- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Activity-based Protein Profiling of Serine Hydrolase Superfamily Enzymes
Published: Vol 12, Iss 6, Mar 20, 2022 DOI: 10.21769/BioProtoc.4356 Views: 2776
Reviewed by: Agnieszka ZienkiewiczAntoni William JamesAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
Assessing Metabolite Interactions With Chloroplastic Proteins via the PISA Assay
Anna Karlsson [...] Elton P. Hudson
May 5, 2025 2039 Views
Advancing 2-DE Techniques: High-Efficiency Protein Extraction From Lupine Roots
Sebastian Burchardt [...] Emilia Wilmowicz
Oct 5, 2025 1792 Views

Quantitative Analysis of the Arabidopsis Leaf Secretory Proteome via TMT-Based Mass Spectrometry
Sakharam Waghmare [...] Rucha Karnik
Nov 20, 2025 2049 Views
Abstract
Activity-based protein profiling (ABPP) is a chemoproteomics platform to assess the functional state of enzymes in complex biological systems. Over the two decades, ABPP has emerged from a gel-based to gel-free platform, for in-depth proteome analysis with enhanced resolution, sensitivity for target detection, and discovery of small molecule inhibitors. The gel-free format of ABPP coupled with advanced mass spectrometry is highly sensitive and provides more comprehensive knowledge for the targeted enzyme family than the gel-based method. ABPP strategy is applied across microbe, plant, and animal models. It can be performed both in vitro and in vivo studies, and there is no limitation on sample origin. Here, we report an ultrasensitive, gel-free format of ABPP called active site peptide profiling. This protocol describes the identification of authentic functional proteins, by tagging their active sites in a native biological system. It is high throughput in nature and helps enrich even low abundance functional proteins. Since protein identification is virtually based on a single peptide, the identified peptide should be a unique peptide to identify its parent protein. It can be performed in a facile manner and offers to consolidate identification of protein targets as well as the site of probe modification. We have validated this approach using a fluorophosphonate (FP) serine hydrolase probe in the native proteome of the cereal crop Oryza sativa.
Graphic abstract:
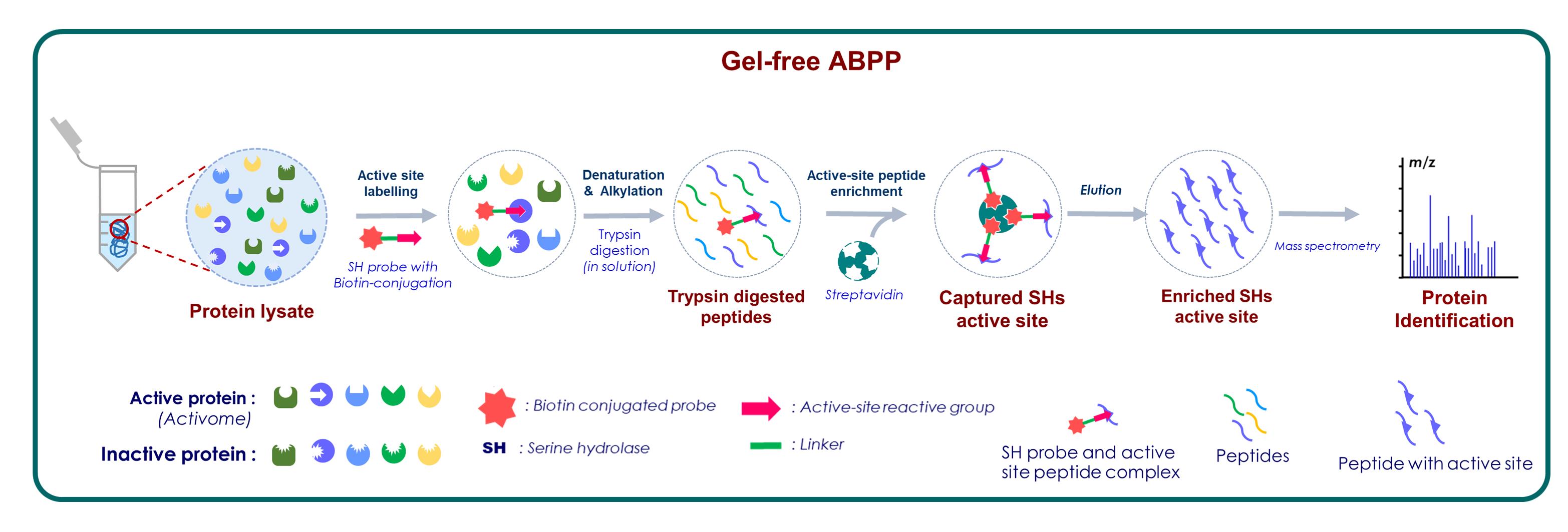
Serine hydrolase active site peptide profiling
Background
The advances in next-generation sequencing and long-read sequencing have opened the floodgate of many complex genomes. Inferring protein functions based on genome annotation may misannotate the catalytic properties to the uncharacterized genes, and it must be experimentally validated (Gerlt, 2016). Studying global analysis of gene transcription and translational changes by genomic and proteomic approaches gives only partial and indirect information about protein function. Further, protein activities are subjected to various post-translational modifications and regulations, which have pronounced effects on protein function (Bürkle, 2001). Therefore, conventional proteomic or genomic methods provide only an indirect assessment of the functional state of the enzyme. The lack of effective small-molecule probes for functional characterization has hindered our understanding of a myriad of functions and regulation modes. Activity-based proteomics is an analytical platform ideally suited for the global profiling of protein function (Schmidinger et al., 2006). In this approach, the functional annotation of uncharacterized proteins can be done precisely in an activity-dependent manner. Activity-based protein profiling (ABPP), pioneered by Cravatt and Bogyo, emerged as an efficient chemical proteomic approach for the analysis of functional states of target proteins in native biological systems (Evans and Cravatt, 2006, Cravatt et al., 2008, Niphakis and Cravatt, 2014). The application of ABPP depends on the quality of the probe, a small molecule that covalently binds to the active site of the target protein. In principle, an ABPP probe contains three parts: i) the reactive group, that forms a covalent bond with a conserved active site nucleophile, ii) a linker region, that attaches the reactive group with a reporter tag, and iii) a reporter tag, that provides a handle to detect and measure the protein labelled by the reactive group.
Since its inception, the most mature format for the visualization and characterization of probe-labelled proteomes in ABPP experiments is the in-gel fluorescence scanning (IGFS) (Patricelli et al., 2001). The identity of the proteins detected in IGFS can be achieved by enrichment followed by classical in-gel trypsin digestion (Dolui and Vijayaraj, 2020a; Latha et al., 2021). For example, replacement of a fluorescence tag with desthiobiotin facilitates the enrichment of the labelled enzymes via streptavidin/avidin affinity purification and subsequent identification through the mass spectrometric platform (Cravatt et al., 2008). This gel-based ABPP method directly links the labelling patterns of in-gel fluorescence signals and their identities (Phillips and Bogyo, 2005). However, there are some limitations to the gel-based ABPP method. The major limitation is the poor resolution of proteins that co-migrate on the gel, and identifying low-abundance proteins is difficult. With the advancement in proteomic techniques, ABPP has also significantly progressed from a gel-based to a virtually gel-free mode (Park et al., 2012). Since the activity-based probe covalently binds to the active site of functional proteins, the identity of the protein with the precise active-site residue information can be determined.
In this study, we described a gel-free format of ABPP for serine hydrolases active-site peptide profiling, using an active-site targeted probe (Dolui and Vijayaraj, 2020a). It provides comprehensive information for the active serine hydrolases with specific active-site residues mapping. In this approach, the number of probe-labelled active site peptides (in the trypsin digest of a probe-labelled proteome) logically equals the number of labelled proteins present in the sample (Okerberg et al., 2005). This gel-free ABPP format offers many advantages. First, performing an affinity enrichment step before mass spectrometry analysis helps to identify a low copy number of target enzymes. Therefore, it is high throughput in nature. Second, every single unique peptide will be able to reveal the identity of the parent target proteins. Since this peptide carries active site information, it is advantageous, and this information can be utilized for designing inhibitors for those target proteins. However, one of the prerequisites of this method is the accessibility for the active sites of enzymes in a native biological system. Therefore, pre-clearing the sample is an inevitable step prior to active site peptide profiling. Hence, we have developed a single-step pre-clearing method for lipid-rich samples for serine hydrolase labelling (Dolui and Vijayaraj, 2020b). Finally, multiple controls along with unbiased selection criteria should be adopted, to rule out any background and nonspecific proteins.
Materials and Reagents
1.7 mL microtubes (Axygen, catalog number: GEN-MT-175-C)
15 mL centrifuge tubes (Axygen, catalog number: GEN-CT-15 mL)
Gloves (Tarsons, catalog number: 370110)
Pipette tips (Axygen, catalog numbers: T-300 [10 µL], T-200-Y [200 µL], T-1000-B [1,000 µL])
Lab Apron (Medifit)
Zeba spin desalting columns, 7K MWCO, 5 mL (Thermo Scientific, catalog number: 89892)
ActiveX Desthiobiotin-FP Serine Hydrolase (Thermo Scientific, catalog number: 88317)
High capacity streptavidin agarose (Thermo Scientific, catalog number: 20357)
Dimethyl Sulphoxide, DMSO (Sigma, catalog number: D2650)
Dithiothreitol, DTT (Sigma, catalog number: D9779)
Iodoacetamide (Sigma, catalog number: I6125)
MS grade trypsin from porcine pancreas (Sigma, catalog number: T6567)
Trifluoroacetic acid,TFA (Sigma, catalog number: T6508-10AMP-KC)
LC-MS grade water (J.T. Baker, catalog number: 39253)
LC-MS grade acetonitrile, ACN (J.T. Baker, catalog number: 9829-03)
Urea (MP Biomedicals, catalog number: 194857)
Phosphate-buffered saline (PBS, pH 7.4)
NP-40 (Sigma, catalog number: NP40S)
Ethylenediaminetetraacetic acid, EDTA (MP Biomedicals, catalog number: 194822)
Tris (Himedia, catalog number: MB029-500G)
Sodium chloride, NaCl (Sigma, catalog number: S3014-500G)
Protein extraction buffer (see Recipes)
Assay buffer (see Recipes)
Washing buffer (see Recipes)
Elution buffer (see Recipes)
Solvent A (see Recipes)
Solvent B (see Recipes)
Equipment
-80°C freezer (New Brunswick, Ultra-low temperature freezer, model-U410), for long term storage of the probe
-20°C freezer (ARCTIKO)
Single-channel pipette (Eppendorf, model: Research Plus, catalog numbers: K31824D [10 µL capacity], L24004D [100 µL capacity], L25129D [1,000 µL capacity])
Ice flaking machine (Model-CB 246 HC)
Ice bucket
UV-Spectrophotometer (Jasco, V-630Bio) for protein quantification
ROTOSPINTM – End to end Test Tube Rotator (Tarsons, model: 3071)
Water bath (Grant, SAP5, SL. no. UD1541001)
AccuBlock digital dry bath (Labnet, model: D1100)
Centrifuge (Eppendorf, model: 5418)
Centrifuge (Eppendorf, model: 5810R) with 15 mL adapter
Ultracentrifuge (Beckman Coulter, model: Optima XPN-100)
Shaker (New Brunswick Scientific, SL. No. SE24FG001727)
Concentrator plus (Eppendorf, Code: E5305000363)
Vortex mixer (Velp Scientifica, Code No. F202A0176)
Weighing balance (Sartorius, model: CP423S)
Thermo Scientific LTQ or LTQ Orbitrap XL Mass Spectrometer (for analysis)
Pre-column (75 µm × 2 cm; Nanoviper C18, 3 µm; 100Å)
Liquid chromatography analytical column (RSLC C18 3 µm; 50 cm × 75 µm; 100Å)
Procedure
Active site peptide enrichment
Aliquot 1 mg of protein in a 1.7 mL microtube, and make up the volume to 500 μL with assay buffer (described in Recipes section).
Add 10 μL (20 µM ) of desthiobiotin–FP serine hydrolase probe or DMSO (No probe control) as control and incubate at 37°C for 1 h.
Stop the reaction by adding 500 μL of 10 M urea. Next, add 10 μL of 500 mM DTT, and incubate the sample at 65°C for 30 min.
Allow the sample to cool down to room temperature (RT). Add 40 μL of 1 M iodoacetamide and incubate protected from light at RT for 30 min.
Desalt the protein using a Zeba spin desalting column, to reduce the urea to a suitable concentration for trypsin digestion (as per the manufacturer’s instruction).
Digest the desalted protein with 20 μg of MS grade trypsin at 150 rpm and 37°C for 7 h.
Affinity captur the labelled peptides on 75 μL of 50% high capacity streptavidin agarose, by incubating the matrix with the digested peptides at constant rotation at RT for 1 h.
Pellet the labelled peptides bound to streptavidin agarose beads by centrifugation at 1,000 × g at RT for 1 min .
Wash the beads three times with 1.5 mL of washing buffer (see Recipes). Mix the beads by briefly vortexing after the addition of the buffer. Centrifuge the sample at 1,000 × g at RT for 1 min to collect the beads, and carefully remove the supernatant without disturbing them. This precaution was followed for all the subsequent steps of washing.
Wash the beads three times with 1.5 mL of PBS buffer (pH 7.4, see Recipes) followed by three washes with 1.5 mL of LC-MS grade water.
Elute the captured peptides by adding 100 μL of elution buffer (see Recipes) and incubating the sample at RT for 3 min. This step was repeated two additional times and the eluted fractions were pooled together and lyophilized by vacuum concentration.
Mass spectrometry analysis of enriched probe targets
Suspend the lyophilized and labelled peptides in 0.1% TFA and inject them directly onto a Thermo LTQ Orbitrap coupled with an EASY nano LC1200 system (Agilent).
Set the column temperature to 35°C and limit the flow rate to 250 nL/min.
The LC solvent system consists of two mobile phases, such as Solvent A (see Recipes) and Solvent B (see Recipes).
Separate the peptides on the analytical column“[EASY SPRAY PREAMP (RSLC C18 3 µm; 50cm × 75µm; 100Å) silica column]”, by running a 180 min gradient of solvents A and B (Start with 10% B for 2 min, gradient 10–45% for 166 min, gradient 45–95% for up to 172 min).
Set the mass spectrometer in positive ion mode with the following parameters: precursor ion scanning in an orbitrap analyzer in the range of 375–1,700, at a resolution of 120,000, with a maximum ion injection time of 50 ms; high collision energy (CID) to 30% for the MS/MS spectra generation; MS2 scan range (m/z) to 100–2,000, with a maximum ion injection time of 35 ms.
Data analysis
Peptide and protein identification
Convert raw data files to Mascot Generic File (MGF) files.
Search the MS/MS spectra data using a Mascot search engine (version 2.7.1.0).
The search parameters included: fixed modification of carbamidomethyl (57.02146 Da); variable modifications of acetyl (protein N-term) (42.01057 Da); oxidation of methionine (15.99492 Da); biotin: Thermo-88317 (S) (443.2913Da). Set enzyme specificity to Trypsin/P, and allow a maximum missed cleavage of 2.
Set the peptide mass tolerance to 10 ppm, fragment mass tolerance to 0.6 Da, ion score cut-off to 30, and the significance threshold to 0.05.
Statistical Analysis
Data are represented as mean ± standard deviation of three independent experiments. The significance of difference at p <0.01 was determined by one-way ANOVA with Tukey post hoc test using IBM SPSS Statistics software.
Recipes
Protein extraction buffer
50 mM Tris-Cl (pH 8.0)
150 mM NaCl
2.0 mM KCl
2.0 mM MgCl2
10% glycerol
Autoclave at 121°C for 20 min and store at 4°C afterward.
Assay buffer
50 mM Tris-Cl (pH 8.0)
50 mM NaCl
10% glycerol
Autoclave at 121°C for 20 min.
Washing buffer
25 mM Tris-Cl (pH 7.4)
350 mM NaCl
1% NP-40
5% glycerol
Active site peptide elution buffer
50% ACN
0.2% of TFA
Dilute 20 μL of TFA with 5.0 mL of ACN and 5.0 mL of LC-MS grade water.
Solvent A
0.1% Formic acid in HPLC grade water
Solvent B
80% acetonitrile and 0.1% formic acid in HPLC grade water
Acknowledgments
The work was supported by the Department of Science and Technology, New Delhi, India, under the DST-INSPIRE Faculty Scheme (IFA14-LSPA28). AKD was supported by CSIR-UGC-JRF. M.L was sponsored by UGC-Rajiv Gandhi National Fellowship (RGNF) Award.
Competing interests
The authors declare no competing interests.
References
- Bürkle, A. (2001). Posttranslational Modification. In: Brenner, S. and Miller, J. H. (Eds.). Encyclopedia of Genetics. Academic Press, New York, NY, USA. p1533.
- Cravatt, B. F., Wright, A. T. and Kozarich, J. W. (2008). Activity-based protein profiling: from enzyme chemistry to proteomic chemistry. Annu Rev Biochem 77: 383-414.
- Dolui, A. K. and Vijayaraj, P. (2020a). Functional Omics Identifies Serine Hydrolases That Mobilize Storage Lipids during Rice Seed Germination. Plant Physiol 184(2): 693-708.
- Dolui, A. K. and Vijayaraj, P. (2020b). A solvent-free delipidation method for functional validation of lipases. 3 Biotech 10(8): 343.
- Evans, M. J. and Cravatt, B. F. (2006). Mechanism-based profiling of enzyme families. Chem Rev 06(8): 3279-301.
- Gerlt, J. A. (2016). Tools and strategies for discovering novel enzymes and metabolic pathways. Perspect Sci 9: 24-32.
- Latha, M., Dolui, A. K. and Vijayaraj, P. (2021). Proteoform of Arabidopsis seed storage protein identified by functional proteomics approach exhibits acyl hydrolase activity during germination. Int J Biol Macromol 172: 452-463.
- Niphakis, M. J. and Cravatt, B. F. (2014). Enzyme inhibitor discovery by activity-based protein profiling. Annu Rev Biochem 83: 341-77.
- Okerberg, E. S., Wu, J., Zhang, B., Samii, B., Blackford, K., Winn, D. T., Shreder, K. R., Burbaum, J. J. and Patricelli, M. P. (2005). High-resolution functional proteomics by active-site peptide profiling. Proc Natl Acad Sci U S A 102(14): 4996-5001.
- Park, J., Oh, S. and Park, S. B. (2012). Discovery and target identification of an antiproliferative agent in live cells using fluorescence difference in two-dimensional gel electrophoresis. Angew Chem Int Ed Engl 51(22): 5447-5451.
- Patricelli, M. P., Giang, D. K., Stamp, L. M. and Burbaum, J. J. (2001). Direct visualization of serine hydrolase activities in complex proteomes using fluorescent active site-directed probes. Proteomics 1(9): 1067-71.
- Phillips, C. I. and Bogyo, M. (2005). Proteomics meets microbiology. Technical advances in the global mapping of protein expression and function. Cell Microbiol 7(8): 1061-1076.
- Schmidinger, H., Hermetter, A. and Birner-Gruenberger, R. (2006). Activity-based proteomics: enzymatic activity profiling in complex proteomes. Amino Acids 30(4): 333-50.
Article Information
Copyright
© 2022 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Readers should cite both the Bio-protocol article and the original research article where this protocol was used:
- Dolui, A. K., Latha, M. and Vijayaraj, P. (2022). Activity-based Protein Profiling of Serine Hydrolase Superfamily Enzymes. Bio-protocol 12(6): e4356. DOI: 10.21769/BioProtoc.4356.
- Dolui, A. K. and Vijayaraj, P. (2020a). Functional Omics Identifies Serine Hydrolases That Mobilize Storage Lipids during Rice Seed Germination. Plant Physiol 184(2): 693-708.
Category
Plant Science > Plant biochemistry > Protein
Biochemistry > Protein > Activity
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link









