- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
High Throughput Blood-brain Barrier Organoid Generation and Assessment of Receptor-Mediated Antibody Transcytosis
(*contributed equally to this work) Published: Vol 12, Iss 8, Apr 20, 2022 DOI: 10.21769/BioProtoc.4399 Views: 4641
Reviewed by: Pilar Villacampa AlcubierreWilliam C. W. ChenAnand Ramesh Patwardhan

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
Accurate Identification of Cell Cycle Stages in RPE1 Cells Using the ImmunoCellCycle-ID Method
Syon Reddy [...] Aussie Suzuki
Aug 5, 2025 1908 Views
Isolation and Imaging of Microvessels From Brain Tissue
Josephine K. Buff [...] Sophia M. Shi
Aug 5, 2025 2700 Views
High Content In Vitro Survival Assay of Cortical Neurons
Paolo V. Fioretti [...] Manuela Basso
Feb 5, 2026 240 Views
Abstract
Targeting receptor-mediated transcytosis (RMT) is a successful strategy for drug delivery of biologic agents across the blood-brain barrier (BBB). The recent development of human BBB organoid models is a major advancement to help characterize the mechanisms of RMT and thus accelerate the design of brain delivery technologies. BBB organoids exhibit self-organization, which resembles the architecture of the neurovascular unit, and low paracellular permeability, due to the formation of tight junctions between endothelial cells. However, current methods of organoid generation have low throughput, exhibit substantial heterogeneity across experiments, and require extensive manual handling. These limitations prevent the use of BBB organoids as a screening tool for discovery and optimization of therapeutic molecules. In this protocol, we use hydrogel-based arrays to generate human BBB organoids, with a 35-fold increase in organoid yield as compared to previous protocols using 96-well plates. We incubate BBB organoid arrays with monoclonal antibody-based constructs and use a custom semi-automated imaging assay to assess RMT within the organoid core. The experimental and analytical tools described in this protocol provide a scalable platform that can be incorporated in the early stages of drug discovery to accelerate the development and optimization of brain delivery technologies to cross the BBB.
Keywords: Blood-brain barrierBackground
The blood-brain barrier (BBB) is a highly selective biological interface composed of brain endothelial cells, pericytes, and astrocytes, that tightly regulates the transport of molecules from the blood into the brain parenchyma. Brain delivery of therapeutic proteins is hindered by their low rate of transport across the BBB. A clinically-validated approach to increase transport across the BBB is to target endogenous receptors expressed on brain microvascular endothelial cells with protein shuttles. The development of such protein shuttle platforms requires scalable in vitro systems that allow high-throughput screening for both compound optimization and systematic discovery of transport mechanisms.
Previously described in vitro systems to investigate transcytosis across the BBB include transwell models, microfluidic systems, or organoids. Previous work showed that organoids are superior to transwell models for recapitulating angiopep-2 transcytosis (Cho et al., 2017). The direct cell-cell interaction in BBB organoid arrays also increases the expression of receptors involved in transcytosis compared to cells in transwells (Urich et al., 2013). On the other hand, microfluidic systems recapitulate complex vascular networks and can generate quantitative permeability data (Hajal et al., 2022). However, this type of models is not yet scalable for high-throughput screening.
Different groups described the generation of BBB organoids by the self-assembly of human brain endothelial cells, astrocytes, and pericytes (Urich et al., 2013; Cho et al., 2017; Nzou et al., 2018). Importantly, BBB organoids recapitulated key functional properties of the BBB, including tight junction formation, expression of efflux transporters, and low permeability to macromolecules. However, previous protocols require extensive manual handling, and exhibit high inter-experimental variability that prevent their widespread adoption for high-throughput screening. This protocol solves these limitations by: 1) increasing organoid generation yield by 35-fold, 2) enabling the growth of homogeneous organoids in a 96-well format, 3) developing a simple microscopy-based assay to assess transport across the BBB, and 4) automating image analysis. The focus of this protocol is on evaluating receptor-mediated transcytosis of antibodies. However, it can be modified to evaluate transport of other biologic agents, including peptides, nanoparticles, and viruses. Since the main readout is microscopy-based, users can combine it with viability probes to evaluate acute cytotoxic effects in endothelial cells, astrocytes, or pericytes. Overall, these characteristics make BBB organoid arrays a flexible assay platform for screening applications in a drug discovery setting.
Materials and Reagents
50 mL polypropylene conical tube (Falcon, catalog number: 352070)
Tissue culture flask, 250 mL, 75 cm2 (Falcon, catalog number: 353136)
GRI3D 96-well plate (VWR, catalog number: MSPPGRI3D-96P-S, order plates with a well diameter of 600 µm), store at 4°C
Eppendorf tubes 5.0 mL with snap cap, 5.0 mL, sterile, colorless (Eppendorf, catalog number: 0030119487)
Eppendorf Safe-Lock tubes, 1.5 mL (Eppendorf, catalog number: 0030121589)
Eppendorf Protein LoBind Tube 1.5 mL (Eppendorf, catalog number: 022431081)
Countess cell counting chamber slide (Invitrogen, catalog number: C10228)
Adhesive microscope slides for tissue sections, ground edges, twin frosted end (Marienfeld Superior, catalog number: 0800001)
High Precision Microscope cover glasses 22 × 22 mm No 1.5H (Marienfeld Superior, catalog number: 0107052)
Blood-brain barrier hCMEC/D3 endothelial cell line (Millipore Sigma, catalog number: SCC066)
Human Astrocytes (ScienCell, catalog number: 1800)
Human Brain Vascular Pericytes (ScienCell, catalog number: 1200)
Endothelial Media: EGM-2 Endothelial Cell Growth Medium-2 BulletKit (Lonza, catalog number: CC-3156), store at 4°C, do not use for longer than 1 month
Astrocyte Media (ScienCell, catalog number: 1801), store at 4°C, do not use for longer than 1 month
Pericyte Media (ScienCell, catalog number: 1201), store at 4°C, do not use for longer than 1 month
Organoid Media: EGM-2 Endothelial Cell Growth Medium-2 BulletKit (Lonza, catalog number: CC-3156) without VEGF, store at 4°C, do not use for longer than 1 month
Trypsin 0.05% (Gibco, catalog number: 25300-054), store at 4°C after initial thaw from -20°C
Dulbecco’s Phosphate Buffered Saline (DPBS) (Gibco, catalog number: 14190-094), store at room temperature
Poly-L-Lysine (PLL) solution, mol wt 150,000–300,000, 0.01% sterile filtered (Sigma, catalog number: P4832-50ml), store at 4°C
Ultrapure Distilled Water DNase/RNase Free (UPW; Invitrogen, catalog number: 10977-035), store at room temperature
Eppendorf Protein LoBind 0.5 mL (Eppendorf, catalog number: 022431064)
Trypan blue stain 0.4% (Invitrogen, catalog number: T10282), store at room temperature
Brain Shuttle – custom-made anti-transferrin receptor antibody (Sade et al., 2013). Store at -80°C. This molecule can be replaced by any other anti-transferrin receptor antibody, for example MEM-189 (Abcam, catalog number: ab1086).
CellEventTM Caspase-3/7 Green Detection Reagent (Thermo Fisher Scientific, catalog number: C10423), store at -20°C
16% Paraformaldehyde Aqueous Solution EM Grade (PFA) (Electron Microscopy Sciences, catalog number: 50-980-487), store at room temperature
Triton X-100 aqueous solution 10% (w/v) (Roche, catalog number: 11332481001), store at 4°C
Donkey Serum sterile filtered (Canvax Biotech, catalog number: SUD004), store at -20°C
Human IgG (H+L) AlexaFluor 488-coupled goat AffiniPure Fab Fragment (Jackson Immunoresearch, catalog number: 109-547-003), resuspend with glycerol and dH2O as per the supplier’s instructions, and store at -20°C.
DAPI (Sigma, catalog number: D9542-5mg), store at 4°C
Disposable wipes KIMTECH® Science precision wipes, 7552 (Carl Roth, catalog number: AA64.1)
Fluoromount-G (Electron Microscopy Sciences, catalog number: 17984-25)
Leica Microsystems Type F Immersion Oil for Microscopes (Fisher Scientific, catalog number: 11944399)
Poly-L-Lysine solution (see Recipes)
4% PFA solution (see Recipes)
Permeabilization Buffer (see Recipes)
Dilution Buffer (see Recipes)
Wash Buffer (see Recipes)
Equipment
Thermo Scientific Precision Water Bath GP 15-D 5L and 10L (Dual) (Thermo Scientific, catalog number: TSGP15D)
CountessTM II FL automated cell counter (Thermo Fisher Scientific, Invitrogen, catalog number: AMQAF1000)
Centrifuge 5804R (Eppendorf, catalog number: 5804000520)
Zeiss Axio Observer 3 LED or DMi8 Leica widefield microscope
VWR Rotator Multimix 230V Swis (VWR Collection, catalog number: 444-0501)
Curved and sharp tweezers (VWR, catalog number: 232-0078)
Scanning laser confocal microscope (Leica, model: SP5)
Software
ImageJ/Fiji software versions 1.53n (available for download: https://imagej.net/)
Procedure
Thawing of cells
Prepare human astrocyte media, human pericyte media, and endothelial media, by adding all supplements provided by the supplier to the basal media bottles.
Pre-warm all media at room temperature prior to use.
Pre-coat two T75-flasks with PLL solution (see Table 1) for subcutlturing of the human pericyte and human astrocyte cell lines.
Incubate flasks at room temperature for at least 1 h.
After incubation, aspirate the PLL solution, add 15 mL of the appropriate media in each flask, and place them in the incubator (37°C, 5% CO2) to equilibrate the media.
In parallel, add 15 mL of endothelial media in a T75 flask, and place in the incubator.
While the media is equilibrating, retrieve the vials of cells (Astrocytes, Pericytes and hCMEC/D3) from liquid nitrogen.
Thaw vials using a water bath set at 37°C, until only a small amount of ice is left.
Pipette the cell suspension from the cryovial into the T75 flask with the appropriate media. Flush the vial with 1 mL of fresh media, to make sure all cells are transferred to the T75 flask.
Change the media of all flasks the next day.
Note: We do not recommend warming up the media in the water bath, but instead allow components to reach room temperature, and then equilibrate them in the incubator, as described here. Always record the lot number of each cell line used. Cells thawed on a Thursday will be 90% confluent, and ready to use for subsequent steps by the next Monday.
Preparation of 96-well GRI3D Plate
Prepare organoid media by adding all components of the EGM-2 Endothelial Cell Growth Medium-2 BulletKit provided by the supplier to the basal media bottle, except VEGF. Let it reach room temperature prior to use.
In a sterile environment, remove the 96-well GRI3D plate from its cover and open the lid.
Using a pipette tip, gently remove the film protection that is found over the plate.
Aspirate the liquid by placing an aspirator in the reservoir of each well. Once the liquid is removed from the reservoir, place the aspirator inside the well, and remove excess liquid found above the microwells. Be careful not to touch the microwells, as the aspirator will damage them (Figure 1A – side view schematic).
Once the liquid is removed from all wells, add 200 μL of organoid media in each well, and place the plate in the incubator until the organoid cell suspension is ready (Step D).
Note: For aspirating the liquid from the top of the microwells, place a P20 tip on the aspirator to ensure that the hydrogel is not damaged.
Trypsinizing Astrocytes, Pericytes, and Endothelial cells
Allow the 0.05% trypsin solution to reach room temperature prior to use.
Prepare three 50-mL Falcon tubes with 10 mL of pre-warmed organoid media. Label tubes as either astrocyte, pericyte, or hCMEC/D3 cells.
For each cell line, take its flask out of the incubator, aspirate the media, wash with 10 mL of DPBS, remove DPBS, and add 5 mL of 0.05% Trypsin. Place flasks back in incubator for 3–5 min.
While flasks are trypsinizing, prepare three 0.5-mL tubes with 10 μL of trypan blue stain, and label three cell counting chamber slides with either astrocyte, pericyte, or hCMEC/D3 cells.
For the hCMEC/D3 cells, check that the cells in the flask have been trypsinized, and add 5 mL of organoid media in the flask to neutralize the 5 mL of trypsin. Pipette a few times to break cell clumps.
Pipette 10 mL of hCMEC/D3 cell suspension (5 mL trypsin + 5 mL of organoid media) from the T75 flask to the corresponding labeled Falcon tube. The final volume in the tube should be 20 mL.
Mix the cell suspension adequately to ensure that there are no cell clumps and that the cells are equally distributed throughout the entire volume.
Pipette 10 μL from the hCMEC/D3 cell suspension and mix with 10 μL of trypan blue stain in the 0.5-mL tube. Pipette 10 μL of the trypan blue stained cell suspension to each side of the labelled cell counting chamber slide.
Repeat steps 5–8 for the astrocytes and pericytes.
Centrifuge the three Falcon tubes at 180 × g for 5 min.
Calculation of cell concentrations, and preparation of organoid cell suspension
Place the cell counting chamber slide into the cell counter.
Select the appropriate cell counting profile, and press Count.
Record the number of live cells per mililiter, and percentage of live cells, for each side of the slide. Calculate the average number of live cells. We expect 3–5 million astrocytes, 5–7 million pericytes, and 8–12 million hCMEC/D3 cells per confluent T75 flask.
Calculate the media needed to resuspend each cell pellet to a final concentration of 2.75 × 106 cells/mL (check Note 1 at the end of this section for calculation explanation).
Remove Falcon tubes from centrifuge. There should be a visible cell pellet at the bottom of each tube.
Aspirate the organoid media from each Falcon tube without perturbing the cell pellet at the bottom of each tube. It is important to remove as much of the media as possible, to ensure accurate cell numbers.
Resuspend each cell pellet with the appropriate amount of organoid media, as calculated in step 4. Pipette up and down, to ensure that the cells distribute equally throughout the media volume.
In a 5-mL tube, mix equal volumes of astrocyte, pericyte, and hCMEC/D3 cell suspensions. This represents the organoid cell suspension, which is composed of all three cell types at a ratio of 1:1:1.
Notes:
For each well, use a final volume of 60 μL of organoid cell suspension added on top of the microwells. Since the organoid cell suspension is composed of all three cell types at a ratio of 1:1:1, use 20 μL of each cell type suspension per well. We established the optimal cell number and ratios for organoid growth to be 1 × 103 cells of each cell type per organoid. Since each well of the GRI3D plate contains 55 microwells, this requires 5.5 × 103 cells for each cell line, within a 20 μL volume. This gives the desired concentration of 2.75 × 106 cells/mL used in step 4.
It is important that the cell counts are as accurate as possible. We observed that if there are less than 3 × 103 cells per organoid, multiple smaller structures are formed instead of one large structure. For the first experiments, it is advisable to calibrate the cell counts given by the automated cell counter manually.
A 96-well GRI3D plate contains 60 wells with microwells. To use all the wells of the plate, an organoid cell suspension of at least 60 wells × 60 μL = 3.6 × 103 μL is required, which corresponds to mixing 1.2 × 103 μL of each cell line. Always prepare a volume larger than needed.
Organoid formation
Once the organoid cell suspension is ready, remove the 96-well GRI3D plate from the incubator, and aspirate the organoid media from each well. Try to remove as much media as possible, without disturbing the microwells. Use a P20 pipette tip to do so (read Note at the end of this section).
Mix the organoid cell suspension, to ensure the cells are evenly mixed.
Add 60 μL of organoid cell suspension on top of the microwells of each well (not in the reservoir).
Check the wells under the microscope. You should see the cells floating above the microwells (Figure 1B).
Place the plate into the incubator for at least 20 min. Avoid sudden movement of the plate, to prevent that the 60-μL organoid cell suspension leaks into the reservoir.
After 20–30 min, check the cells under the microscope, to ensure that they have fallen into the microwells (Figure 1B).
Gently, add 200 μL of organoid media into the reservoir of each well, ensuring not to flush out the cells from the microwells.
After 24 h, check the organoid formation under the microscope. The cells should have already self-assembled into organoids (Figure 1B). If cells have not aggregated (e.g., due to low viability of the starting cell suspension, bacterial contamination, or cytotoxicity), the organoid formation needs to be repeated.
Aspirate the organoid media from the reservoir, and add 200 μL of fresh pre-warmed organoid media. There is no need to aspirate the media from the top of the microwells as well.
Place the organoids back in the incubator for 24 h. At 48 h, the organoids are ready for the experiment (Figure 1B).
Note: If organoid media is not aspirated fully, then there is a chance that the 60 μL will leak into the reservoir, and there will be less cells per organoid. Try to remove as much of the media as possible, prior to adding the cells.
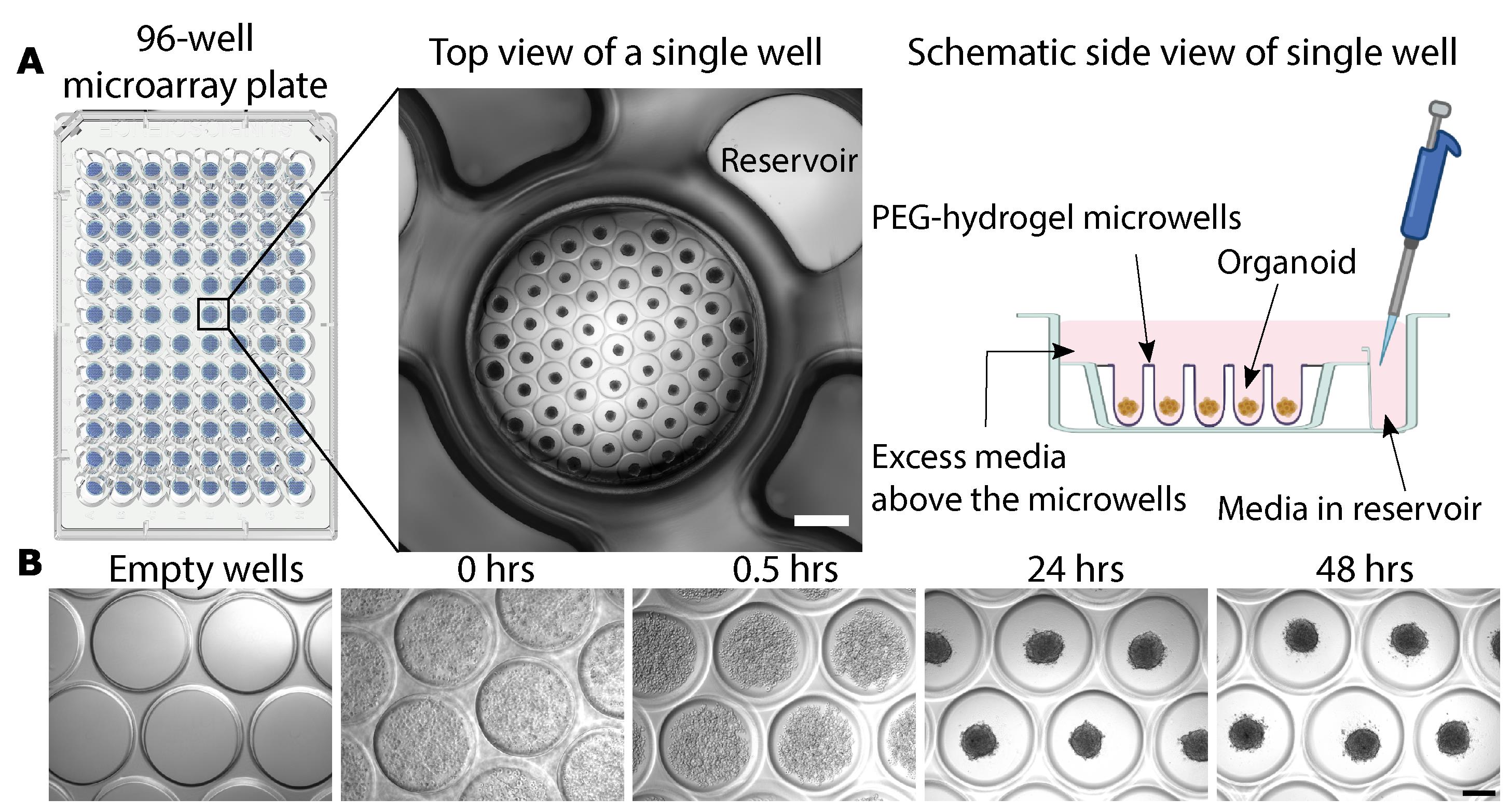
Figure 1. Blood-brain barrier organoid formation using the 96-well SunBioscience GRI3D microarray plate. A. Schematic of a 96-well GRI3D microarray plate and a representative phase contrast image of a single well with its microarray and associated reservoir. Scale bar, 1 mm. Modified from Simonneau et al. (2021). A side view schematic (not to scale) highlights the presence of media both in the reservoir, as well as above the PEG-hydrogel microwells. B. Phase contrast images of the microarray during organoid formation. At 0 h, the cells are added on top of the empty wells and are evenly distributed throughout the media. At 0.5 h, the cells have fallen into the microwells and cluster closer together, leading to the formation of a self-assembled organoid within 24 h. At 48 h, the organoids are ready for treatment. Scale bar, 200 μm.Assessment of organoid viability using CellEvent Caspase-3/7 dye
Note: After 48 h, assess the viability of the organoids. To do so, we treat one or two wells with CellEvent Caspase-3/7 dye. It is important to check organoid viability prior to each experiment, especially when using new cell lot numbers.
After 48 h, prepare 200 μL of 2 μM of CellEvent Caspase-3/7 dye per well (dilution of 1:1,000) in organoid media.
Remove the media from the reservoir of the well, and add the freshly prepared media containing the CellEvent Caspase-3/7 dye.
Place the plate back in the incubator for at least 30 min.
After 30 min, assess the signal of CellEvent Caspase-3/7 dye using a DMi8 Leica widefield with filter set for GFP/FITC/AlexaFluor 488. If there is no bright core, the organoids are viable, and can be used for further experiments (Figure 2).
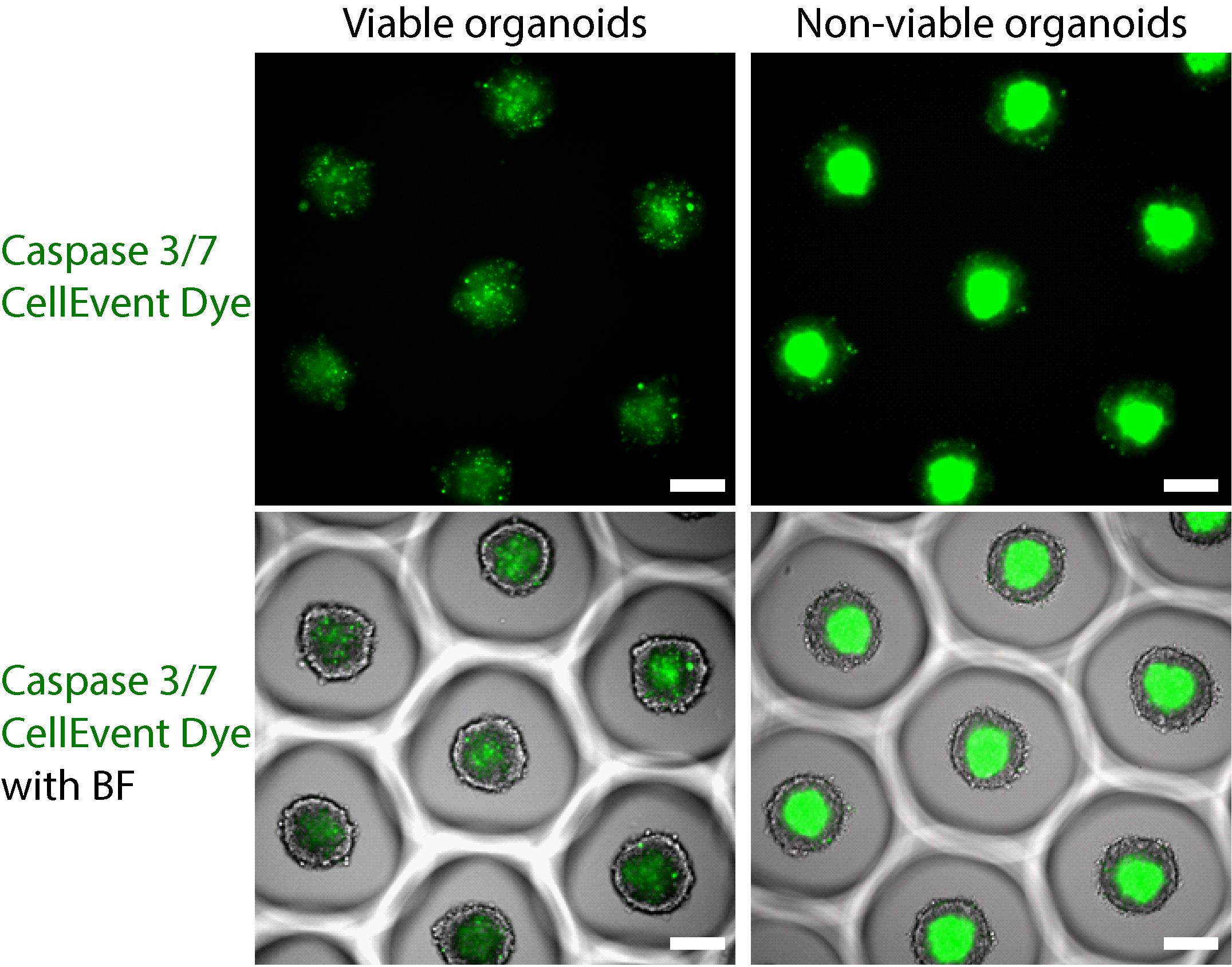
Figure 2. Assessment of blood-brain barrier organoid viability using Caspase 3/7 CellEvent Dye at 48 h. The first row shows the signal obtained from Caspase 3/7 CellEvent dye, and the second row shows the caspase signal merged with a brightfield image of the organoids. The first column shows the signal of Caspase 3/7 CellEvent dye in viable organoids. The signal is weak and shows puncta for individual apoptotic cells. The second column shows a bright apoptotic core in non-viable organoids. The intensity of Caspase 3/7 CellEvent dye images has been adjusted to the same dynamic range. Scale bars, 150 µm.Transcytosis Assay with BBB organoid arrays
Prepare four tubes of the following solutions (calculate for a volume of 150 μL per well and always prepare a total volume in excess of what is need per condition):
Tube 1: non-treated sample containing only organoid media.
Tube 2: Mouse Brain Shuttle treatment containing organoid media with 100 nM of custom-made humanized monovalent antibody targeting the mouse transferrin receptor.
Tube 3: Human Brain Shuttle treatment containing organoid media with 100 nM of custom-made humanized monovalent antibody targeting the human transferrin receptor.
Tube 4: IgG control containing organoid media with 100 nM of non-targeting human IgG.
Aspirate organoid media from the reservoir of wells. Wait a few minutes, so that excess media on top of the microwells leaks into the reservoir. Aspirate media from the reservoir again.
In each well, add 150 μL of the corresponding treatment in the reservoir.
Incubate the plate at 37°C, 5% CO2 for 4 h.
During the incubation step, prepare 4% paraformaldehyde (PFA) solution (see Table 2).
After 4 h, aspirate the treatment solutions from the reservoir of each well, and wash with 150 μL of organoid media. Incubate organoids for 5 min, aspirate media, and wash again. Repeat six times.
After the final wash, aspirate media, and add 200 μL of 4% PFA solution per well.
After incubating with 4% PFA solution for 45 min, remove the 4% PFA solution, and add 200 μL of DPBS per well.
After a few minutes, remove the DPBS from the reservoir, and add 200 μL of fresh DPBS.
Store plate overnight at 4°C prior to staining.
Immunostaining of BBB organoids, to visualize the Brain Shuttle constructs
Prepare permeabilization buffer (PB), dilution buffer (DB), and wash buffer (WB), as described in the Recipe section.
Aspirate DPBS from the well reservoir.
Add 150 μL of PB per well, and incubate at room temperature for 1 h.
After the incubation, aspirate PB from the reservoir, and add 150 μL of WB. Wash one more time with WB, to remove any traces of PB. Leave the last wash in the plate until the next step.
Flush organoids from the plate into 1.5-mL Protein LoBind tubes. To flush organoids, pipette 150 μL of WB directly on top of the microwells. Repeat a few times, to get all organoids out of the microwells. Pipette the solution and transfer it into the Protein LoBind tube. Remember to collect any solution that has leaked from the top of the microwells into the reservoir, as it will also contain organoids. You can pool organoids from wells with the same treatment into the same tube.
Repeat step 5, by adding 150 μL of WB into the reservoir.
Once the organoids have been transferred to the 1.5-mL tubes, allow 2–3 min for them to settle to the bottom (Figure 3).
While the organoids are settling, dilute the secondary antibody solution (Goat anti-Human AF488 1:250), and DAPI (1:5,000, stock: 5 mg) in DB. Prepare 150 μL of solution per tube.
Using a pipette tip connected to an aspirator, remove the WB from the Protein LoBind tubes containing the organoids, leaving 50 μL at the bottom of the tubes. Do not touch the pipette tip to the bottom of the tube, to avoid aspirating the organoids.
Add 150 μL of antibody solution per tube.
Incubate on a rotary shaker in the dark at room temperature for 1 h.
After incubation, allow the organoids to settle to the bottom of the tube.
Add 1 mL of WB, and invert the tubes a couple of times to resuspend the organoids.
Allow the organoids to settle to the bottom, and remove the WB.
Repeat steps 13 and 14 thrice.
Note: The organoids are visible by eye, so it should be easy to see whether you have flushed out all the organoids successfully. Try to avoid the formation of bubbles. If needed, repeat the process more than once. Once the organoids are in the Eppendorf tube, it should be easy for the user to identify them in suspension or/and at the bottom of the tube (Figure 3).
Mounting of organoids on imaging slides
Aspirate the WB, but leave 50 μL of solution in the tube.
Set a 20-µL pipette to 20 µL, and pipette the solution up and down, to resuspend the organoids into the 50 μL solution.
Take 20 μL of solution, and allow the organoids to settle at the bottom of the pipette tip.
Transfer the organoids onto a microscope slide and reduce the amount of liquid added on the slide as much as possible.
You can repeat steps 3 and 4 if you wish, to have more organoids on the slide.
Carefully remove excess WB using a kimwipe tissue, while avoiding touching the organoids (Figure 3).
Add 40 μL of mounting media on top of the organoids, and, using tweezers, gently add a cover glass on top.
Allow the slides to dry at room temperature overnight.
Image the next day, or store at 4°C.
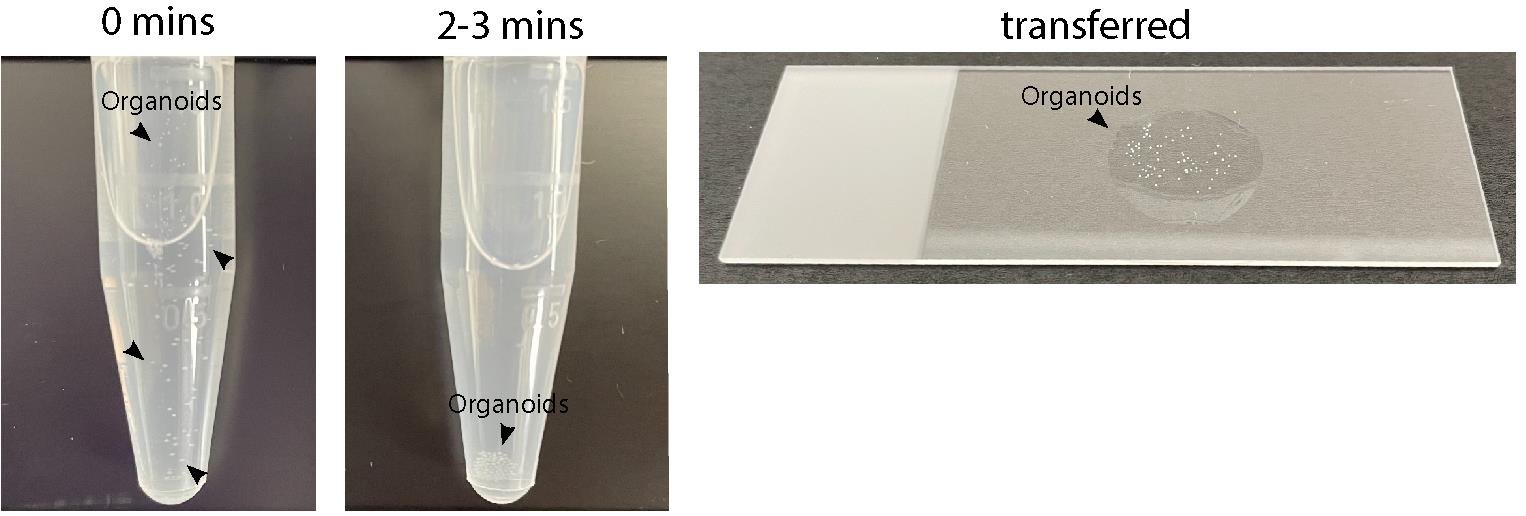

Figure 3. Blood-brain barrier organoids in suspension and transferred onto a slide. The image on the left shows visible BBB organoids in suspension directly transferred to a 1.5-mL tube. The central image shows BBB organoids settled at the bottom of the tube after 2–3 min. The image on the right shows visible BBB organoids transferred onto a microscopy slide.Confocal imaging of organoids
Image organoids using a laser scanning confocal microscope (e.g., Leica SP5), with an HCX PL APO CS 40×/1.3 oil objective, and zoom = 1.26. This results in a pixel size of 604.5 nm, for a 512 × 512 frame.
Locate an organoid, and set up the acquisition settings accordingly. At least two channels need to be set up—one for DAPI (UV laser), and one that matches the fluorophore of the secondary antibody used, in this case AlexaFluor 488 (488 laser). On a Leica SP5, adjust the range of the spectral detectors, so that there is no overlap between the two fluorophores.
Set the pinhole to achieve an optical section thickness of 1 µm.
Focus on the organoid’s core, and check that the laser power and gain maximize the 12-bit dynamic range without pixel saturation.
Set up a z-stack covering a total depth of 3.5 μm, using 8 steps with the core placed at the center (0.5 µm step size).
Image at least 20 organoids per condition per experiment. All images are stored within one .lif file, with individual names that differentiate the conditions.
Note: On the Leica SP5 microscope, the user can choose between 8-, 12-, and 16-bit images. Choose 12-bit images for all image acquisitions.
Data analysis
The code used for analysis is available here: https://github.com/phagozyt/Fiji/blob/fb365d7c1275a6b013dc32b982504dfef9653d42/MIP75ROI.
Briefly, the user provides the location of the input directory where the .lif file is stored, and the location of the output directory where the results should be stored (Figure 3). The user then chooses which channel will be used as a mask (DAPI), and which channel will be used for analysis (488 channel). This is based on the order of acquisition, for example, if DAPI was acquired as the second channel, choose Ch2. After receiving user input, the macro opens an individual z-stack stored within the .lif file. It converts the multi-channel z-stack into a multi-channel maximum projection image (Figure 4). It then splits the multi-channel maximum projection image into individual channel images, and takes the DAPI maximum projection image to create a mask via thresholding (Figure 4). The macro then converts the mask into a region of interest (ROI) based on its size and shape. Organoids that are too small or are found at the edges of the image, do not meet the ROI criteria, and are excluded. The ROIs are then reduced to 75%, to cover only the core of the organoid, and exclude measurements from the endothelial surface of the organoid. The shrunk ROIs are overlaid on top of the channel of interest (488 channel), and relevant measurements are calculated (Figure 4). The script then moves on to the next image stored within the .lif file. The macro stores an image that shows the mask generated using the DAPI images with the 75% ROI, an overlay of the maximum projection of all channels, and an image of the maximum projection of the channel of interest with the 75% ROI. After all images are processed, a .csv file is generated that contains the measurements of all objects identified within the images. We report fluorescence intensity per µm2, by dividing raw integrated density over area (µm2) (gray column in Figure 4).
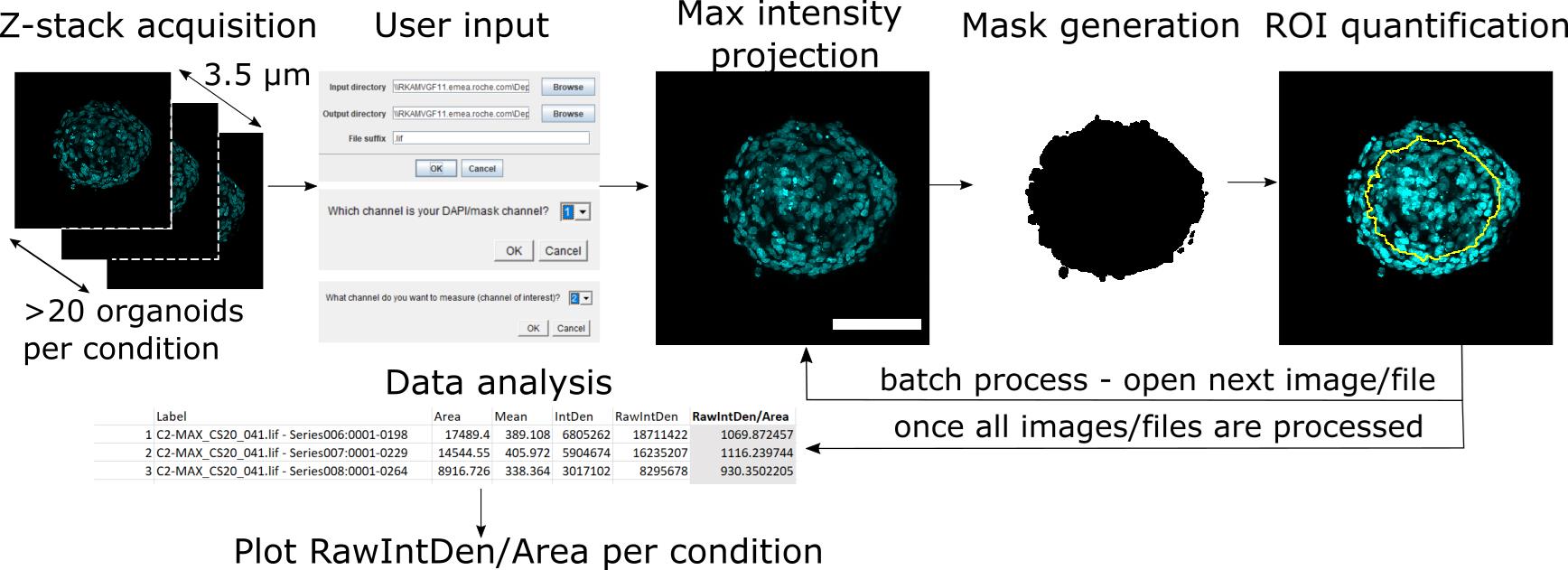

Figure 4. Standardized workflow for imaging acquisition and analysis for assessing transport into blood-brain barrier organoids. A z-stack of 3.5 µm around the core of the organoid is acquired. To initiate analysis, the user defines the input and output directory, the mask channel (DAPI), and the channel of interest (488). The z-stack is converted to a maximum intensity projection, and segmented based on DAPI, to generate the mask. The mask is then shrunk to 75% to define the ROI, which is used to measure the channel of interest. The process is repeated automatically for the next image. Once all images are processed, the user opens the saved .csv file, calculates RawIntDen/Area (gray column), and plots the results as fluorescence intensity per µm2. In all images, nuclei are labelled with DAPI (cyan). Scale bar, 100 µm.
Prior to starting the analysis for the first time, the user needs to set the measurements in Fiji. To do so, the user can go to Analyze → Set Measurements… and select Area, Mean, Integrated density, and Display labels. These are the minimum selections required for our analysis, but the user can choose more if they wish to do so.
The user may need to adjust the macro in the following ways:
The macro is set to open .lif files, which are a Leica proprietary file. If the images are obtained with a different microscope, the user has to modify the code accordingly.
The threshold used is “Triangle dark” (line 138). If the user sees that this threshold does not perform well on their images, other options can be explored. Open a representative DAPI maximum projection image in Fiji, go to Image → Adjust → Auto Threshold and, in the method, choose Try all. This will open an image with 20 different thresholded results. The user can choose which one performs the best, and modify the code accordingly.
The user may choose to work with a different type of projection. This can be modified at line 105.
If the organoids are of a different size, the user can adjust this at line 148.
If the user wishes to look at a different size of ROI, they can modify line 165.
Notes
We only use primary cells between P0 (thawing) to P3, and hCMEC/D3 cells from P4 to P15. We make fresh media every time we thaw a new vial, and we do not use any media for more than a month.
Cell source is very important, as we have found that different lot numbers from the same product can behave differently. Prior to experiments, and especially if using new lot numbers, the user needs to assess each cell source via immunostaining of common markers, such as GFAP for astrocytes, NG2 or CD13 for pericytes, and CD31 for hCMEC/D3 cells. In addition, whenever new lot numbers are used, users should perform a viability assay to ensure that there is no apoptotic core within the organoids.
Recipes
Poly-L-Lysine solution (Table 1)
Table 1. Poly-L-Lysine solution (10 mL per one T75 flask)
Reagent Amount for 10 mL Supplier Product No. Final Conc. UPW 9.850 mL Invitrogen C10283 Poly-L-Lysine 150 μL Sigma P4832-50ml 2 μg/cm2 4% PFA solution (Table 2)
Table 2. 4% Paraformaldehyde solution
Reagent Amount Supplier Product No. Final Conc. 16% Paraformaldehyde 10 mL Electron Microscopy Sciences 50-980-487 4% PFA DPBS 30 mL Gibco 14190-094 Permeabilization Buffer (Table 3)
Table 3. Permeabilization Buffer
Reagent Final % wtv Stock Amount Supplier Product No. Donkey Serum 10% 100% 1 mL Canvax Biotech SUD004 Triton X 0.6% 10% 0.6 mL Roche 11332481001 DPBS 8.4 mL Gibco 14190-094 Dilution Buffer (Table 4)
Table 4. Dilution Buffer
Reagent Final % wtv Stock Amount Supplier Product No. Donkey Serum 10% 100% 0.1 mL Canvax Biotech SUD004 Triton X 0.1% 10% 0.01 mL Roche 11332481001 DPBS 0.89 mL Gibco 14190-094 Wash Buffer (Table 5)
Table 5. Wash Buffer
Reagent Final % wtv Stock Amount Supplier Product No. Triton X 0.1% 10% 0.5 mL Roche 11332481001 DPBS 49.5 mL Gibco 14190-094
Acknowledgments
The original protocol was described in Simonneau et al. (2021). We thank SUN bioscience for their support in adapting Gri3D plates for the growth of BBB organoids in the original publication. C.S. was supported by a Roche Postdoctoral Fellowship (RPF ID:491 2018-2020).
Competing interests
All authors were paid employees of F. Hoffmann-La Roche during the preparation of this protocol.
References
- Cho, C. F., Wolfe, J. M., Fadzen, C. M., Calligaris, D., Hornburg, K., Chiocca, E. A., Agar, N. Y. R., Pentelute, B. L. and Lawler, S. E. (2017). Blood-brain-barrier spheroids as an in vitro screening platform for brain-penetrating agents. Nat Commun 8: 15623.
- Hajal, C., Offeddu, G. S., Shin, Y., Zhang, S., Morozova, O., Hickman, D., Knutson, C. G. and Kamm, R. D. (2022). Engineered human blood-brain barrier microfluidic model for vascular permeability analyses. Nat Protoc 17(1): 95-128.
- Nzou, G., Wicks, R. T., Wicks, E. E., Seale, S. A., Sane, C. H., Chen, A., Murphy, S. V., Jackson, J. D. and Atala, A. J. (2018). Human Cortex Spheroid with a Functional Blood Brain Barrier for High-Throughput Neurotoxicity Screening and Disease Modeling. Sci Rep 8(1): 7413.
- Sade, H., Baumgartner, C., Hugenmatter, A., Moessner, E., Freskgard, P. O. and Niewoehner, J. (2014). A human blood-brain barrier transcytosis assay reveals antibody transcytosis influenced by pH-dependent receptor binding. PLoS One 9(4): e96340.
- Simonneau, C., Duschmale, M., Gavrilov, A., Brandenberg, N., Hoehnel, S., Ceroni, C., Lassalle, E., Kassianidou, E., Knoetgen, H., Niewoehner, J., et al. (2021). Investigating receptor-mediated antibody transcytosis using blood-brain barrier organoid arrays. Fluids Barriers CNS 18(1): 43.
- Urich, E., Patsch, C., Aigner, S., Graf, M., Iacone, R. and Freskgard, P. O. (2013). Multicellular self-assembled spheroidal model of the blood brain barrier. Sci Rep 3: 1500.
Article Information
Copyright
© 2022 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Kassianidou, E., Simonneau, C., Gavrilov, A. and Villaseñor, R. (2022). High Throughput Blood-brain Barrier Organoid Generation and Assessment of Receptor-Mediated Antibody Transcytosis. Bio-protocol 12(8): e4399. DOI: 10.21769/BioProtoc.4399.
Category
Neuroscience > Basic technology > High-throughput screening
Neuroscience > Nervous system disorders > Blood brain barrier
Cell Biology > Cell imaging > Fixed-cell imaging
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










