- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Metabolic Labeling of Yeast RNA with Radioactive Uracil
Published: Vol 3, Iss 7, Apr 5, 2013 DOI: 10.21769/BioProtoc.623 Views: 10645

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
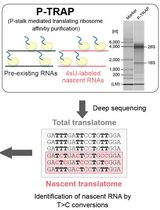
Metabolic RNA Labeling and Translating Ribosome Affinity Purification for Measurement of Nascent RNA Translation
Hirotatsu Imai and Akio Yamashita
Oct 20, 2024 2568 Views
Versatile Click Chemistry-based Approaches to Illuminate DNA and RNA G-Quadruplexes in Human Cells
Angélique Pipier and David Monchaud
Feb 5, 2025 2602 Views
APEX2 RNA Proximity Labeling in Mammalian Cell Lines With Low Biotin Permeability
Adrian Beat Tschan [...] Alexa B.R. McIntyre
Jul 5, 2025 2428 Views
Abstract
To examine gene expression, Northern blot or Real-Time PCR can be used to detect low abundant RNA such as mRNA. However, for high abundant RNAs such as rRNA and tRNA, Northern blot will not be able to discriminate the newly synthesized RNA from total RNA. Therefore, metabolic labeling is necessary to evaluate the expression of rRNA and tRNA genes. In this protocol, I describe a step-by-step method for labeling yeast RNA with radioactive uracil and examine the synthesis of these high abundant RNAs.
Keywords: AutoradiographyMaterials and Reagents
- Yeast strain of interest
- RapidGel (500 ml) (Affymetrix, catalog number: 75848 )
- Urea (CO(NH2)2) (Sigma-Aldrich, catalog number: U6504 )
- Dextrose (C6H12O6) (Sigma-Aldrich, catalog number: G7021 )
- Yeast nitrogen base without amino acids (Sigma-Aldrich, catalog number: Y0626 )
- Synthetic dropout supplement without uracil (Sigma-Aldrich, catalog number: Y1501 )
- Uracil (C4H4N2O2) (Sigma-Aldrich, catalog number: U1128 )
- Rapamycin
- TEMED/Tetramethylethylenediamine (C6H16N2) (Thermo Fisher Scientific, catalog number: 110-18-9 )
- Ammonium persulfate (APS) ((NH4)2S2O8) (Sigma-Aldrich, catalog number: A3678 )
- Formamide (CH3NO) (Thermo Fisher Scientific, catalog number: 75-12-7 )
- DEPC/ Diethylpyrocarbonate (O(COOC2H5)2) (Sigma-Aldrich, catalog number: D5758 )
- Bromophenol Blue (C19H10Br4O5S) (Sigma-Aldrich, catalog number: B0126 )
- Xylene Cyanol FF (C25H27N2NaO6S2) (Sigma-Aldrich, catalog number: X4126 )
- [5, 6-3H]-Uracil (PerkinElmer, catalog number: NET368250UC )
- RNA marker (Promega Corporation, catalog number: G3191 )
- Tris base (Thermo Fisher Scientific, catalog number: 77-86-1 )
- Boric acid (Thermo Fisher Scientific, catalog number: 10043-35-3 )
- EDTA (Sigma-Aldrich, catalog number: EDS-1KG )
- Formamide loading dye
- Small RNA separating gel
- SD-Ura- (see Recipes)
- SD-1/3 Uracil (see Recipes)
- DEPC water (see Recipes)
- 10x TBE in DEPC water (see Recipes)
- Small RNA separating gel (see Recipes)
- 2x Formamide loading dye (see Recipes)
Equipment
- Bench top centrifuge
- 30 °C Shaker
- Power supply with constant voltage > 450 V
- Gel dryer (Bio-Rad Laboratories, catalog number: 165-1745 )
- Exposure cosset/intensifier screen (Sigma-Aldrich, catalog number: C5479-1EA )
- 50 ml conical tubes
- 1.5 ml Eppendorf tube
Procedure
- Inoculate yeast single colony in SD medium (or SD with appropriate dropouts). Shake 300 rpm at 30 °C overnight.
- Dilute overnight culture in 50 ml conical tubes to 10 ml, OD600 =0.1 with SD-1/3 uracil and continue shaking until OD600 =0.4 (note: Reduction in cold uracil will allow hot uracil to be taken up by cells easily).
- Cells were treated with drug and control vehicle, for example 100 nM rapamycin (final concentration) and its solvent methanol, continue shaking 300 rpm at 30 °C for desired time. In this specific experiment, rapamycin were added for 30 min.
- Collect yeast cells by spinning down at 1,000 x g for 1 min at room temperature, remove supernatant (critical: Avoid putting yeast cells on ice. This is because ice will slow down growth, which will reduce significantly the uptake of hot uracil in the step 8 below).
- Re-suspend yeast cells in 1 ml SD-Ura- (critical: Pre-warm medium to 30 °C), transfer to 1.5 ml eppendorf tube.
- Spin down briefly by a bench top centrifuge at 5,000 x g, 15 sec, remove supernatant. Re-suspend yeast cells in 1 ml pre-warmed SD-Ura-.
- Repeat 6 for 2 times and with the final re-suspension in 0.5 ml pre-warmed SD-Ura- (from the next step, collect radioactive liquid and solid waste in all steps, dispose according to environmental regulation).
- Carefully add [5, 6-3H]-Uracil into each tube to the final concentration of 15 μCi/ml, vortex to mix, then put on a rack at 30 °C for 5 min.
- Briefly spin down by a bench top centrifuge at 5,000 x g, 15 sec, remove supernatant.
- Wash cells with SD-Ura-3 times as in step 5, ready to extract total RNA.
- Total RNA was extracted by hot phenol method described in Wei (2012).
- RNAs can be stored at -80 °C for up to 6 months.
- Prepare ''small RNA separating gel'' on a large gel set (around 20 x 30 cm, mini gel did not work well).
- Pre-run the gel for about 1 h at constant 450 V until the gel is heated to 50 °C.
Note: I found this step to be critical. One reason could be that pre-running the gel to this temperature could help get rid of excessive Urea in the gel, making RNA possible to go through. I usually attached a thermometer to ensure that the temperature has reached 50 °C. - Mix RNA samples with ''2x Formamide loading dye'' and heat at 70 °C for 2 min, then put samples on ice.
Turn off power. Rinse the wells with 1x TBE using a syringe with needle, make sure all urea is rinsed out from the wells.
Note: Urea is very dense and it will be impossible to load samples if urea is not rinsed out. If residual urea remains in the well, the resulting bands will be waving. - Load RNA samples (25 μg) with appropriate RNA marker and run gel at constant 450 V for about 2 h (BPB runs around 12 nt and cyanol around 55 nt).
- Stain the gel with EtBr and take picture under UV light. This is total RNA which served as controls for newly synthesized RNA.
- Sandwich the gel with autoclaved filter paper on one side and Saran-Wrap on the other side, put on a gel-dryer with filter paper side attach to the vacuum surface. Dry the gel at 80 °C for at least 2 h. Sometimes gels get cracked, may be because of insufficient drying or leaky vacuum.
- The dried gel will stick to the filter paper. Wrap them in Sara-Wrap. 3H autoradiography in an exposure cassette with appropriate intensifying screen, for example Sigma Transcreen.
- Develop after 4-7 days of exposure.
Recipes
- SD-Ura-
Synthetic dextrose medium with Uracil dropout:
20 g Dextrose
1.7g Yeast Nitrogen Base
1.92 g synthetic dropout supplement without Uracil
5.0 g Ammonium Sulfate
Add ddH2O2 to 1 L and autoclave. - SD-1/3 Uracil: SD medium with 1/3 of regular Uracil
Similar to SD-Ura-except adding 25 mg Uracil. - DEPC water
Add 1 ml DEPC to 1 L ddH2O2, mix and put at room temperature overnight. Autoclave. - 10x TBE in DEPC water
In 800 ml DEPC water, add:
108 g Tris base
55 g boric acid
40 ml of 0.5 M EDTA (pH 8.0)
Mix to dissolve and add DEPC water to 1 L. Autoclave. - Small RNA separating gel
- Mix 2.5 ml 10x TBE, 6.25 ml RapidGel (40%) and 15 g Urea, heat to 50 °C and mix to dissolve.
- Add DEPC water to 25 ml then filter through 0.45 μM Syringe.
- Add 25 μl TEMED and 50 μl 25% APS, mix vigorously transfer to gel set with appropriate comb.
- Mix 2.5 ml 10x TBE, 6.25 ml RapidGel (40%) and 15 g Urea, heat to 50 °C and mix to dissolve.
- 2x Formamide loading dye
95% (v/v) formamide in DEPC water, add tiny amount of Bromophenol Blue (0.01~0.1%) and Xylene Cyanol FF (0.01~0.1%), vortex to mix.
Acknowledgments
This protocol is derived from the following papers, Wei et al. (2009a) and Wei et al. (2009b), and the relevant references therein. The work was supported by NIH grants R01-CA099004 and R01-CA123391 to Dr. X.F. Steven Zheng at Robert Wood Johnson Medical School, University of Medicine and Dentistry of New Jersey.
References
- Wei, Y., Tsang, C. K. and Zheng, X. F. (2009a). Mechanisms of regulation of RNA polymerase III-dependent transcription by TORC1. EMBO J 28 (15): 2220-2230.
- Wei, Y. and Zheng, X. F. (2009b). Sch9 partially mediates TORC1 signaling to control ribosomal RNA synthesis. Cell Cycle 8(24): 4085-4090.
- Wei, Y. (2012). A Simple Preparation of RNA from Yeast by Hot Phenol for Northern Blot. Bio-protocol 2(9): e209.
Article Information
Copyright
© 2013 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Readers should cite both the Bio-protocol article and the original research article where this protocol was used:
- Wei, Y. (2013). Metabolic Labeling of Yeast RNA with Radioactive Uracil. Bio-protocol 3(7): e623. DOI: 10.21769/BioProtoc.623.
- Wei, Y., Tsang, C. K. and Zheng, X. F. (2009a). Mechanisms of regulation of RNA polymerase III-dependent transcription by TORC1. EMBO J 28 (15): 2220-2230.
Category
Molecular Biology > RNA > RNA labeling
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link











