- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Wax Analysis of Stem and Rosette Leaves in Arabidopsis thaliana
Published: Vol 3, Iss 11, Jun 5, 2013 DOI: 10.21769/BioProtoc.782 Views: 15210

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
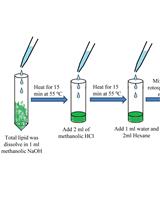
GC-MS-Based Analysis of Methanol: Chloroform-extracted Fatty Acids from Plant Tissues
Manish Kumar Patel [...] Jitendra Kumar Thakur
Sep 20, 2018 15438 Views
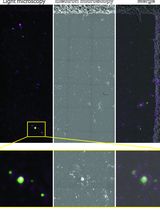
Sorghum bicolor Extracellular Vesicle Isolation, Labeling, and Correlative Light and Electron Microscopy
Deji Adekanye [...] Jeffrey L. Caplan
Oct 5, 2024 2108 Views

PhosphoLIMBO: An Easy and Efficient Protocol to Separate and Analyze Phospholipids by HPTLC From Plant Material
Louise Fougère [...] Yohann Boutté
Sep 5, 2025 1334 Views
Abstract
The primary aerial surfaces of all land plants are coated by a lipidic cuticle, which restricts non-stomatal water loss and protects the plant from pathogens, herbivores, and ultraviolet radiation. The cuticle is made up of two components: cutin, a polymer of hydroxy- and epoxy- long-chain fatty acid derivatives and glycerol, and cuticular waxes, which are derivatives of very-long-chain fatty acids. While chemical analysis of cutin can be a lengthy and technically challenging task, analysis of cuticular waxes is relatively simple, and can be routinely used to characterize different plant species, adaptations of a given species to environmental conditions, or mutant phenotypes. Here, we present a protocol tailored for the analysis of cuticular waxes on the surface of the model organism Arabidopsis thaliana. Because cuticular waxes are found on the outermost surface of the plant, the wax extraction process is very simple, and sample processing can be completed in less than one day. Chemical analysis involves quantitation of wax monomers by gas chromatography coupled with flame ionization detection (GC/FID), and identification of wax monomers by either mass spectrometry or comparison of retention times of individual wax components to those of known standards.
Keywords: ArabidopsisMaterials and Reagents
- Chloroform, ACS grade (Sigma-Aldrich, catalog number: 319988-4L )
- N,O-bis(trimethylsilyl)trifluoroacetamide with trimethylchlorosilane (99:1 BSTFA+TMCS) (Sigma-Aldrich, catalog number: 33148 )
- Pyridine, ACS grade (Thermo Fisher Scientific, catalog number: P368-500 )
- Tetracosane (solid, for use as an internal standard) (Sigma-Aldrich, catalog number: 87089-1G )
Equipment
- Ruler or sticker of known dimensions
- Overhead transparent sheets (only needed for leaf wax analysis)
- Camera, ideally with a small tripod
- Scissors
- Forceps
- 11 ml glass tubes (Wheaton, catalog number: 358606 )
- 1.5 ml GC vials (Agilent, catalog number: 5182-0715 )
- 250 μl GC vial inserts (Agilent, catalog number: 5183-2085 )
- GC vial rack
- 100 μl glass syringe (Hamilton, catalog number: 80630 )
- 500 μl glass syringe (Hamilton, catalog number: 80865 )
- Oven
- Reacti-Vap evaporator (Thermo Fisher Scientific, catalog number: TS-18825 ) with Reacti-Therm heating module (Thermo Fisher Scientific, catalog number: TS-18822 )
- Nitrogen gas (Praxair NI M-T)
- Adobe Photoshop
- Microsoft Excel
- Gas chromatography system
- Agilent 7890A GC system with FID
- HP1 methyl siloxane column (Agilent, 19091Z-313 )
- hydrogen (GC carrier gas and FID fuel) and compressed air (for FID)
- Agilent 7890A GC system with FID
Procedure
Note: A few steps of the protocol vary between leaves and stems. Chloroform, pyridine, and BSTFA should all be handled in a fume hood.
- Prepare a 1 μg μl-1 internal standard by dissolving 500 μg tetracosane in 500 μl chloroform. If you expect to be analyzing waxes routinely, scale this up to make a large stock, and aliquot known amounts of the stock in 1.5 ml GC vials. Allow the chloroform to evaporate off, seal the vials, and store at 4 °C. Resuspend the aliquots in chloroform to their initial volume as required for immediate use.
- Prepare one 11 ml glass tube for each sample you will collect. We recommend using 3-6 technical replicates for each genotype you are investigating. Rinse all glass tubes with chloroform three times before you begin. Then, fill the tubes with approximately 10 ml chloroform, and add 10 μl internal standard to each sample. It is critical to remember to include a set of wild-type samples as a positive control every time you do a wax analysis experiment. Wax load and composition can vary greatly depending on the conditions your plants are grown in, so this control is necessary to distinguish between changes caused by different genotypes vs. changes in the environmental conditions or developmental stage of the plants at the time of harvest. If you wish to check the GC trace of the solvent and standard alone, you can prepare an extra tube at this step, which will not have a tissue sample dipped in it.
- STEMS: Stem wax extraction and image capture for surface area calculation. Take care to minimize sample handling, as handling will remove wax from your sample. Use forceps, not your hands, whenever possible.
- Cut stems approximately 10cm from the apical meristem, remove leaves and branches, and lie flat on a light, even background with a ruler or item of known dimensions for size comparison. Label each sample by writing on the background. Only one stem for each replicate is needed.
- Photograph the samples so that you have all stems, their labels, as well as the ruler or size marker in view (Figure 1). Photographs will be used later to calculate stem surface area.
- Dip stems in solvent tubes for 30 s to extract wax. Stems can be thrown out after dipping.

Figure 1. How to collect stem area data. Stems are trimmed and photographed for measurement before dipping in chloroform. The round green sticker was used as a reference for size.
ROSETTE LEAVES: Rosette leaf wax extraction and image capture for area calculation. As with stems, take care to minimize sample handling, and use forceps, not your hands, whenever possible.
- Arabidopsis rosette leaves carry approximately 10% of the wax load found on inflorescence stems. Therefore, you will need to collect more leaf tissue to complete your analysis. Aim to collect at least 6 leaves for each replicate. Also, you will likely find more variability in leaf wax composition and load than you will for stem, so using more technical replicates for each genotype (5-8) is advisable.
- As rosette leaves of Arabidopsis are often curved and warped, it is difficult to calculate their area based on a 2D image. Our solution is to first dip leaves in chloroform for 30 s, then, after the wax has been extracted and the leaf is limp and coated with solvent, spread the leaf flat on a plastic transparency sheet. The transparency sheet will melt very slightly with the chloroform, helping to hold the leaf flat. Label each sample by writing on the transparency sheet (Figure 2).
- Repeat for all leaves for each replicate, and all replicates for each genotype. Photograph the samples after wax extraction, with the labels and ruler or size marker in view. Photographs will be used later to calculate stem surface area.
- Try to work quickly, as the leaves will eventually dry out after wax extraction, and your area calculation will be inaccurate if you do not photograph the samples quickly enough.

Figure 2. How to collect leaf area data. Leaves are laid flat (which is sometimes easier if small notches are cut in the leaf blade) and photographed for measurement after dipping in chloroform.
- Cut stems approximately 10cm from the apical meristem, remove leaves and branches, and lie flat on a light, even background with a ruler or item of known dimensions for size comparison. Label each sample by writing on the background. Only one stem for each replicate is needed.
- Evaporate off all of the solvent by passing a gentle stream of nitrogen gas from an evaporator manifold over each sample. Clean the steel needles of the manifold in chloroform before and after each use.
Note: A greater flow of nitrogen will not necessarily reduce the time required to evaporate your sample. Use a low flow and position the needles so that there is a shallow dimple in the top of your sample from the outflow of gas. If your sample is splashing or bubbling, you are wasting nitrogen, and risk losing your sample. - After the solvent has dried off, you should be able to see a white film of wax at the bottom of each tube. Resuspend this wax residue in 100-200 μl fresh chloroform using a glass syringe, and transfer to a GC vial insert in a GC vial. Evaporate solvent off as above. Be very careful with the outflow of nitrogen this time; because the vial inserts are narrow, it is very easy to apply too much nitrogen and lose your concentrated sample.
- After the solvent has dried off completely, add 10 μl pyridine and 10 μl BSTFA+TMCS to each sample using a glass syringe. Seal the GC vials tightly, and incubate for 1 h in an oven set to 80 °C.
- Remove the vials from the oven and allow them to cool to room temperature for 1-2 min. Evaporate off the solvent again as above.
Note: Pyridine, BSTFA, and TMCS will evaporate much more slowly than chloroform. - Resuspend the final derivatized residues in 40 μl chloroform, and seal the GC vials tightly.
- Inject samples on the GC column. We use a 2.7:1 split for stem wax analysis, and run leaf waxes in splitless mode. For both, we inject 1 μl sample. The program we use is as follows:
50 °C for 2 min
ramp 40 °C/min to 200 °C
hold 1 min
ramp 3 °C/min to 320 °C
hold 15 min - Calculate the area of your samples. Upload your photos to Adobe Photoshop, and use the magic wand tool to select and record the area of each of your samples and the size marker in pixels. As the size of the marker is known, you can use a simple ratio to determine the 2D areas of your samples in centimeters or inches. For leaves, 2D area is used as is. For stems, the 2D area represents the stem diameter multiplied by length. Therefore, you will need to multiply the 2D area by pi to obtain the surface area for the entire stem.
- After the GC has finished running your samples, download your data. Typically, a wild-type sample will be run through GC/MS using the same program as for GC/FID so that you can determine the chemical identities of each of the peaks in your trace from GC/FID. Alternatively, you can compare the peaks in the GC/FID chromatogram from your wild-type sample to a chromatogram where the peaks have been identified, and determine the exact retention times that you expect for each monomer on your system. Identify the peaks for the wax monomers you are interested in, and the internal standard, and copy their retention times and peak areas into a Microsoft Excel spreadsheet. Repeat this for all of your replicates and different genotypes.
- 10 μg tetracosane standard was added to each sample. Therefore, you can use a ratio with the tetracosane standard mass and peak area and the peak area of each of your wax monomers to determine the mass of each monomer for every sample. Finally, the amount of each monomer is divided by the sample area, so that you express wax monomer amount in μg/cm2 for stems, or μg/dm2 for leaves. If you include all wax monomers in your analysis, you can tally these values to determine the total wax load for each sample.
References
- Haslam, T. M., Manas-Fernandez, A., Zhao, L. and Kunst, L. (2012). Arabidopsis ECERIFERUM2 is a component of the fatty acid elongation machinery required for fatty acid extension to exceptional lengths. Plant Physiol 160(3): 1164-1174.
Article Information
Copyright
© 2013 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Readers should cite both the Bio-protocol article and the original research article where this protocol was used:
- Haslam, T. M. and Kunst, L. (2013). Wax Analysis of Stem and Rosette Leaves in Arabidopsis thaliana. Bio-protocol 3(11): e782. DOI: 10.21769/BioProtoc.782.
- Haslam, T. M., Manas-Fernandez, A., Zhao, L. and Kunst, L. (2012). Arabidopsis ECERIFERUM2 is a component of the fatty acid elongation machinery required for fatty acid extension to exceptional lengths. Plant Physiol 160(3): 1164-1174.
Category
Plant Science > Plant biochemistry > Lipid
Biochemistry > Lipid > Extracellular lipids
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link











