- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
A Simple Method for Estimating the Spatiotemporal Distribution of Phenoloxidase Proteins in Insect Tissues
Published: Vol 16, Iss 7, Apr 5, 2026 DOI: 10.21769/BioProtoc.5646 Views: 37
Reviewed by: Anonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
In situ Dephosphorylation Assay with Recombinant Nil Phosphatase
Nilay Nandi [...] Helmut Krämer
Sep 20, 2022 2161 Views
Isolation and Quantification of Mandelonitrile from Arabidopsis thaliana Using Gas Chromatography/Mass Spectrometry
Ana Arnaiz [...] Isabel Diaz
Jun 20, 2023 1610 Views
Live Imaging and Analysis of Meiotic Cytokinesis in Drosophila Testes
Govind Kunduri and Jairaj K. Acharya
Jan 20, 2024 3112 Views
Abstract
Laccase2 (Lac2), a member of the phenoloxidase (PO) family, is an essential oxidase for melanin pigmentation in insects. The identification of the in vivo spatial distribution of Lac2 is crucial for understanding the molecular mechanisms underlying color pattern formation. However, it is technically difficult to determine the distribution because Lac2 expression peaks at late pupal stages, when adult cuticle sclerotization has already begun. Here, we report a simple and rapid protocol for estimating the distribution of endogenous PO proteins, prophenoloxidases (proPOs) and phenoloxidases (POs), in insect tissues. In this method, the spatial distribution of endogenous PO proteins is estimated based on staining patterns formed by dopamine melanin synthesis in tissues incubated in a solution containing isopropanol and dopamine. We validated that tissues collected at approximately 80% of the total pupal duration yielded staining patterns corresponding to adult melanin-forming regions in three insect species. By comparing staining patterns across developmental stages, this protocol enables estimation of the timing of color pattern formation. Furthermore, the contrast between stained and unstained regions within the same tissue allows region-specific sampling, thereby facilitating an investigation of the underlying molecular mechanisms regulating spatial PO distribution. Taken together, this method facilitates the study of melanin biosynthesis and enables the identification of the genes involved in regulating color pattern formation. This protocol does not require antibodies, transgenic lines, or specialized equipment and can be completed within a short time frame. Its effectiveness has been validated in multiple coleopteran and lepidopteran species, demonstrating its broad applicability as a versatile tool for studying insect pigmentation and color pattern formation.
Key features
• A simple tissue staining protocol to estimate the spatial distribution of endogenous PO proteins without the use of antibodies or transgenic lines.
• Comparing staining patterns across different developmental stages to estimate both the spatial distribution of PO proteins within tissues and the timing of color pattern formation.
• RNAs can be extracted from the tissue after staining, enabling gene expression analyses between stained and unstained regions.
• This protocol has been validated in two coleopteran species and one lepidopteran species, demonstrating broad applicability across diverse insect taxa.
Keywords: PhenoloxidaseBackground
Insects exhibit a remarkable diversity of body coloration patterns, showing extensive variation both among and within species. For example, the ladybird beetle Harmonia axyridis displays approximately 200 distinct color patterns within one species [1]. These patterns often function as warning coloration or mimicry and play important roles in ecological interactions and evolutionary adaptation. Insect color patterns are composed of various pigments, among which melanin pigments are the most widely used in color pattern formation [2]. In insects, the melanin biosynthesis pathway produces four major products—dopa melanin (black), dopamine melanin (brown), NBAD (yellow), and NADA (colorless)—through a highly conserved biochemical process [3].
Previous studies have shown that Lac2, a member of the PO family, catalyzes the reactions leading to the formation of the four major products of the melanin biosynthesis pathway and is essential for cuticle pigmentation and sclerotization [2]. Therefore, identifying the timing and spatial distribution of Lac2 protein expression is crucial for understanding the molecular mechanisms underlying insect melanin pattern formation. Conventional studies of PO activity have largely relied on enzymatic assays using hemolymph or homogenized tissues, typically quantifying total PO activity with substrates such as L-DOPA or dopamine [4,5]. However, these approaches do not provide spatial information on where PO proteins are distributed or potentially functional within tissues.
Identification of Lac2 protein distribution is technically challenging because Lac2 expression peaks immediately before adult eclosion, when the cuticle has already thickened and begun to harden [6]. At this stage, detection of mRNA or protein localization by in situ hybridization or immunostaining is difficult, and transgenic reporter approaches are not established for most non-model insects.
The molecular mechanisms underlying the activation of PO proteins have been well characterized. In insects, PO proteins are synthesized as an inactive precursor, proPOs, and are irreversibly activated through proteolytic cleavage at the N-terminal region by a protease cascade centered on serine proteases, which exposes the catalytic active site [7]. This activation process is tightly regulated in a developmental stage- and tissue-specific manner and is closely associated with immune responses and cuticle sclerotization [7–9]. In contrast, an in vitro method for artificial activation has been developed, in which isopropanol induces conformational changes in proPOs that expose the active site and activate the enzyme [10]. This activation mechanism differs from the physiological in vivo process but can be easily induced experimentally. Taking advantage of this property, True et al. [11] employed proPO as a transgenic reporter gene and established a method in which insect tissues are incubated in a staining solution containing isopropanol and dopamine, leading to artificial activation of proPO and subsequent dopamine melanin formation, thereby visualizing the spatial pattern of PO reporter activity. However, this approach has not been adapted to estimate the distribution of endogenous PO proteins.
Here, we established a protocol to estimate the distribution of endogenous PO proteins. This protocol is based on the method developed by True et al. [11] and allows estimation of the distribution of endogenous PO proteins based on the spatial pattern of dopamine melanin formed by PO activity staining. A major advantage of this protocol is that it does not require antibodies, transgenic lines, or specialized equipment, and clear staining patterns can be detected within 1–2 h. Because dopamine can be oxidized by multiple phenoloxidase/laccase-type enzymes [12], it is challenging to determine which enzymes are responsible. At late pupal stages, the genes encoding laccase-type enzymes (e.g., Lac2) are highly expressed in the epithelial tissues [5,6,13], but contributions from other oxidases cannot be excluded without genetic or biochemical validation. By applying this protocol across multiple developmental stages, the developmental dynamics of staining patterns can be tracked, which may allow estimation of the onset of spatially defined color pattern formation [1]. Furthermore, we have previously demonstrated that RNAs can be extracted from stained tissues [1]. By separating stained and unstained regions, the samples can be used for region-specific gene expression analysis, thereby facilitating the identification of genes expressed in a pattern-specific manner [1]. This approach enables the identification of genes involved not only in melanin biosynthesis but also in pattern-specific gene regulation.
The protocol has been validated in multiple non-model insect species, including coleopterans such as H. axyridis and Propylea japonica, as well as the lepidopteran Eurema hecabe. Therefore, this method represents a broadly applicable and versatile tool for advancing research on insect pigmentation, color pattern formation, and the evolution of color patterns.
Materials and reagents
Biological materials
1. Harmonia axyridis laboratory stock established from individuals collected from the field in Aichi Prefecture, Japan, in spring, and maintained under laboratory conditions as described in Nakamura et al. [14]
2. Propylea japonica larvae purchased from Sumika Techno-Service and reared under the same conditions as H. axyridis, following the method described by Nakamura et al. [14]
3. Eurema hecabe pupae provided by Dr. Tatsuro Konagaya, who reared the insects to the pupal stage according to the method described by Konagaya and Maruyama [15]
4. Pea aphid Acyrthosiphon pisum parthenogenetic colony maintained on broad bean plants used as prey for H. axyridis, as described in Nakamura et al. [14]
Reagents
1. Potassium dihydrogen phosphate (KH2PO4) (Wako, catalog number: 169-04245)
2. Dipotassium hydrogen phosphate (K2HPO4) (Wako, catalog number: 164-04295)
3. 3,4-dihydroxyphenethylamine hydrochloride (dopamine) (Wako, catalog number: 040-15433)
4. Triton X-100 (Sigma-Aldrich, catalog number: X100)
5. 2-Propanol (isopropanol) (Wako, catalog number: 168-21675)
6. Sodium chloride (NaCl) (Wako, catalog number: 195-01663)
Safety note: When handling isopropanol, dopamine, and Triton X-100, appropriate personal protective equipment (e.g., gloves and safety goggles) should be worn. Isopropanol should be used in a well-ventilated area, avoiding inhalation of vapors, and should be kept away from sources of ignition.
Solutions
1. 0.5 M KH2PO4/K2HPO4 (see Recipes)
2. K-PO4 buffer (see Recipes)
3. K-PO4 T buffer (see Recipes)
4. Dopamine solution (see Recipes)
5. Staining solution (see Recipes)
Recipes
1. 0.5 M KH2PO4/K2HPO4
Add 0.5 M K2HPO4 dropwise to 100 mL of 0.5 M KH2PO4 until the pH reaches 6.3 (approximately 88 mL of 0.5 M K2HPO4). Sterilize the buffer by autoclaving at 121 °C and 1.5 atm for 20 min. The solution is stable at room temperature, and fresh preparation prior to the assay is unnecessary.
2. K-PO4 buffer
| Reagent | Final concentration | Quantity or volume |
|---|---|---|
| 0.5 M KH2PO4/K2HPO4 | 100 mM | 3 mL |
| 5 M NaCl | 150 mM | 450 μL |
| H2O | 11.55 mL |
The solution is stable at room temperature, and fresh preparation prior to the assay is unnecessary.
3. K-PO4 T buffer
| Reagent | Final concentration | Quantity or volume |
|---|---|---|
| 10% Triton X-100 | 0.3% (v/v) | 30 μL |
| K-PO4 buffer | 970 μL |
The solution is stable at room temperature, and fresh preparation prior to the assay is unnecessary.
4. Dopamine solution
| Reagent | Final concentration | Quantity or volume |
|---|---|---|
| Dopamine | 10 mg/mL | X mg (4 ≤ X ≤ 15) |
| K-PO4 buffer | 100× X μL |
Because dopamine is required in a very small amount, weigh dopamine within the range of 4–15 mg. Calculate the volume of K-PO4 buffer to be added according to the measured mass of dopamine. The solution should be freshly prepared prior to each assay.
5. Staining solution
| Reagent | Final concentration | Quantity or volume |
|---|---|---|
| Dopamine solution | 0.4 mg/mL | 40 μL |
| K-PO4 buffer | 360 μL | |
| Isopropanol | 600 μL |
The solution should be freshly prepared prior to each assay.
Laboratory supplies
1. 200 and 1,000 μL pipette tips (Watson, catalog numbers: 123R-757CS and 110-705C)
2. Slide glass (Matsunami, catalog number: S1111)
3. Cover glass (Matsunami, catalog number: C018181)
4. 1.5 mL microtube flat-bottom slight-stiff-touch cap (Watson, catalog number: 131-415C)
5. 24-well and 48-well microplates (Corning, catalog numbers: 258201 and 258301)
6. Paper wipe (Daio Paper, catalog number: DSI20703128)
7. Artificial diet for ladybird beetles [16]
Equipment
1. Dissection/stereomicroscope (Leica or equivalent)
2. Pipetman [GILSON, catalog numbers: F123602 (P1000) and F123601 (P200)]
3. Analytical balance (Shimadzu, model: AUW120D)
4. pH meter (METTLER TOLEDO, model: S220)
5. Vortex mixer (LMS, model: VTX-3000L)
6. E-Centrifuge (Wealtec, catalog number: 1090003)
7. Petri dish (MonotaRO, catalog number: 34680572)
8. Forceps (Fine Science Tools, catalog number: 11252-00)
9. Time-lapse camera (SANWA SUPPLY, model: 400-CAM109)
10. Stir bar (AS ONE, catalog number: 9-870-07)
11. Magnet stirrer (AS ONE, model: CT -1AT)
Procedure
Here, we describe the protocol using the pupal forewing primordia of H. axyridis as an example; we validated this protocol with the other two species, P. japonica and E. hecabe.
A. Determination of pupal duration
Note: Lac2, a PO protein involved in melanin synthesis and cuticle sclerotization, is expressed during the late pupal stage in many insect species [6,17–19]. Therefore, it is important to determine the pupal duration in advance to ensure reproducible staining patterns. Here, we describe a method to determine pupal duration, using H. axyridis as an example.
1. Rear H. axyridis at 25 °C under 16L:8D and feed with aphids and an artificial diet (Niimi et al. [16]), as described previously [14].
2. Position a time-lapse camera approximately 40 cm above the rearing container. Set the camera to a resolution of 1,280 × 960 pixels, with the LED flash turned on (allowing image acquisition under dark conditions), and programmed to automatically capture images at 3-min intervals to obtain time-lapse images (Figure 1A, B).
Note: To facilitate accurate tracking of pupal position in the images, the angle of light is adjusted to minimize reflected light, and obstructions within the field of view are removed as much as possible (Figure 1A, B).
3. Define the time point of pupation as 0 h after pupation (0 h AP) and, in H. axyridis, collect pupae at approximately 80% of the total pupal duration.
Note: In a previous study applying this protocol to the wing primordia of H. axyridis, weak staining across the entire wing primordium was observed at 70 h after pupation (approximately 65% of the total pupal duration), whereas reproducible staining patterns corresponding to adult melanin-forming regions were obtained at 80 h after pupation (approximately 75% of the total pupal duration) and later stages [1]. Thus, in this section, the protocol is demonstrated using pupae at approximately 80% of the total pupal duration.
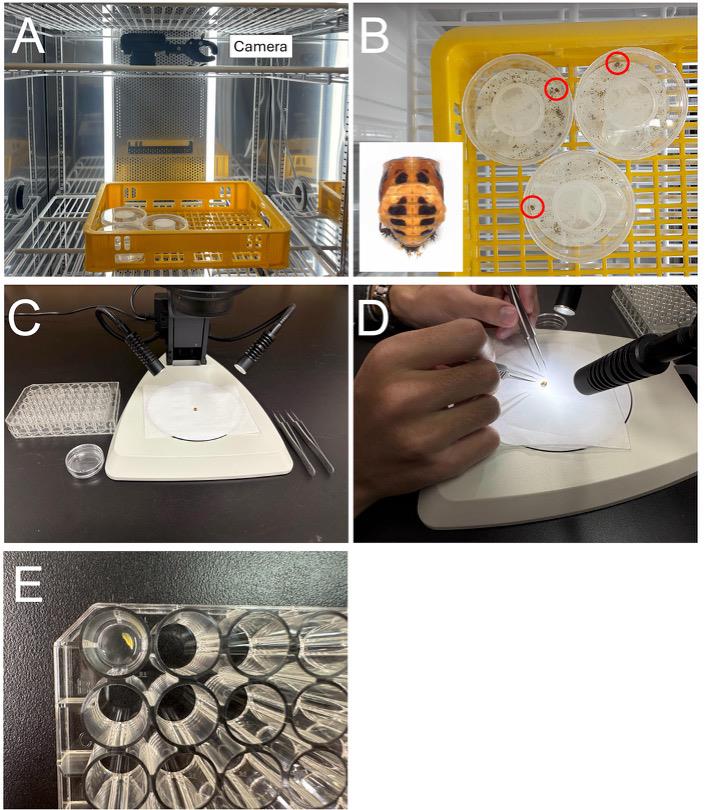

Figure 1. Experimental setup and procedure. (A) Time-lapse imaging using a time-lapse camera. (B) Representative images captured by the time-lapse camera (red circles indicate the position of H. axyridis pupa). The bottom-left image shows the pupa. (C) Instruments used for dissection and staining. (D) Dissection procedure. (E) Well plate containing wing primordia during phenoloxidase (PO) activity staining.
B. PO activity staining
Here, we describe the dissection and staining protocol for pupal forewing primordia of H. axyridis.
1. Place a pupa on a paper wipe and, under a stereomicroscope, grasp the proximal region of the pupal forewing primordia with forceps. Dissect the forewing primordia from the pupa together with the pupal cuticle (Video 1, Figure 1D).
Notes:
1. Using a piece of paper as a support allows us to easily position the pupa within the microscope field of view without direct manual handling.
2. Depending on the species and tissue, use forceps or scissors as appropriate.
2. Hold the proximal region of the dissected forewing primordia with forceps and transfer it into the K-PO4 buffer in a Petri dish. Carefully isolate the wing primordia from the pupal cuticle (Video 1).
3. Transfer the isolated wing primordia into a 48-well plate containing 500 μL of staining solution per well or a 24-well plate containing 1 mL of staining solution per well (Figure 1E).
Notes:
1. The volume of staining solution and well size are adjusted according to tissue size. For H. axyridis and P. japonica, we used a 48-well plate with 500 μL of staining solution. For E. hecabe, which has larger wing primordia, we used a 24-well plate with 1 mL of staining solution.
2. Staining can also be performed in microcentrifuge tubes; however, visual monitoring of staining progression is difficult.
3. ProPOs are artificially activated by isopropanol in the staining solution [10]; therefore, the observed activity may not fully reflect endogenous PO activity under physiological conditions.
4. Put the lid on the plate and incubate at room temperature (25–27 °C), checking the staining progression every 20 min (Figure 1E).
Notes:
1. Because the timing of PO activity staining may vary depending on pupal stage and species, staining should be monitored at 20-min intervals to determine the optimal time point at which to terminate the staining reaction. In H. axyridis, staining beyond 180 min leads to nonspecific darkening of regions unrelated to future adult melanization (Figure 2), which could be caused by the presence of PO proteins involved in cuticle sclerotization outside the adult melanin-forming regions, as well as by spontaneous oxidation of dopamine.
2. In H. axyridis, staining post-80 h pupal stages for 120 min consistently produced staining patterns corresponding to adult melanized regions, and the patterns became clearer as pupal development progressed [1].
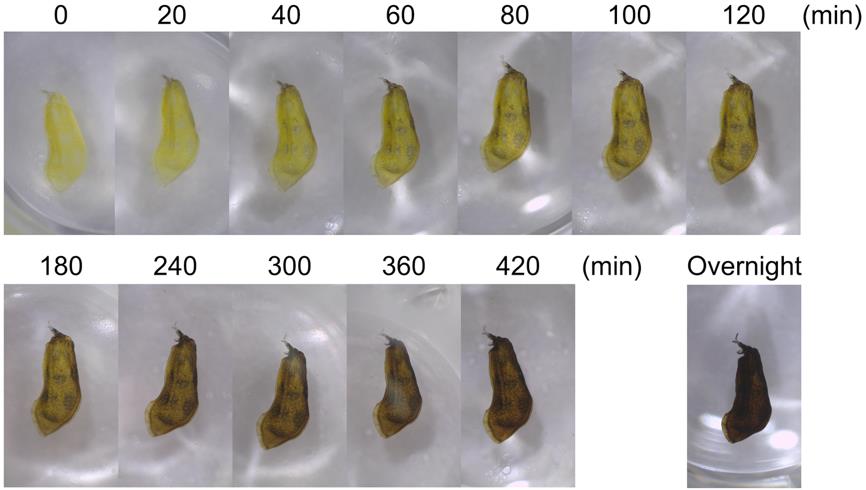

Figure 2. Time-lapse images of the phenoloxidase (PO) activity staining process in pupal forewing primordia of H. axyridis. Images were captured at 20-min intervals up to 120 min, at 1-h intervals thereafter, and again after overnight incubation.
5. Once staining is observed exclusively in regions corresponding to future melanized regions in adults (Figure 2), terminate the reaction by washing the tissue three times with K-PO4 T buffer.
Notes:
1. Although the reaction is terminated by washing, the tissue may continue to darken gradually over time; images should be acquired immediately after washing.
2. Extraction of RNAs should be performed after washing.
C. Observation of staining results
To validate this protocol, we performed staining in three insect species (Figure 3). As described in the previous section, we performed staining at approximately 80% of the total pupal duration for all three examined species. Specifically, pupae were sampled at 4 days after pupation for H. axyridis (total pupal duration: 4.5 days [20]), 3 days after pupation for P. japonica (total pupal duration: 3.76 ± 0.04 days [21]), and 5 days after pupation for E. hecabe (total pupal duration: 6.5 ± 0.4 days [22]). We obtained PO activity staining patterns corresponding to adult melanin-forming regions in all three species by incubating wing primordia for 120 min (Figure 3G–I). Based on these results, we propose that performing staining at approximately 80% of the total pupal duration serves as a valuable baseline when applying this protocol to other insect species, although incubation times may need optimization.
Conducting stage-specific analyses in other species could allow for the precise identification of the timing of color pattern determination, as previously demonstrated in H. axyridis [1], in which we obtained a transition in the staining pattern from weak staining across the entire wing primordium to a pattern corresponding to adult melanin-forming regions between 70 and 80 h after pupation [1]. Applying this approach to define critical developmental time points in other insects would facilitate further analyses, such as RNA-seq, aiming to identify genes involved in color pattern formation.
The staining patterns obtained at this stage are considered to reflect the spatial distribution of proPOs, rather than endogenous active POs. This is supported by the observation that staining without isopropanol-mediated artificial activation yielded no patterns corresponding to adult melanin-forming regions (data not shown). Consequently, while this method effectively estimates proPO distribution, it does not provide insight into the precise timing of in vivo proPO conversion to active POs.
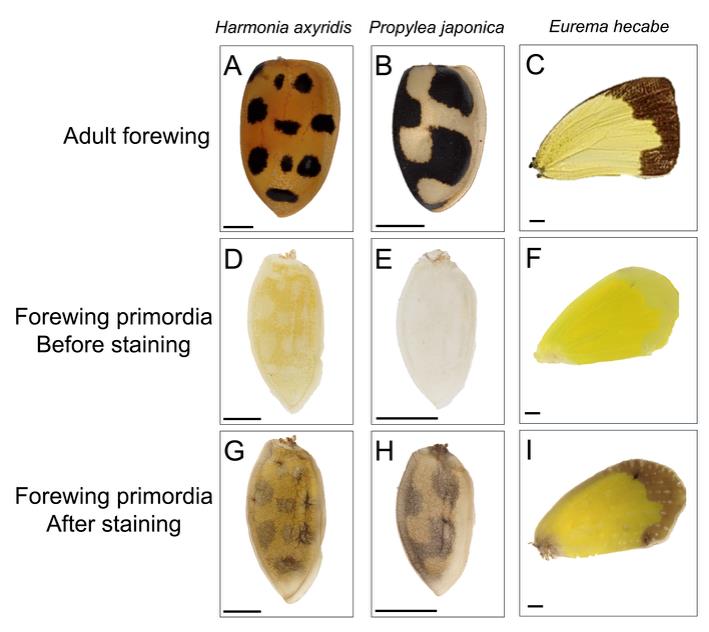
Figure 3. Phenoloxidase (PO) activity staining in pupal wing primordia of different insect species. (A–C) Adult right forewings. (D–F) Right pupal forewing primordia before staining. (G–I) Right pupal forewing primordia after staining for 120 min. Scale bars: 1 mm.
Validation of protocol
This protocol has been used and validated in the following research article:
• Ando et al. [1]. Repeated inversions within a pannier intron drive diversification of intraspecific colour patterns of ladybird beetles. Nat Commun. (Figure 2a and Supplementary Figures 1 and 9)
General notes and troubleshooting
General notes
We observed that the pupal stage, suitable for estimating the spatial distribution of PO proteins, may be associated with external morphological features. In all three species examined, wing primordia were easily dissected at approximately 80% of the total pupal duration, and staining for 120 min yielded clear patterns corresponding to adult melanin-forming regions. Based on external morphological features, H. axyridis pupae at this stage (later than 80% development) exhibit both black compound eyes and black hindwing primordia, whereas earlier stages (>60%) show dark brown eyes with no wing pigmentation (Figure 4). Similar pigmentation changes in the compound eyes and hindwings were observed in P. japonica at comparable stages (data not shown). In E. hecabe, while eye pigmentation was distinct, yellow pigmentation of the wings became apparent at this stage (data not shown). These external markers may thus serve as practical indicators for identifying the optimal timing for staining.

Figure 4. Developmental stages of H. axyridis pupae. Immediately after pupation, the pupa is entirely white; after 1 h, black melanin pigmentation appears on the dorsal side. The compound eye primordia progressively darken during development and become fully black by day 4. In addition, melanization of the hindwing primordia located beneath the forewing primordia becomes apparent by day 4. The numbers above the images indicate the percentage (%) of the total pupal duration (4.5 days), and the numbers below indicate the time after pupation for each image.
Troubleshooting
We caution that staining patterns corresponding to adult melanin-forming regions cannot be reliably obtained at earlier developmental stages. In our previous paper, in H. axyridis wing primordia at 70 h after pupation (~65% of the total pupal duration), weak staining was observed across the entire wing primordium rather than patterns corresponding to adult melanin-forming regions [1]. Therefore, we recommend performing staining at later developmental stages.
Acknowledgments
T.N. conceived and developed the protocol. Y.M., S.M., and T.N. supervised the study. Y.N. performed experiments to validate the protocol. Y.N. wrote the manuscript with feedback from all authors. All authors reviewed and approved the final version of the manuscript. We are grateful to Tatsuro Konagaya for providing E. hecabe pupae and photographs of adults, and to Toshiya Ando for helpful advice. This protocol was adapted from True et al. [11] and the master’s thesis of Kumiko Goto (2009). This work was supported by JST SPRING, Japan Grant Number JPMJSP2104 (to Y.N.), and JSPS KAKENHI grant numbers 19H01004 and 23H02227 (to T.N.). We thank the Model Plant Research Facility/NIBB BioResource Center and the Emerging Model Organisms Facility of NIBB Trans-Scale Biology Center for technical assistance. This protocol was used in [1].
Competing interests
No conflict of interest is declared regarding this article.
Ethical considerations
No ethical requirements are needed regarding this article.
References
- Ando, T., Matsuda, T., Goto, K., Hara, K., Ito, A., Hirata, J., Yatomi, J., Kajitani, R., Okuno, M., Yamaguchi, K., et al. (2018). Repeated inversions within a pannier intron drive diversification of intraspecific colour patterns of ladybird beetles. Nat Commun. 9(1): 3843. https://doi.org/10.1038/s41467-018-06116-1
- Sugumaran, M. and Barek, H. (2016). Critical analysis of the melanogenic pathway in insects and higher animals. Int J Mol Sci. 17(10): 1753. https://doi.org/10.3390/ijms17101753
- Futahashi, R., Koshikawa, S., Okude, G. and Osanai-Futahashi, M. (2022). Diversity of melanin synthesis genes in insects. In Sugumaran, M. Advances in Insect Physiology. Academic Press, 339–376. https://doi.org/10.1016/bs.aiip.2022.03.003
- Sorrentino, R. P., Small, C. N. and Govind, S. (2002). Quantitative analysis of phenol oxidase activity in insect hemolymph. Biotechniques. 32(4): 815–823. https://doi.org/10.2144/02324st08
- Tabunoki, H., Dittmer, N. T., Gorman, M. J. and Kanost, M. R. (2019). Development of a new method for collecting hemolymph and measuring phenoloxidase activity in Tribolium castaneum. BMC Res Notes. 12(1): 7. https://doi.org/10.1186/s13104-018-4041-y
- Arakane, Y., Muthukrishnan, S., Beeman, R. W., Kanost, M. R. and Kramer, K. J. (2005). Laccase 2 is the phenoloxidase gene required for beetle cuticle tanning. Proc Natl Acad Sci USA. 102(32): 11337–11342. https://doi.org/10.1073/pnas.0504982102
- Cerenius, L. and Söderhäll, K. (2004). The prophenoloxidase-activating system in invertebrates. Immunol Rev. 198(1): 116–126. https://doi.org/10.1111/j.0105-2896.2004.00116.x
- Asano, T., Taoka, M., Yamauchi, Y., Craig Everroad, R., Seto, Y., Isobe, T., Kamo, M. and Chosa, N. (2014). Re-examination of a α-chymotrypsin-solubilized laccase in the pupal cuticle of the silkworm, Bombyx mori: Insights into the regulation system for laccase activation during the ecdysis process. Insect Biochm and Mol Biol. 55: 61–69. https://doi.org/10.1016/j.ibmb.2014.10.004
- Yatsu, J. and Asano, T. (2009). Cuticle laccase of the silkworm, Bombyx mori: Purification, gene identification and presence of its inactive precursor in the cuticle. Insect Biochem and Mol Biol. 39(4): 254–262. https://doi.org/10.1016/j.ibmb.2008.12.005
- Asada, N. (1998). Reversible activation of prophenoloxidase with 2-propanol in Drosophila melanogaster. J Exp Zool. 282(1–2): 28–31. https://doi.org/10.1002/(SICI)1097-010X(199809/10)282:1/2%253C28::AID-JEZ6%253E3.0.CO;2-P
- True, J. R., Kopp, A. and Carroll, S. B. (2001). Prophenoloxidase as a reporter of gene expression in Drosophila. Biotechniques. 30(5): 1004–1009. https://doi.org/10.2144/01305st06
- Kanost, M. R. and Gorman, M. J. (2008). Phenoloxidases in insect immunity. In Beckage N. Insect Immunology. Academic Press, 69–96. https://doi.org/10.1016/B978-012373976-6.50006-9
- Binggeli, O., Neyen, C., Poidevin, M. and Lemaitre, B. (2014). Prophenoloxidase activation is required for survival to microbial infections in Drosophila. PLoS Pathog. 10(5): e1004067. https://doi.org/10.1371/journal.ppat.1004067
- Nakamura, T., Matsuoka, Y. and Niimi, T. (2024). Egg microinjection for the ladybird beetle Harmonia axyridis. Bio Protoc. 14(24): e5136. https://doi.org/10.21769/BioProtoc.5136
- Konagaya, T. and Maruyama, M. (2025). Mature egg production in cryopreserved ovaries of a pierid butterfly, Eurema mandarina. J Insect Conserv. 29(2): 29. https://doi.org/10.1007/s10841-025-00667-7
- Niimi, T., Kuwayama, H. and Yaginuma, T. (2005). Larval RNAi applied to the analysis of postembryonic development in the ladybird beetle, Harmonia axyridis. J insect biotechnol sericology. 74(3): 95–102. https://doi.org/10.11416/jibs.74.95
- Du, M.-H., Yan, Z.-W., Hao, Y.-J., Yan, Z.-T., Si, F.-L., Chen, B. and Qiao, L. (2017). Suppression of Laccase 2 severely impairs cuticle tanning and pathogen resistance during the pupal metamorphosis of Anopheles sinensis (Diptera: Culicidae). Parasit Vectors. 10(1): 171. https://doi.org/10.1186/s13071-017-2118-4
- Niu, B.-L., Shen, W.-F., Liu, Y., Weng, H.-B., He, L.-H., Mu, J.-J., Wu, Z.-L., Jiang, P., Tao, Y.-Z. and Meng, Z.-Q. (2008). Cloning and RNAi-mediated functional characterization of MaLac2 of the pine sawyer, Monochamus alternatus. Insect Mol Biol. 17(3): 303–312. https://doi.org/10.1111/j.1365-2583.2008.00803.x
- Ze, L.-J., Jin, L. and Li, G.-Q. (2023). The compatible effects of RNA interference targeting laccase2 with biocontrol in Henosepilachna vigintioctopunctata. Entomol Gen. 43(1): 117–126. https://doi.org/10.1127/entomologia/2023/1644
- Lamana, M. L. and Miller, J. C. (1998). Temperature-dependent development in an Oregon population of Harmonia axyridis (Coleoptera: Coccinellidae). Environ Entomol. 27(4): 1001–1005. https://doi.org/10.1093/ee/27.4.1001
- Zhang, X., Li, Y., Romeis, J., Yin, X., Wu, K. and Peng, Y. (2014). Use of a pollen-based diet to expose the ladybird beetle Propylea japonica to insecticidal proteins. PLoS One. 9(1): e85395. https://doi.org/10.1371/journal.pone.0085395
- Arju, M. H., Miah, M. K., Parven, N. and Bashar, M. (2015). Developmental stages of a common grass yellow butterfly, Eurema hecabe. Dhaka Univ J Biol Sci. 24(1): 73–81. https://doi.org/10.3329/dujbs.v24i1.46311
Article Information
Publication history
Received: Dec 22, 2025
Accepted: Feb 19, 2026
Available online: Mar 10, 2026
Published: Apr 5, 2026
Copyright
© 2026 The Author(s); This is an open access article under the CC BY-NC license (https://creativecommons.org/licenses/by-nc/4.0/).
How to cite
Nakatani, Y., Matsuoka, Y., Morita, S. and Niimi, T. (2026). A Simple Method for Estimating the Spatiotemporal Distribution of Phenoloxidase Proteins in Insect Tissues. Bio-protocol 16(7): e5646. DOI: 10.21769/BioProtoc.5646.
Category
Developmental Biology > Morphogenesis
Biological Sciences > Biological techniques
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










