- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
First-line Evaluation of Sperm Parameters in Mice (Mus musculus)
Published: Vol 12, Iss 20, Oct 20, 2022 DOI: 10.21769/BioProtoc.4529 Views: 3238
Reviewed by: Chiara AmbrogioAnonymous reviewer(s)
Abstract
Infertility has become a major public health problem, with a male factor involved in about half the cases. Mice are the most widely used animal model in reproductive biology research laboratories, but changes in sperm parameters in mice can be subtle and, in the absence of official guidelines, it is important that analyses are carried out in a strict and reproductive manner. This protocol successively details the different steps required to obtain spermatozoa under good conditions, the measurement of sperm motility using a Computer Assisted Sperm Analysis System (CASA) device, the calculation of sperm concentration in the epididymides using a sperm counting cell, and the examination of sperm morphology. The combination of these assays provides an overview of the basic sperm parameters in mice. This is both a diagnostic and a decision-making tool for researchers to orient their scientific strategy according to the observed abnormalities.
Keywords: SpermBackground
Mice are the most widely used model organism in laboratories around the world because of their relatively simple breeding conditions, short reproduction cycles, and physiology close to humans. Several fields of science use mice as model organisms, and the advent of genetically manipulated mouse breeding has further expanded the possibilities of this model (Olson et al., 2018; Ito et al., 2018; Gill et al., 2020). For example, CRISPR technology now allows the efficient generation of mouse knock-out models to functionally validate newly identified genetic mutations in a wide range of diseases (Abbasi et al., 2017; Hochheiser et al., 2018; Zhao et al., 2020), including infertility (Beurois et al., 2020; Touré et al., 2021).
Medically assisted reproduction techniques are implemented in the murine model (Stein et al., 2010), establishing it as a model of choice for reproductive biologists (Giritharan et al., 2010), for whom the analysis of sperm parameters became essential a long time ago (Osterloh et al., 1983; Tessler et al., 1985). These techniques have since been adopted by a large portion of scientists who use them for different purposes, such as the evaluation of the impact of diseases and/or toxic substances on reproduction (Sampannang et al., 2020; Abdollahi et al., 2021) and the curative and/or protective effects of molecules against induced deleterious effects (Feyli et al., 2017; Moghadam et al., 2021; Shahedi et al., 2021). Other noteworthy examples include the analysis of nutrition influence on sperm parameters (Zhao et al., 2017; Gómez-Elías et al., 2019) and their heritability (Crisóstomo et al., 2021), or the potential of cryobiology for reproduction (Sztein et al., 2018).
However, this model suffers from limitations in its transfer to humans. One of the drawbacks of the mouse model is that mice have a much more efficient spermatogenesis and much higher fertility than humans—already the primate with the worst known sperm parameters (Martinez and Garcia, 2020). This implies that even subtle defects in mouse sperm parameters may eventually translate into a much more deleterious impact in humans (Martinez et al., 2022). That is why descriptions of animal reproductive phenotypes are becoming increasingly sophisticated and standardized, with each specialty creating its own phenotyping framework (Houston et al., 2021). Here, we propose a rapid, accessible, and reproducible protocol for the first-line assessment of sperm parameters in mice, which provides the researcher with a first diagnosis to guide their scientific strategy and the framework of their future experiments.
This protocol sequentially describes the surgical steps of sperm recovery, sperm motility measurement using a Computer Assisted Sperm Analysis System (CASA) device, sperm concentration measurement, and the examination of sperm morphology.
Materials and Reagents
Petri dish polystyrene (Falcon, catalog number: 351008)
Box with instruments for minor surgery (LD Medical, catalog number: 01748)
Needles, MicrolanceTM 3, 18 G (Beckton Dickinson, catalog number: 304622)
Microtube safety lock 2 mL Clearline (Dutscher, catalog number: 2519649)
SureOne specialty tips (Fisherbrand, catalog number: 02-707-464)
Counting chamber Leja 100 μm (Leja Products B.V., Gynemed, catalog number: SC100-01-02-A)
Thoma Cell Counting chamber (VWR, catalog number: HECH40447702)
Tubes 15 mL (Falcon, catalog number: 352096)
SuperFrost PlusTM adhesion slides (Epredia, catalog number: J7800AMNZ)
Soda lime glass staining troughs with lid (Brand, catalog number: 472200)
EUKITT® classic 100 mL flask (Dutscher, catalog number: 124103)
Mice (Charles River Laboratories)
M2 medium (Sigma-Aldrich, catalog number: M7167-50ML)
Ethanol 96% (Carlo Erba, Labelians, catalog number: CE529141)
Sterile water (Versylene, Fresenius)
Dulbecco′s phosphate buffered saline (PBS) (Sigma-Aldrich, catalog number: D8537-500ML)
Papanicolaou′s solution 1a Harris′ hematoxylin solution (Sigma-Aldrich, catalog number: 1092530500)
Shorr staining solution stain for histology (Sigma-Aldrich, catalog number: 1092750500)
Ammonia solution 25% (Sigma-Aldrich, catalog number: 105432)
Immersion oil for microscopy type N (Nikon, MXA22166)
75% ethanol (see Recipes)
50% ethanol (see Recipes)
Ethanol ammoniacal (see Recipes)
Equipment
Surgical scissors
Incubator binder model CB56 (Binder, catalog number: 9640-0006)
Binocular loupe (Nikon, catalog number: SM2800N)
IKA dry block heater 3 (IKA, catalog number: 0004025300)
Computer Assisted Sperm Analysis System (CASA) CEROS II (Hamilton Thorne Research, IMV Technologies, catalog number: 024905) with its heated blade holder
Centrifuge Sorvall ST 16R (ThermoFisher, catalog number: 10688725)
Nikon Eclipse 80i microscope equipped with a Nikon DS-Ri1 camera with NIS-Elements software and 40× and 100× objectives
Procedure
Notes:
The first two steps of this protocol (sperm recovery and motility analysis) are performed under time and temperature constraints. Sperm motility is a physiological phenomenon that evolves over time and is impacted by the temperature of the environment. To obtain reproducible motility data, the time between sperm extraction and motility measurement has to be the same for all individuals in the study (optimal time would be between 15 and 20 min), and a temperature of 37 °C has to be maintained throughout. As concentration and morphology measurements are not impacted by these factors, they can then be carried out comfortably afterwards, without restrictions.
When handling sperm, always use a cut or wide-opening pipette tip (see Figure 1). The use of normal tips induces deleterious effects on the motility and morphology of spermatozoa.
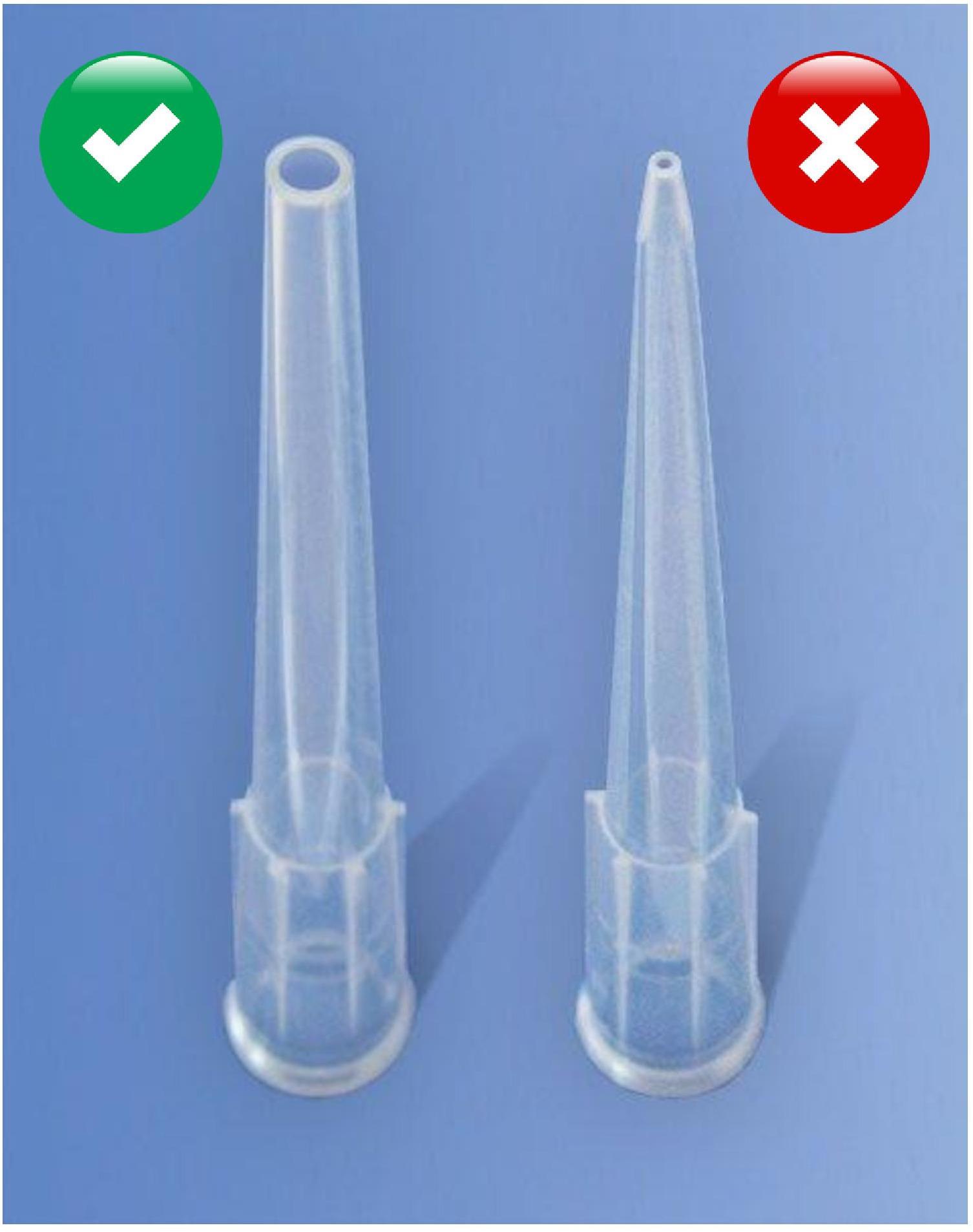
Figure 1. Use wide-opening pipette tips
Before beginning the procedure
Prepare Petri dishes (two per individual, one per epididymis) with 1 mL of M2 medium and one tube with 5 mL of M2 medium. Heat them to 37 °C for at least 2 h before starting the experiment.
Prepare the dissection field.
Preheat the heating block to 37 °C and place 2 mL microtubes in it (see Figure 2).
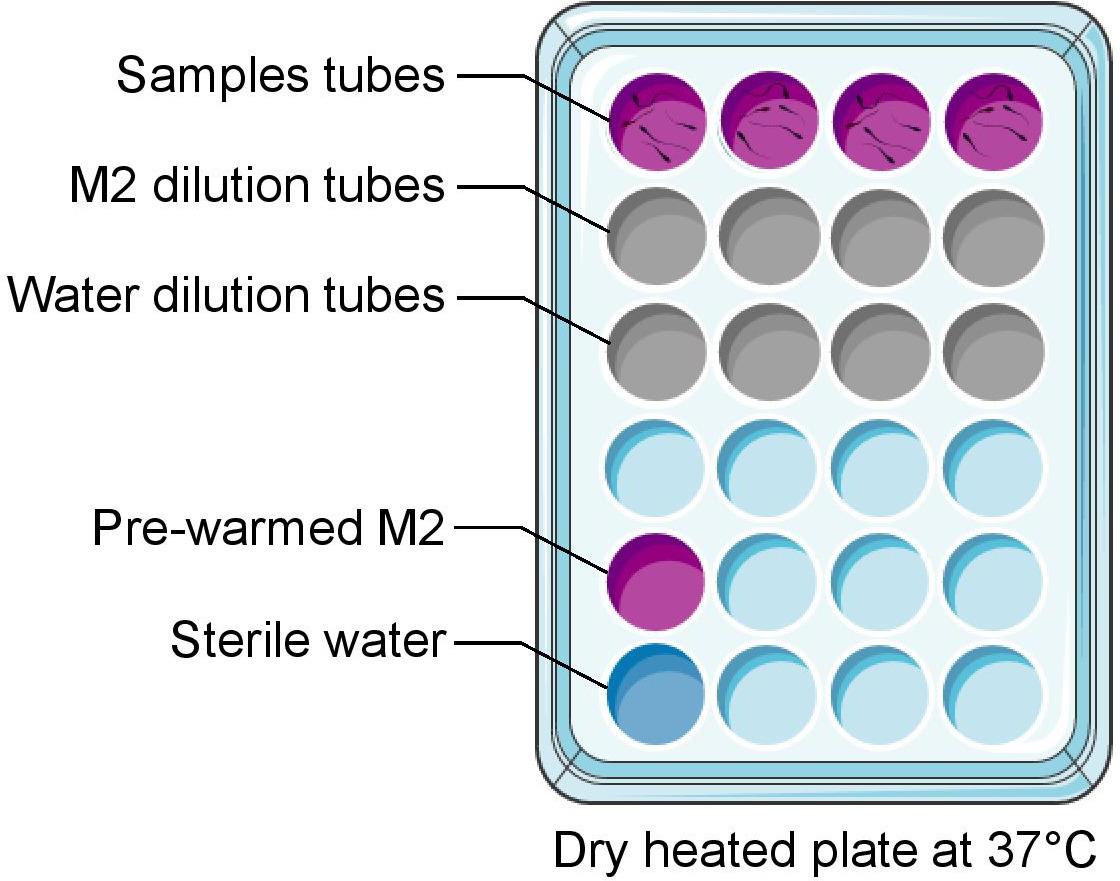
Figure 2. Suggested organization of the heating blockConnect the heated blade holder of the CASA.
Turn on the computer, start the CASA software (Figure 3), and enter the sample numbers if appropriate.
Note: The sperm analysis software must have the animal analysis module and be configured according to its user manual. We provide the parameters to be used for mice sperm analysis in Table 1, but we will not go over all the basic configuration elements. For this, please refer to the CEROS II manual.
Table 1. Settings for CASA analysis
Parameters Value Parameters Value acquisition rate 60 Hz low static–intensity gate 0.5 number of frames 45 high static–intensity gate 1.3 minimum contrast 50 minimum elongation gate 0 minimum cell size 5 maximum elongation gate 87 low static-size gate 0.3 magnification factor 0.7 high static-size gate 1.95 motile sperm VAP >1 progressive motility VAP >30 + STR >70 
Figure 3. Screenshot of the software window
Sperm recovery
Sacrifice the animal according to the procedures established in your country and validated by your local ethics committees.
Note: To obtain reproducible and exploitable data, all animals in the same study should be sacrificed at the same age (we recommend between 10 and 12 weeks for an evaluation of the reproductive function in the mature male).
Immediately put the animal on its back on a sterile surgical field and disinfect the abdominal and genital area with ethanol.
Note: Simply spray or apply ethanol with a sterile compress.
Use surgical scissors to make a horizontal skin incision from one lumbar region to the other (Figure 4A), and a large vertical incision through the hypogastric region to the pelvis region (Figure 4B).
Expose the organs by incising vertically through the abdominal muscle wall (Figure 4C).
Note: A quick check will allow you to spot unexpected flagrant anomalies such as a situs invertus.
Locate (Figure 4D) and pull upward with surgical clamps the fat deposits located on each side of the pelvic fossae to elevate and expose the testicles (Figure 4E).
Note: Be careful not to puncture the bladder.
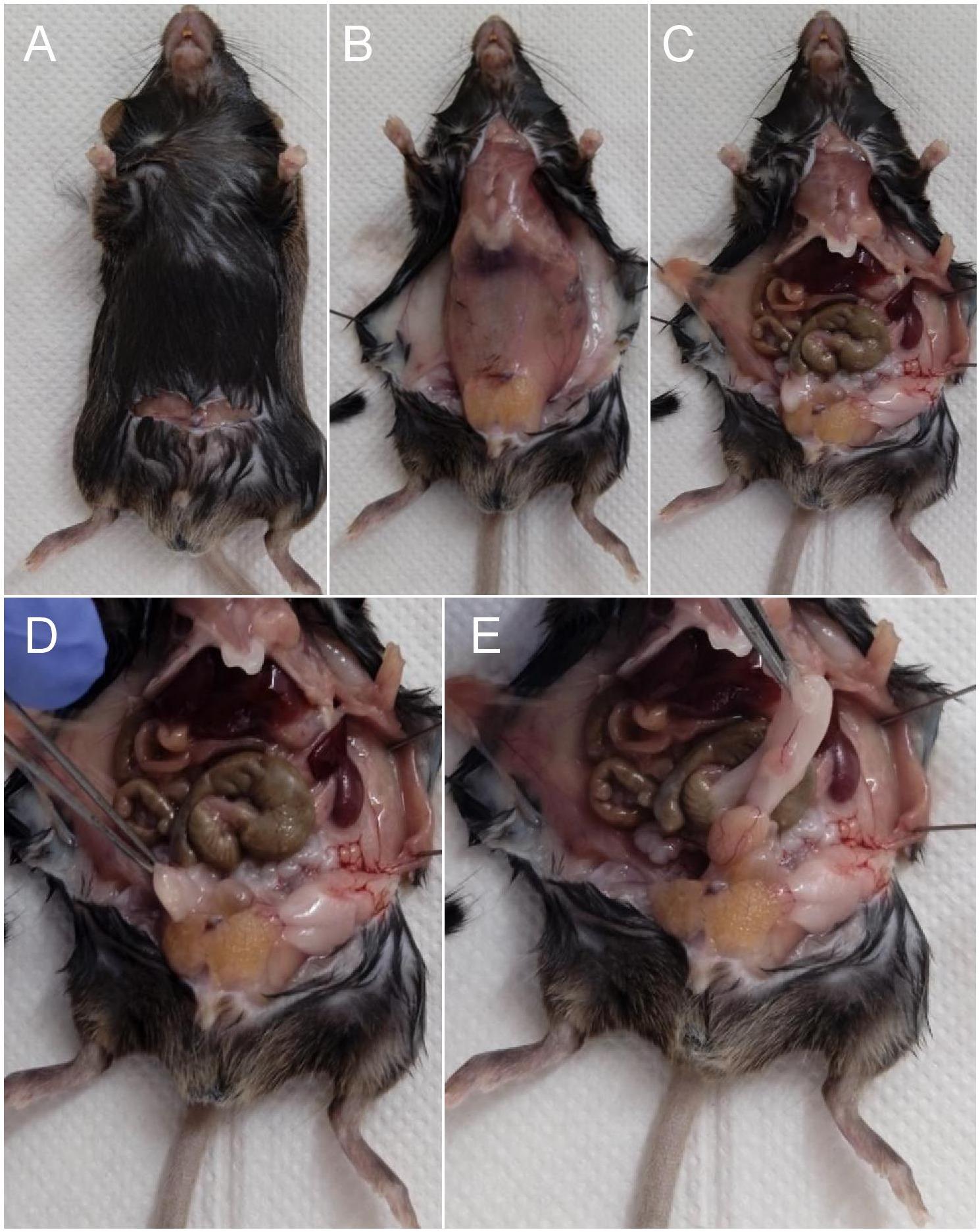
Figure 4. Sequence of surgical procedures for exposure of reproductive organsLocate the different structures of the reproductive tract from the testis (Figure 5A). Start with the head of the epididymis visible on the testicle and then proceed to the body, tail, and then the efferent canal.
Isolate the tail of the epididymis by incising at one millimeter past the tail-canal junction distally (Figure 5B) and at the body-tail junction proximally (Figure 5C), and place it immediately in the pre-warmed M2 medium in Petri dish.
Note: Optimal time for steps 1–7 is less than 10 min.
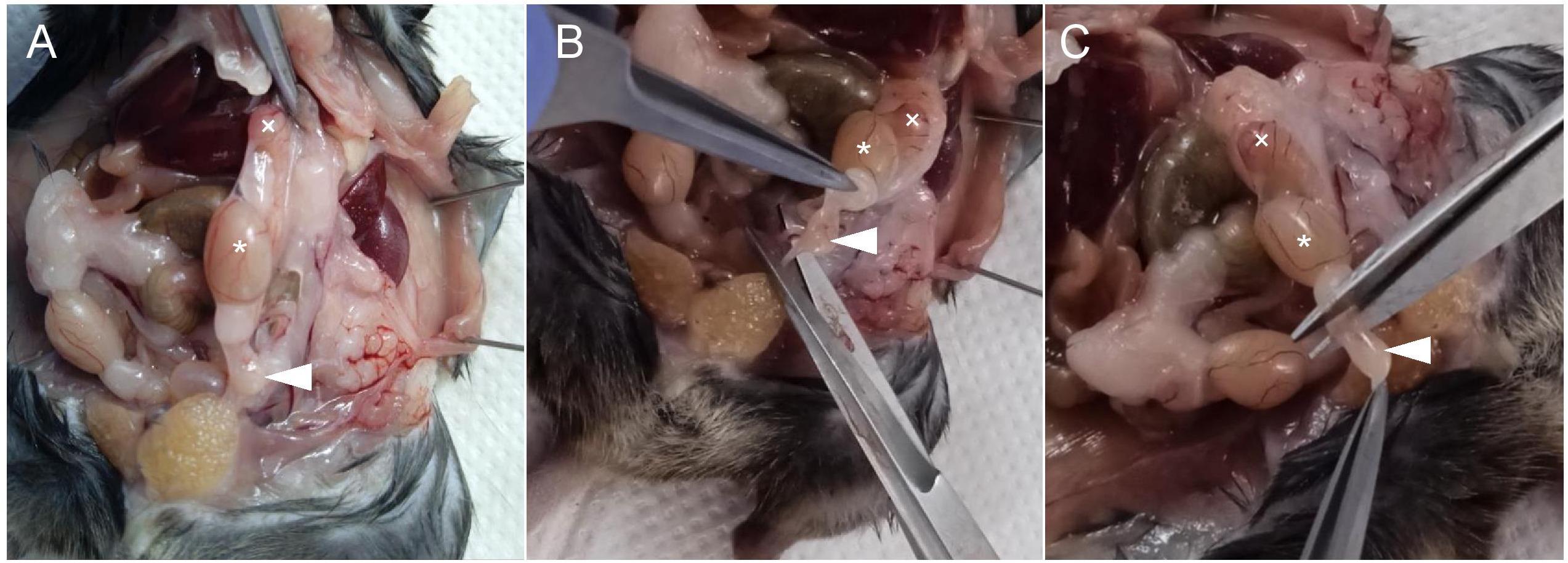
Figure 5. Sequence of surgical procedures for epididymal tail isolation with indication of testis (asterisk), epididymal head (cross), and tail (arrow)Dissect the epididymis with two needles under a binocular loupe with a preheated plate at 37 °C (Figure 6).
Note: It is important to completely drain the tail of the epididymis to obtain the correct concentration of spermatozoa. The dissection must be done very quickly, which can be challenging for a beginner. In this case, always take the same amount of time to dilate the epididymes of the different individuals of the same study. Optimal time for step 8 is less than 10 min (for both epididymis).
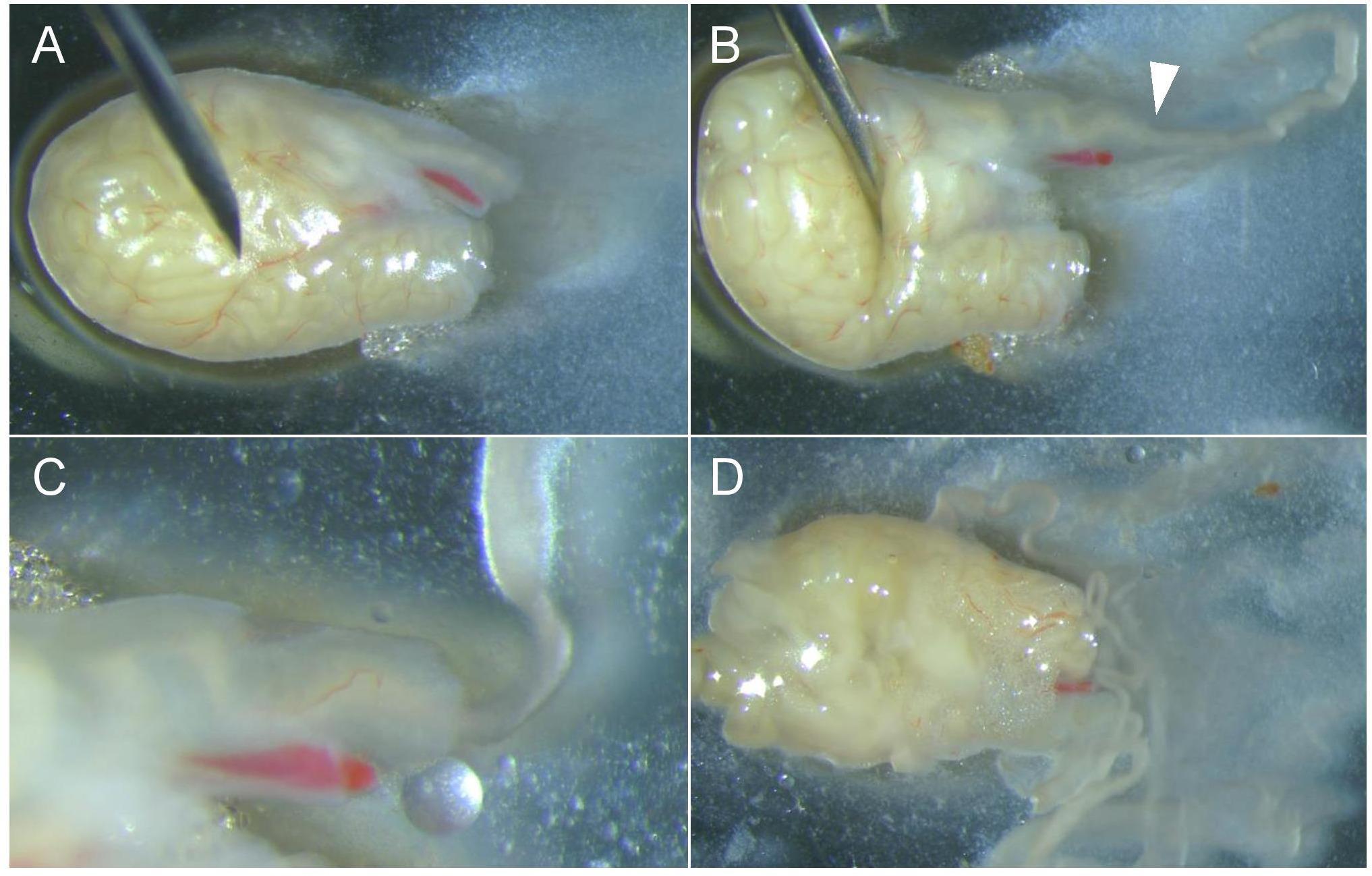
Figure 6. Place the tail of the epididymis so that the incisions are clearly visible. (A) Apply pressure to the tail of the epididymis with a needle to remove the sperm through the vas deferens. (B) The spermatozoa will come out as a jelly that will disintegrate in the medium. (C) Empty the epididymis completely by applying repeated pressure. (D) It may be necessary to pierce the organ in several places to empty it completely.Place the Petri dishes in an incubator at 37 °C for 10 min to allow the sperm to gain motility.
Note: These ten minutes can be used to adjust the CASA settings and fill in the sample identification in the software, if not already done beforehand.
Sperm motility assessment
Transfer the sperm in M2 medium from the Petri dish to the 2 mL microtube preheated in a 37 °C heating block.
Note: You can pool the two epididymides of the same individual or treat them separately according to your needs.
Take 20 µL of suspended semen, place it in a second Eppendorf, then dilute it with preheated M2 medium.
Note: The dilution performed depends on the mice strain used. Perform an experiment beforehand on a control individual to identify the dilution to be used for the rest of the study. For example, we use a standard 1:50 dilution for our Black 6 mice but a 1:100 dilution for our OF1 mice.
Homogenize and then transfer the sample to a Leja slide. Put the slide in the heated blade holder under the microscope.
Note: Wait a few moments for the flow to stabilize in the chamber to prevent the sperm trajectories from being affected by any residual Brownian motion.
Confirm that the sperm density is in the acceptable range of the software and check that the sample identification is correct.
Note: The software acceptability cut-off is relatively high. We advise to use a concentration that does not lead to a systematic crossing of the trajectories to allow an easier analysis afterwards.
Acquire with the software as much field as you may need.
Note: For a statistically robust analysis, we recommend acquiring a minimum of 200 motile spermatozoa. Use a “forward walk” reading during the acquisition to avoid taking the same field several times (Figure 7).
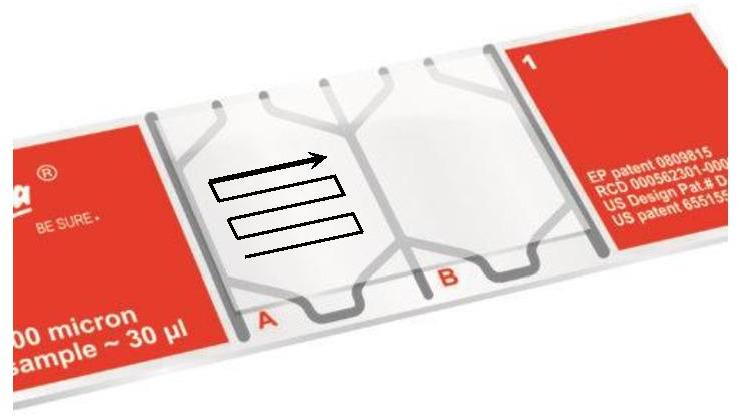

Figure 7. “Forward walk” reading path on a Leja slideProcess all samples successively in the same way.
Note: The purpose here is to acquire the data (the videos) to being able to interpret them later. Do not waste time interpreting now and focus on acquiring all the samples sequentially.
Sperm concentration assessment
Note: We describe here the method used for a Thoma chamber; however, you can use any other type of counting chamber (e.g., Mallassez or Makler) as long as you follow the associated instructions.
Take 10 µL of suspended semen, place it in a third Eppendorf, and add 90 µL of sterile H2O to immobilize the spermatozoa.
Note: You can use another dilution, but you will have to take it into account when calculating the final concentration.
Homogenize and transfer 10 µL to the Thoma counting chamber.
Count the number of sperm present on two lines to obtain the sperm concentration expressed in millions per milliliter (Figure 8).
Note: Count the sperm heads; single flagella are not counted or counted separately. This is also an opportunity to calculate the concentration of any round cells in the ejaculate (germline or immune cells). If you have pooled epididymides, do not forget to multiply by two. You can count more lines and average the results for more accuracy.
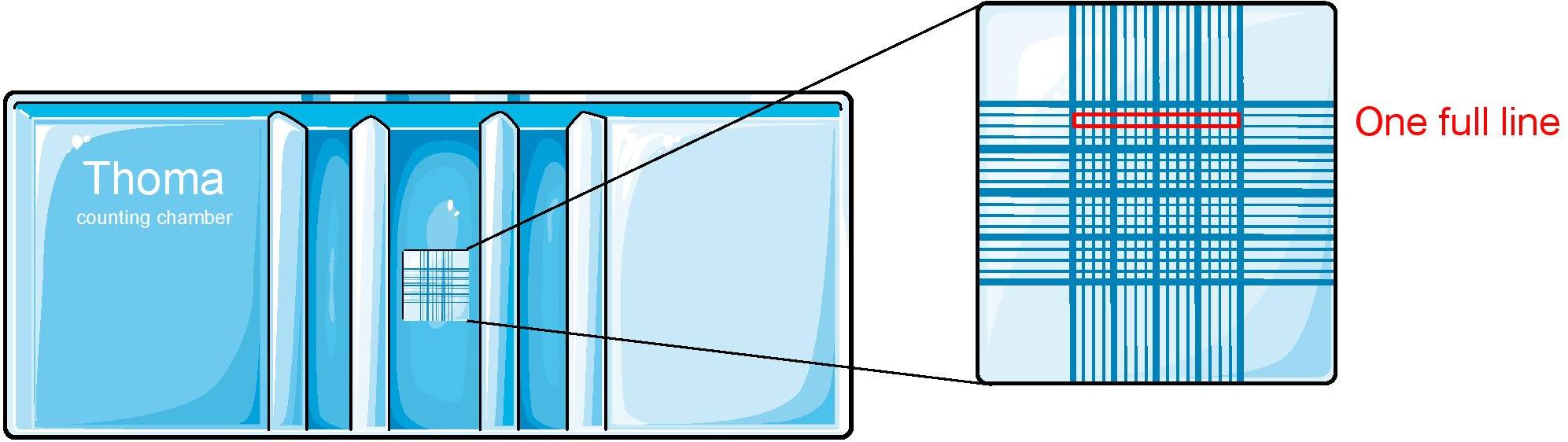
Figure 8. Count two full lines of a 1:10 diluted sample to obtain the cell concentration in millions per milliliter
Sperm morphology assessment
Transfer the remaining samples to 15 mL tubes. Add 3 mL of 1× PBS. Homogenize by gentle pipetting.
Centrifuge at 400 × g for 5 min. Discard the supernatant.
Add another 3 mL of 1× PBS to the pellet.
Homogenize by gentle pipetting.
Centrifuge at 400 × g for 5 min. Discard the supernatant.
Add another 200 µL of 1× PBS to the pellet.
Homogenize by gentle pipetting.
Put 10 µL (this volume can be adjusted according to the desired concentration of cells on the slide) at the base of a clean slide and make a smear with a second slide.
Note: The drop must be "pulled" and not "pushed" by the second blade, which would break the sperm flagella (see Figure 9).
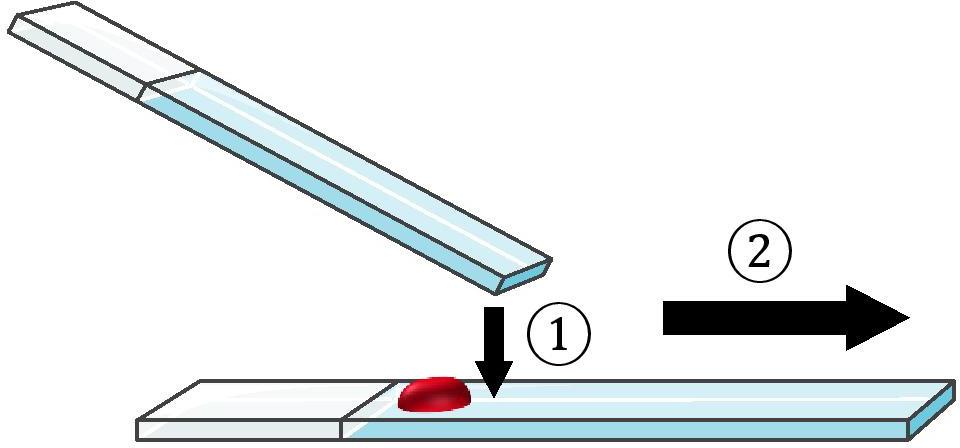
Figure 9. The smearing blade should be placed in front of the drop and pull it forward. It must not push or cut through the drop.Let dry at room temperature.
Immerse in a 75% ethanol bath for at least 1 h.
Let dry at room temperature.
Perform a Harris-Schorr staining by carrying out the following sequence of baths (one immersion corresponds to a duration of approximately 1 s):
Tap water—15 immersions
Hematoxylin—2 min
Tap water—15 immersions
Ethanol ammoniacal—10 immersions
Tap water—15 immersions
Ethanol 50%—5 min
Shorr—5 min
Ethanol 50%—5 min
Ethanol 75%—5 min
Ethanol 95%—5 min
Allow the stained slides to dry completely.
Note: Any alcohol residue will interfere with the interpretation.
Mount the slides with Eukitt and a 24 mm × 60 mm cover slide.
Note: Avoid trapping air bubbles and remove them by applying gentle pressure to the cover slide if necessary.
Let the mounted smear dry horizontally on absorbent paper for 24 h before interpretation.
Data analysis
Concentration analysis
Data can be presented as cell concentration in million cells per milliliter, number of cells per epididymis, or total numeration of cells per individual.
Motility analysis
Review each acquired video (Figure 10) to ensure that the selected sperm paths are correct, i.e., eliminate possible experiment artifacts (air bubble, residue, aggregates) and intersecting sperm paths.
Note: For more details, please refer to the CEROS II user manual.
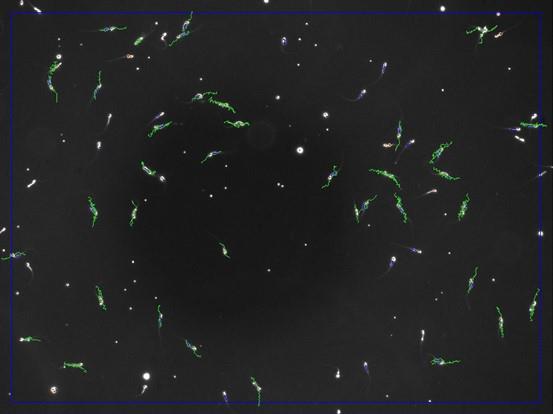
Figure 10. Screenshot of pattern analysis from the software. Motile sperm pathways are highlighted in green.The software will provide you with all the information about sperm motility (Figure 11): a first part with the percentages of motile and progressive motile sperm, a second part (if necessary) with the main parameters—the curvilinear velocity (VCL) or average path velocity (VAP), straight-line velocity (VSL), and amplitude lateral of the head (ALH)—and then if necessary the subsequent findings (e.g., the percentage of capacitated sperm).
Note: Be careful with how you handle the results given by the software, as they are in the form of mean/median ± standard error/deviation. Depending on the analysis you want to do, it could be wiser, statistically speaking, to start directly from the raw data of each trajectory extractable from the software.
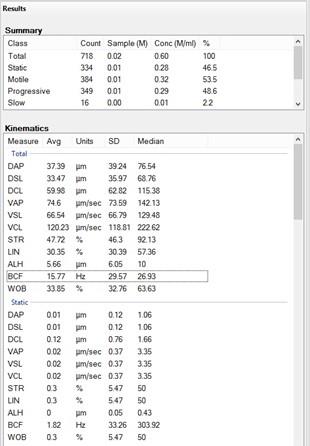
Figure 11. Screenshot of results provided by the software post-analysis
Morphology analysis
Analyze the slides using a brightfield light microscope with the 100× objective and immersion oil.
The head is stained pale blue in the acrosome region and dark blue in the post-acrosomal region. The intermediate piece is stained red and the flagellum blue or red. The cytoplasmic residues are colored red–orange.
Differentiate spermatozoa according to their typical/atypical morphology (Figure 12) on a minimum of 100 cells (200 for more precise analysis).
Note: It is possible to limit the analysis to the typical/atypical character of the spermatozoa, but it is interesting to carry out an in-depth analysis by separating the abnormalities of the head and the flagellum, or even going into the details of the abnormalities. For this purpose, it is necessary to distinguish abnormalities of the head [elongated, thinned, microcephalic, macrocephalic, multiple, with an abnormal or absent acrosome, or with an abnormal base (post-acrosomal region], the intermediate piece (which may be small, angular, or with cytoplasmic remains), and the flagellum (which may be absent, shortened, of irregular caliber, coiled, and/or multiple).
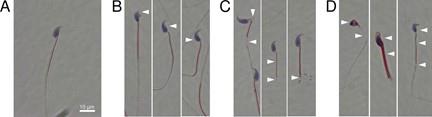
Figure 12. Light microscopy pictures of Harris-Schorr stained spermatozoa (from Martinez et al., 2022). (A) Typical morphology. (B) Head abnormalities. (C) Flagellum abnormalities. (D) Head and flagellum abnormalities. Defects are pointed by white arrows.
Notes
In an effort to refine and reduce the number of animals used in the studies, always consider additional analyses whenever possible: fix the testicles to make histological sections or the remaining spermatozoa pellets to make immunofluorescence slides, freeze for proteomics, extract DNA from other organs, and run toxicology analyses. Whatever your field of study, try to plan for simultaneous manipulations.
Recipes
75% ethanol
100 mL of 96% ethanol + 31 mL of sterile water
50% ethanol
100 mL of 96% ethanol + 98 mL of sterile water
Ethanol ammoniacal
95 mL of 75% ethanol + 5 mL of ammonia solution at 25%
Acknowledgments
This protocol is derived from the original research paper “Oligogenic heterozygous inheritance of sperm abnormalities in mouse” from Martinez et al. published in eLife on April 22, 2022 (Martinez et al., 2022; doi:10.7554/eLife.75373) and funded by Agence Nationale de la Recherche ANR-19-CE17-0014 and ANR-21-CE17-0007.
Competing interests
The authors declare that no competing interests exist.
Ethics
All animal procedures presented in the original paper were conducted according to a protocol approved by the local Ethics Committee (ComEth Grenoble No. 318), by the French government (ministry agreement number #7128 UHTA-U1209-CA), and by the Direction Générale de la Santé (DGS) for the State of Geneva.
References
- Abbasi, F., Miyata, H. and Ikawa, M. (2018). Revolutionizing male fertility factor research in mice by using the genome editing tool CRISPR/Cas9. Reprod Med Biol 17(1): 3-10.
- Abdollahi, M. B., Dehghan, S. F., Balochkhaneh, F. A., Moghadam, M. A. and Mohammadi, H. (2021). Comparison of mice' sperm parameters exposed to some hazardous physical agents. Environ Anal Health Toxicol 36(3): e2021013-2021010.
- Beurois, J., Cazin, C., Kherraf, Z. E., Martinez, G., Celse, T., Toure, A., Arnoult, C., Ray, P. F. and Coutton, C. (2020). Genetics of teratozoospermia: Back to the head. Best Pract Res Clin Endocrinol Metab 34(6): 101473.
- Crisóstomo, L., Jarak, I., Rato, L. P., Raposo, J. F., Batterham, R. L., Oliveira, P. F. and Alves, M. G. (2021). Inheritable testicular metabolic memory of high-fat diet causes transgenerational sperm defects in mice. Sci Rep 11(1): 9444.
- Feyli, S. A., Ghanbari, A. and Keshtmand, Z. (2017). Therapeutic effect of pentoxifylline on reproductive parameters in diabetic male mice. Andrologia 49(1). doi: 10.1111/and.12604.
- Gill, K. K., Rajan, J. R. S., Goldowitz, D. and Zwicker, J. G. (2020). Using a mouse model to gain insights into developmental coordination disorder. Genes Brain Behav 19(4): e12647.
- Giritharan, G., Li, M. W., Di Sebastiano, F., Esteban, F. J., Horcajadas, J. A., Lloyd, K. C., Donjacour, A., Maltepe, E. and Rinaudo, P. F. (2010). Effect of ICSI on gene expression and development of mouse preimplantation embryos. Hum Reprod 25(12): 3012-3024.
- Gómez-Elías, M. D., Rainero Caceres, T. S., Giaccagli, M. M., Guazzone, V. A., Dalton, G. N., De Siervi, A., Cuasnicu, P. S., Cohen, D. J. and Da Ros, V. G. (2019). Association between high-fat diet feeding and male fertility in high reproductive performance mice. Sci Rep 9(1): 18546.
- Hochheiser, K., Kueh, A. J., Gebhardt, T. and Herold, M. J. (2018). CRISPR/Cas9: A tool for immunological research. Eur J Immunol 48(4): 576-583.
- Houston, B. J., Conrad, D. F. and O'Bryan, M. K. (2021). A framework for high-resolution phenotyping of candidate male infertility mutants: from human to mouse. Hum Genet 140(1): 155-182.
- Martinez, G. and Garcia, C. (2020). Sexual selection and sperm diversity in primates. Mol Cell Endocrinol 518: 110974.
- Martinez, G., Coutton, C., Loeuillet, C., Cazin, C., Muronova, J., Boguenet, M., Lambert, E., Dhellemmes, M., Chevalier, G., Hograindleur, J. P., et al. (2022). Oligogenic heterozygous inheritance of sperm abnormalities in mouse. Elife 11: e75373.
- Moghadam, M. T., Dadfar, R. and Khorsandi, L. (2021). The effects of ozone and melatonin on busulfan-induced testicular damage in mice. JBRA Assist Reprod 25(2): 176-184.
- Ito, R., Takahashi, T. and Ito, M. (2018). Humanized mouse models: Application to human diseases. J Cell Physiol 233(5): 3723-3728.
- Olson, B., Li, Y., Lin, Y., Liu, E. T. and Patnaik, A. (2018). Mouse Models for Cancer Immunotherapy Research. Cancer Discov 8(11): 1358-1365.
- Osterloh, J., Letz, G., Pond, S. and Becker, C. (1983). An assessment of the potential testicular toxicity of 10 pesticides using the mouse-sperm morphology assay. Mutat Res 116(3-4): 407-415.
- Sampannang, A., Arun, S., Burawat, J., Sukhorum, W. and Iamsaard, S. (2020). Comparison of male reproductive parameters in mice with type 1 and type 2 diabetes. Clin Exp Reprod Med 47(1): 20-33.
- Shahedi, A., Talebi, A. R., Mirjalili, A. and Pourentezari, M. (2021). Protective effects of curcumin on chromatin quality, sperm parameters, and apoptosis following testicular torsion-detorsion in mice. Clin Exp Reprod Med 48(1): 27-33.
- Stein, P. and Schultz, R. M. (2010). ICSI in the mouse. Methods Enzymol 476: 251-262.
- Sztein, J. M., Takeo, T. and Nakagata, N. (2018). History of cryobiology, with special emphasis in evolution of mouse sperm cryopreservation. Cryobiology 82: 57-63.
- Tessler, S. and Olds-Clarke, P. (1985). Linear and nonlinear mouse sperm motility patterns. A quantitative classification. J Androl 6(1): 35-44.
- Touré, A., Martinez, G., Kherraf, Z. E., Cazin, C., Beurois, J., Arnoult, C., Ray, P. F. and Coutton, C. (2021). The genetic architecture of morphological abnormalities of the sperm tail. Hum Genet 140(1): 21-42.
- Zhao, J. L., Zhao, Y. Y. and Zhu, W. J. (2017). A high-fat, high-protein diet attenuates the negative impact of casein-induced chronic inflammation on testicular steroidogenesis and sperm parameters in adult mice. Gen Comp Endocrinol 252: 48-59.
- Zhao, Y., Qu, H., Wang, Y., Xiao, W., Zhang, Y. and Shi, D. (2020). Small rodent models of atherosclerosis. Biomed Pharmacother 129: 110426.
Article Information
Copyright
Martinez. This article is distributed under the terms of the Creative Commons Attribution License (CC BY 4.0).
How to cite
Readers should cite both the Bio-protocol article and the original research article where this protocol was used:
- Martinez, G. (2022). First-line Evaluation of Sperm Parameters in Mice (Mus musculus). Bio-protocol 12(20): e4529. DOI: 10.21769/BioProtoc.4529.
- Martinez, G., Coutton, C., Loeuillet, C., Cazin, C., Muronova, J., Boguenet, M., Lambert, E., Dhellemmes, M., Chevalier, G., Hograindleur, J. P., et al. (2022). Oligogenic heterozygous inheritance of sperm abnormalities in mouse. Elife 11: e75373.
Category
Developmental Biology > Reproduction
Biological Sciences > Biological techniques
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link









