [...] Rajan Lamichhane
Jan 5, 2026 351 Views
Abstract
If an antibody for your protein of interest is available, immunofluorscence is a useful method to detect the localization and relative abundance of the protein by using a fluorescence microscope. Immunofluoresence can be used in combination with other, non-antibody methods of fluorescence staining, for example, the use of DAPI to label DNA. This protocol describes setting up an immunofluorescence experiment using cells grown on a coverglass.
Keywords: Protein localizationMaterials and Reagents
- Primary antibodies: self-produce or commercially order
Note: For self-produce primary antibodies, optimalize antibody concentration at 1-50 µg/ml); Try 5 µg/ml as a starting point. For commercial primary antibodies, usually 1:1,000 dilution as a starting point, and then adjust the working concentration according to the preliminary data. - Secondary antibodies (below are the most common secondary antibodies):
- Goat anti-rabbit IgG (Life Technologies, Molecular Probes®/Alexa Fluor® 488, catalog number: A-11008 )
- Goat anti-rabbit IgG (Life Technologies, Molecular Probes®/Alexa Fluor® 594, catalog number: A-11012 )
- Goat anti-mosue IgG (Life Technologies, Molecular Probes®/Alexa Fluor® 488, catalog number: A-11001 )
- Goat anti-mosue IgG (Life Technologies, Molecular Probes®/Alexa Fluor® 594, catalog number: A-11005 )
- Goat anti-human IgG (Life Technologies, Molecular Probes®/Alexa Fluor® 488, catalog number: A-11013 )
- Goat anti-human IgG (Life Technologies, Molecular Probes®/ Alexa Fluor® 594, catalog number: A-11014 )
Note: We usually dilute commercial secondary antibodies at 1: 200 as a starting point. Make sure to spin down the stock solution of secondary antibody before dilution to prevent precipitated fluorophores from causing high background. We usually aliquot secondary antibody and stored at -20 °C.
- Goat anti-rabbit IgG (Life Technologies, Molecular Probes®/Alexa Fluor® 488, catalog number: A-11008 )
- 4’, 6-diamidine-2-phenylindole dihydrochloride (DAPI) (Boeringer Manheim, 236 276) (make 1 mg/ml stock, 1,000x)
- Paraformaldehyde (16% solution) (Electron Microscopy Sciences, catalog number: 15170 )
- Methanol (Thermo Fisher Scientific, catalog number: BP 1105-4 )
- Coverglasses (12 mm) (Thermo Fisher Scientific, catalog number: 12-545-82 )
- Poly-L-lysine (MW 150,000-300,000) (Sigma-Aldrich, catalog number: P1399 )
- Fluormount-GTM (mounting solution) (SouthernBiotech, catalog number: 0100-01 )
- Nitric acid
- Hydrochloric acid
- Liquid nitrogen
- PBS-BT solution (see Recipes)
- Normal goat serum (The Jackson Laboratory, catalog number: 005-000-001 ) (see Recipes)
Equipment
- Glass beaker
- Petri dishes
- UV-irradiate
- 24-well plate
- 6-well plate
- 60 mm dish
- Parafilm
Procedure
- Preparing coverglasses
- Make up 100 ml acid solution in a large glass beaker in the hood.
Note: The acid solution is made of 2 parts of nitric acid and 1 part of HCl, and the color is orange-red. - Put the 12 mm coverglasses into the acid solution one by one so that the coverglasses are evenly washed in the acid.
- Let the coverglasses sit in the acid for 2 h or overnight swirling occasionally.
- Decant acid solution into another large glass beaker; neutralize acid with NaOH pellets.
Note: With NaOH, the solution will ‘boil’ vigorously, so be careful. - Wash coverglasses with ddH2O until pH goes up to 7.0.
- Add sufficient 500 µg/ml poly-L-lysine solution to cover all coverglasses, and sit for 2 h with occasionally swirling.
- Decant poly-L-lysine solution (solution can be used repeatedly).
- Wash coverglasses 3 times with ddH2O.
- Dry each coverglass individually in Petri dishes, avoiding coverglasses stuck to the surface.
Note: If coverglasses are stuck to the surface of a Petri dish, they can be easily removed by adding a small amount of liquid nitrogen. - After dry, put all the coverglasses into a Petri dish and UV-irradiate overnight before first use.
Note: For coverglasses that are stored for a long time, re-irradiate for the sake of sterility.
- Make up 100 ml acid solution in a large glass beaker in the hood.
- Growing cells on coverglasses
- Split cells onto tissue culture dishes containing coverglasses or chambered slides.
Notes:- We usually put one coverglass in one well of the 24-well plate; or no more than 2 coverglasses in one well of the 6-well plate, or no more than 5 coverglasses in a 60 mm dish.
- Pipette up and down or shake the dish to make sure cells are not concentrating in the center of the dish well. Make sure there are no air bubbles between the coverglasses and the tissue culture dish.
- We usually put one coverglass in one well of the 24-well plate; or no more than 2 coverglasses in one well of the 6-well plate, or no more than 5 coverglasses in a 60 mm dish.
- Grow cells to 70-100% confluency.
- Split cells onto tissue culture dishes containing coverglasses or chambered slides.
- Fixation and permeablization of the cells
- Transfer the coverglasses or chambered slides into another tissue plate containing sufficient methanol (-20 °C stock), fix for 10 min or a couple of weeks.
Alternately, transfer cells to another plate including PBS. Discard PBS, adding 200 µl PBS plus 20 µl 16% paraformaldehyde. Shaking the plate for 15 min at room temperature (RT).
Notes:- Transfer to methanol directly, no need to do the PBS wash.
- 16% paraformaldehyde needs to be stored at -20 °C, the final working concentration is 1.6% here.
- Formaldehyde preserves antigens by crosslinking the proteins, it has the advantage of preserving most subcellular antigens in their proper localization and GFP. Methanol precipitates antigens by dehydrating cells, it is useful for observing cytoskeletal elements such as microtubles, actin, and other associated structures, but it destroys GFP.
- Transfer to methanol directly, no need to do the PBS wash.
- Carefully transfer the coverglasses or chambered slides from the plates and place cell side up onto secured Parafilm. Wash the coverglasses or chambered slides with 100 µl PBS immediately after transfer, never dry the cells.
- Add 100 µl PBS-BT solution to the coverglasses or chambered slides, let sit for 30 min at RT to permeablize and block cells.
Note: For stringent block, 4-6% normal goat serum (NGS) can be used.
- Transfer the coverglasses or chambered slides into another tissue plate containing sufficient methanol (-20 °C stock), fix for 10 min or a couple of weeks.
- Staining and mounting cells
- Incubate cells in 40 µl primary antibody (1 µg/ml final primary antibody concentration, dilute in PBS-BT) for 30 min at RT.
- Rinse with PBS-BT twice, and then wash with 100 µl PBS-BT twice, 5 min each.
- Cells were incubated in 40 µl secondary antibody (1 µg/ml final secondary antibody concentration, dilute in PBS-BT) for 30 min at RT.
- Rinse with PBS-BT twice, wash with PBS-BT once, 5 min, and then wash with PBS, 5 min.
- Incubate cells in 40 µl DAPI (1 µg/ml final concentration, 1:1,000 dilute in PBS) for 2 min, and then wash with PBS once.
- Add 5-10 µl mounting solution to a clean microscope slide for each coverglass, place stained coverglass cell side down onto mounting solution from one edge; allow mounting solution to cover the entire surface of the coverglass, avoiding air bubbles.
- Let the mounting solution dry and self-seal for 30 min at RT.
- Incubate cells in 40 µl primary antibody (1 µg/ml final primary antibody concentration, dilute in PBS-BT) for 30 min at RT.
Recipes
- PBS-BT solution
10 ml 10x PBS
3 g BSA (to 3%)
1 ml 10% Triton X-100 (to 0.1%)
1 ml 5% NaN3
ddH2O to 100 ml
Stored at 4 °C - Normal goat serum
Make 4-6% solution in PBS
Acknowledgments
This protocol was modified from an immunoprecipitation protocol developed in the laboratory of Dr. Guowei Fang (Department of Biology, Stanford University, Stanford, CA, USA). The protocol was originally developed Dr. Jim Wong. This work was supported by a Burroughs-Wellcome Career Award in Biomedical Research (G.F.) and by grants from National Institutes of Health (GM062852 to G.F.).
References
- Wong, J. and Fang, G. (2006). HURP controls spindle dynamics to promote proper interkinetochore tension and efficient kinetochore capture. J Cell Biol 173(6): 879-891.
- Zhu, H., Coppinger, J. A., Jang, C. Y., Yates, J. R., 3rd and Fang, G. (2008). FAM29A promotes microtubule amplification via recruitment of the NEDD1-gamma-tubulin complex to the mitotic spindle. J Cell Biol 183(5): 835-848.
- Zhu, H., Fang, K. and Fang, G. (2009). FAM29A, a target of Plk1 regulation, controls the partitioning of NEDD1 between the mitotic spindle and the centrosomes. J Cell Sci 122(Pt 15): 2750-2759.
Article Information
Copyright
© 2012 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Zhu, H. (2012). Immunofluorescence (Especially for Cells Growing on a Coverglass). Bio-protocol 2(4): e79. DOI: 10.21769/BioProtoc.79.
Category
Biochemistry > Protein > Fluorescence
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link









