- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Activity-based Crosslinking to Identify Substrates of Thioredoxin-domain Proteins in Malaria Parasites
Published: Vol 12, Iss 4, Feb 20, 2022 DOI: 10.21769/BioProtoc.4322 Views: 2933
Reviewed by: Alexandros AlexandratosMigla MiskinyteDarrell CockburnPetru-Iulian Trasnea

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

Liposome Disruption Assay to Examine Lytic Properties of Biomolecules
John R. Jimah [...] Niraj H. Tolia
Aug 5, 2017 14310 Views
In vitro Di-ubiquitin Formation Assay and E3 Cooperation Assay
Rebecca J. Burge [...] Jeremy C. Mottram
Nov 5, 2022 1904 Views
Monitoring Protein Stability In Vivo Using an Intein-Based Biosensor
John S. Smetana [...] Christopher W. Lennon
Apr 20, 2025 1619 Views
Abstract
Malaria remains a major public health issue, infecting nearly 220 million people every year. The spread of drug-resistant strains of Plasmodium falciparum around the world threatens the progress made against this disease. Therefore, identifying druggable and essential pathways in P. falciparum parasites remains a major area of research. One poorly understood area of parasite biology is the formation of disulfide bonds, which is an essential requirement for the folding of numerous proteins. Specialized chaperones with thioredoxin (Trx) domains catalyze the redox functions necessary for breaking incorrect and forming correct disulfide bonds in proteins. Defining the substrates of these redox chaperones is difficult and immunoprecipitation based assays cannot distinguish between substrates and interacting partners. Further, the substrate or client interactions with the redox chaperones are usually transient in nature. Activity based crosslinkers that rely on the nucleophilic cysteines on Trx domains and the disulfide bond forming cysteines on clients provide an easily scalable method to trap and identify the substrates of Trx-domain containing chaperones. The cell permeable crosslinker divinyl sulfone (DVSF) is active only in the presence of nucleophilic cysteines in proteins and, therefore, traps Trx domains with their substrates, as they form mixed disulfide bonds during the course of their catalytic activity. This allows the identification of substrates that rely on Trx activity for their folding, as well as discovering small molecules that interfere with Trx domain activity.
Graphic abstract:
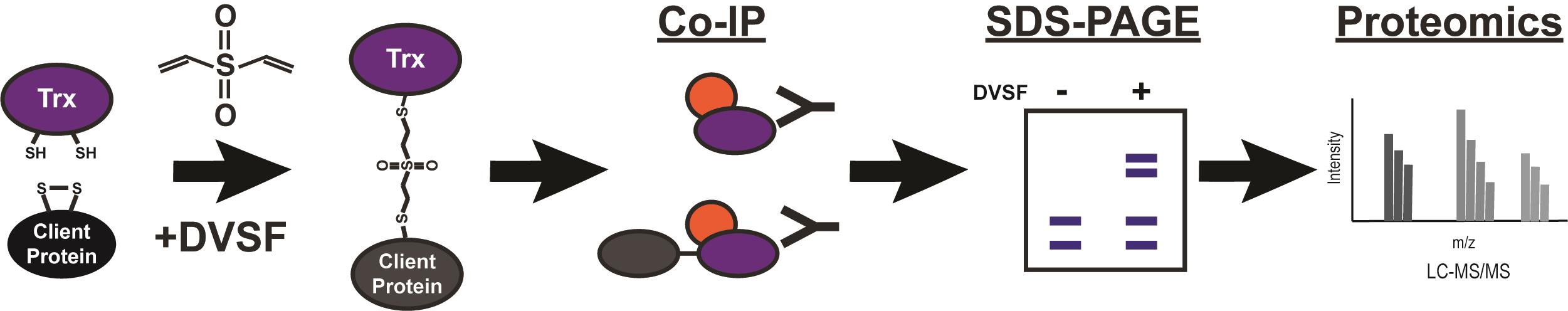
Identification of thioredoxin domain substrates via divinylsulfone crosslinking and immunoprecipitation-mass spectrometry.
Background
Disulfide bonds are crucial for the activity and regulation of many proteins; therefore, disulfide bond formation and reduction is a critical aspect of cellular life. Members of the thioredoxin (Trx) superfamily use the nucleophilic cysteines of their “CXXC” active sites to facilitate the formation and reduction of disulfide bonds (Lu and Holmgren, 2014). Given their importance for cell survival, thioredoxin-domain proteins and the thioredoxin system have been proposed as drug targets against apicomplexan parasites (Becker et al., 2000; Krnajski et al., 2001; Andricopulo et al., 2006; Kehr et al., 2010; Biddau et al., 2018; Biddau and Sheiner, 2019; Cobb et al., 2021). Despite being essential and their potential as drug targets, the substrates of these Trx-domain proteins are often ill-defined, as it can be difficult to discern specific substrates from other interacting partners.
One strategy for identifying the substrates of Trx domains relies on the active site mode of action: the first cysteine of the “CXXC'' active site forms a mixed disulfide bond with a client protein, and the sulhydryl group of the second cysteine is used to resolve that bond. Mutagenesis of the second cysteine, often to alanine or serine, prevents the reduction of the bond between the Trx-domain and client protein, trapping the two proteins together (Jessop et al., 2007; Oka et al., 2013). Subsequently, mass spectrometry can be used to identify the substrate. A major disadvantage to this approach is that it requires modification to the active sites of endogenous enzymes, that may be essential, or exogenous expression of the mutated enzymes, both of which may prove lethal for the organism. Particularly in organisms like Plasmodium falciparum, which is haploid for the majority of its lifecycle and carries single copies of genes for essential Trx-domain proteins, this approach may prove especially difficult.
Given the challenges inherent to a genetic approach, activity based chemical crosslinking of Trx-domains to their substrates is a straightforward and powerful option for identifying those substrates. Divinyl sulfone (DVSF) is a bifunctional, electrophilic, and cell permeable crosslinker shown to have remarkable specificity for nucleophilic cysteines, like those within Trx-domain active sites (West et al., 2011). Treatment of cells with DVSF results in irreversible crosslinking between redox-active cysteines, and it has been used to trap and identify substrates of Trx-domain proteins and other redox-active enzymes in yeast and human cells (Naticchia et al., 2013; Allan et al., 2016; Araki et al., 2017). Demonstrating the power of this activity-based technique for identifying Trx-domain substrates in a non-model organism, we used DVSF to trap and identify substrates of Trx-domain proteins in the endoplasmic reticulum (ER) of P. falciparum parasites (Cobb et al., 2021). We also demonstrated its specificity for redox-active nucleophilic cysteines, as other cysteine-containing proteins did not react with DVSF, and further validated our results by demonstrating ablation of crosslinking, by mutation of reactive cysteine residues to alanine residues (Cobb et al., 2021). In this protocol, we describe the use of DVSF to trap substrates of ER-localized Trx-domain proteins in P. falciparum and the identification of these trapped substrates via mass spectroscopy. Given the robust nature of this protocol, it will be of use for similar experiments in diverse organisms and within multiple cellular compartments. Because this approach can be utilized with minimal genetic modification, requiring only the addition of an affinity tag to the Trx-domain protein if a suitable antibody is not otherwise available, this protocol will allow others working with difficult organisms to study the interactions between Trx-domains and their substrates.
Materials and Reagents
15 mL centrifuge tubes (Genesee Scientific, catalog number: 28-103)
1.7 mL microtubes (Genesse Scientific, catalog number: 28-281)
Filter tip pipette tips: 10 μL (Fisher Scientific, catalog number: 02-707-473), 200 μL (Fisher Scientific, catalog number: 02-707-478), 1 mL (Fisher Scientific, catalog number: 2707480)
Serological pipettes: 5 mL (Genesee Scientific, catalog number: 12-102), 10 mL Genesee Scientific, catalog number: 12-104)
TPP Tissue culture plate, 100 mm (MidSci, catalog number: TP92406)
O+ AS-1 Packed Cells (Interstate Blood Bank, Memphis, TN)
IRDye® 800CW anti-mouse (Li-Cor, catalog number: 926-32210 used at 1:20,000 for Western blot)
IRDye® 680RD anti-rabbit (Li-Cor, catalog number: 926-68071, used at 1:20,000 for Western blot)
Rabbit anti-HA antibody (Life Technologies, catalog number: 715500, used at 1:100 for Western blot)
Mouse anti-PMV antibody (Gift from Daniel Goldberg, Washington University School of Medicine, used at 1:400 for Western blot)
V5-Tag (D3H8Q) Rabbit mAb (Cell Signaling Tech, Inc., catalog number: 13202S, used at 1:1,000 for Western blot)
Anti-HA Magnetic Beads (Thermo Fisher Scientific/PierceTM, catalog number: 88836)
Divinyl sulfone, 97% stab. with 0.05% hydroquinone (Thermo Fisher Scientific/Alfa Aesar, catalog number: L12827)
Saponin Quillaja sp., Sapogenin content 20-35% (Sigma, catalog number: S4521-10G)
Protein sample loading buffer, 4× (Li-Cor, catalog number: 928-40004)
Chameleon Duo pre-stained protein ladder (Li-Cor, catalog number:928-60000)
2-mercaptoethanol (Fisher Scientific, catalog number: 21985023)
10× Tris/Glycine/SDS buffer (Bio-Rad, catalog number: 161-0732)
4-20% Mini-PROTEAN® TGXTM precast protein gels (Bio-Rad, catalog number: 456-1093)
InterceptTM blocking buffer (Li-Cor, catalog number: 927-70003)
Odyssey® nitrocellulose membrane (Li-Cor, catalog number: 926-31092)
Halt protease inhibitor cocktail (Thermo Fisher Scientific/Life Technologies, catalog number: 78429)
HA synthetic peptide (Thermo Fisher Scientific/Life Technologies, catalog number: 26184)
Simply Blue SafeStain (Thermo Fisher Scientific/Life Technologies, catalog number: LC6060)
Tris-HCl (Fisher Bioreagents, catalog number: 1185-53-1)
KCl (Sigma, catalog number: 7447-40-7)
EDTA (Fisher Bioreagents, catalog number: 60-00-4)
NP-40 (Sigma, catalog number: 9002-93-1)
Na2HPO4 (Millipore Sigma, catalog number: 1065861000)
KH2PO4 (Millipore Sigma, catalog number: 529568)
NaCl (Fisher Scientific, catalog number: BP358212)
TRIS base (Fisher Scientific, catalog number: YBP152500)
Glycine (Fisher Scientific, catalog number: ALF-036435-A1)
Methanol (Fisher Scientific, catalog number: A4081)
Tween-20 (Fisher Scientific, catalog number: BP337-100)
Immunoprecipitation extraction buffer (see Recipes)
Immunoprecipitation binding buffer (see Recipes)
Phosphate buffered saline (PBS), pH 7.4 (see Recipes)
Protein loading dye (see Recipes)
10× Transfer buffer (see Recipes)
1× Transfer buffer (see Recipes)
0.1% Tween-20, in 1× PBS (see Recipes)
10× Tris-buffered saline (TBS) (see Recipes)
2 mg/mL HA peptide (see Recipes)
Equipment
SorvallTM LegendTM XTR Centrifuge (Thermo Fisher Scientific, model: 75004506)
Odyssey® CLx Imaging System (Li-Cor, model: 9140)
DynaMagTM-2 Magnet (Life Technologies, model: 12321D)
Branson S450D Digital Sonifier (Thermo Fisher Scientific, model: 101-063-593)
Mini-PROTEAN Tetra Cell System (Bio-Rad, catalog number: 1658005EDU)
PowerPacTM Basic Power Supply (Bio-Rad, catalog number: 1645050)
FormaTM Series II Water-Jacketed CO2 incubator (ThermoFisher Scientific, catalog number: 3110)
Software
Image Studio 5.x CLx (Li-Cor, https://www.licor.com/bio/image-studio-lite/)
Procedure
Trap Trx-domain substrates with DVSF
Incubate cells with DVSF
Transfer cultures of red blood cells (RBCs) infected with P. falciparum parasites and expressing an epitope-tagged Trx-domain protein, into sterile 15 mL conical tubes. As a starting point, use 10 mL of culture at 2% hematocrit, with 10-15% parasitemia.
Pellet the infected RBCs by centrifuging at 800 × g for 3 min.
Remove the supernatant.
Use a serological pipette to resuspend the RBCs in an equal volume of either 3 mM DVSF prepared in PBS, or PBS alone as a control (e.g., pellet 10 mL of culture, and resuspend the cells in 10 mL of 3 mM DVSF or PBS).
Incubate the cells at 37°C, for 30 min in a CO2 incubator.
Pellet the cells by centrifuging at 800 × g for 3 min.
Remove the supernatant.
Dispose of the DVSF supernatant according to your institution’s chemical waste policies.
Wash the cells three times with PBS, using an equivalent volume of PBS (e.g., use 10 mL of PBS if 10 mL of DVSF was used), and performing spins at 800 × g for 3 min.
Extract protein from the cells
Prepare a solution of 0.04% (w/v) saponin in PBS and cool on ice, until the buffer is ice-cold.
Isolate parasites by lysing RBCs in 0.04% saponin.
Resuspend cells in 1 mL of ice-cold saponin per 4 mL of culture that was pelleted (e.g., resuspend cells pelleted from 10 mL of culture in 2.5 mL of saponin).
Incubate on ice for 15 min.
Pellet parasites by centrifuging at 2,000 × g for 5 min, at 4°C.
Remove the supernatant and wash two times with ice-cold PBS, using the same volume of PBS as the volume of saponin used in Step A.2.a, performing spins at 2,000 × g for 5 min.
Resuspend parasites in 50-100 μL of protein loading dye.
Boil parasites at 95°C for 5 min.
Pellet insoluble material, including parasite hemozoin, by centrifuging at full speed on a table-top centrifuge for 10 min, at room temperature.
Transfer supernatant, which contains extracted proteins, to a clean tube.
Demonstrate successful cross-linking by Western blot: compared to the PBS-only control sample, the DVSF-crosslinked sample should contain one or more bands at higher molecular weight, as the Trx-domain protein + substrate will migrate slower during SDS-PAGE than the Trx-domain protein alone (Figure 1).
Load 30-50 μL of protein sample and a protein ladder into a polyacrylamide gel.
Separate the proteins in the DVSF-crosslinked and PBS-only samples by SDS-PAGE, at 180 V for ~45 min, or until the bands of the protein ladder are sufficiently resolved.
Transfer proteins from the gel to a nitrocellulose membrane, at 100 V for 1 h.
Prepare the membrane by cutting it to the approximate size of the gel and pre-wetting in 1× transfer buffer before applying to the gel.
Block the membrane for 1 h, at room temperature with gentle shaking, using 30 mL of 50% InterceptTM blocking buffer (v/v, prepared in 1× PBS).
Remove the blocking buffer, and add 10 mL of primary antibody solution, prepared in 50% blocking buffer described above.
Use primary antibodies directed against the Trx-domain protein of interest and a loading control.
Incubate the membrane in the primary antibody solution at 4°C overnight, in a covered container, with gentle shaking.
Remove the primary antibody solution and wash three times with 30 mL of 0.1% Tween-20 (v/v, prepared in 1× PBS), for 10 min, at room temperature with gentle shaking.
Add 10 mL of secondary antibody solution prepared in 50% blocking buffer, as described in Step A.2.g.iv. above.
Incubate for 1 h, at room temperature, in a covered container, with gentle rocking.
Remove the secondary antibody solution and wash three times with 30 mL of 0.1% Tween-20 (v/v, prepared in 1× PBS), for 10 min, at room temperature with gentle shaking.
Wash the membrane with 30 mL of 1× PBS for 5 min, at room temperature with gentle shaking.
Image the membrane with the Odyssey® CLx Imaging System to visualize cross-linking of proteins to the Trx-domain protein of interest.
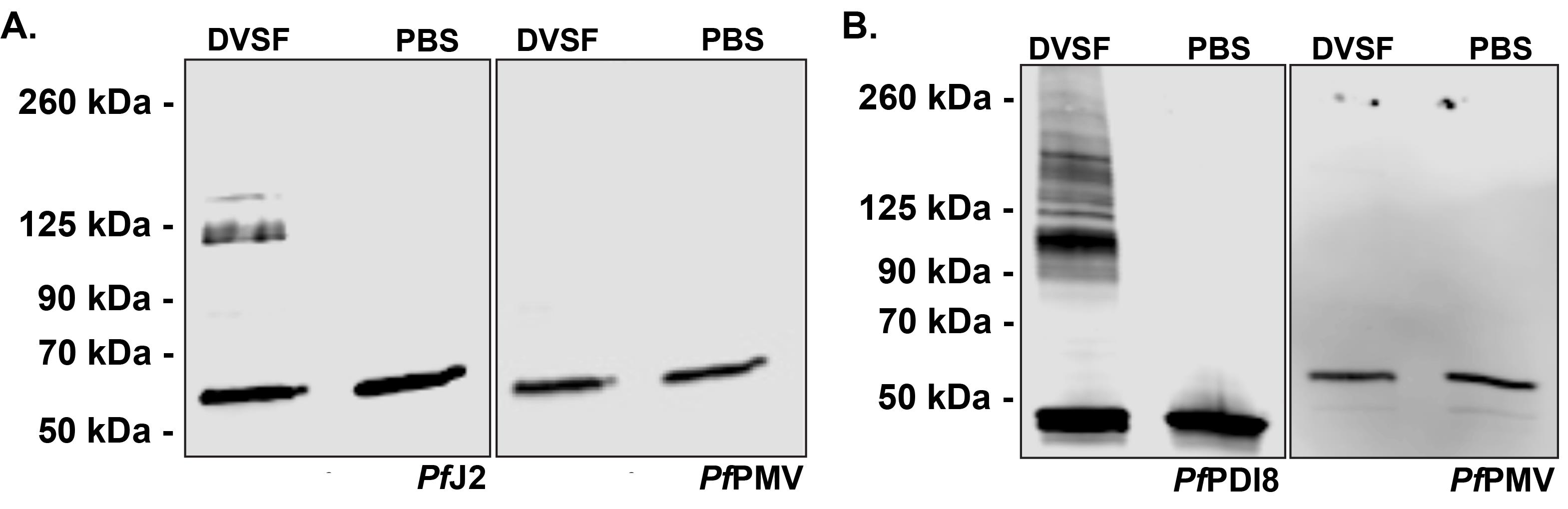
Figure 1. DVSF crosslinking traps redox partners to P. falciparum Trx-domain proteins. Parasites were incubated with 3 mM DVSF for 30 min at 37°C, then proteins were extracted and analyzed via Western blot. The parasites expressed epitope-tagged A.) PfJ2 or B.) PfPDI8. Plasmepsin V (PfPMV) was used as a loading control, and as a non-Trx-domain protein, demonstrating the specificity of DVSF.
Immunoprecipitate Trx-domain proteins with trapped substrates
Extract protein from parasites
Perform DVSF crosslinking and isolate parasites using 0.04% saponin, as described above in section A; importantly, include the PBS-only control parasites. For immunoprecipitation for mass spectroscopy protein identification, we start with 1 × 109 parasites and scale up or down as necessary.
Resuspend the parasite pellet in 300 μL of extraction buffer, then add 15 μL of 10% NP-40, and 3 μL of HALT protease inhibitor.
Incubate on ice for 15 min.
Lyse parasites by sonication.
Sonicate parasites with an amplitude of 10%, for 5 s.
Let the sample rest on ice for 30 s.
Repeat Steps B.1.d.i and -ii two more times, for a total of three rounds of sonication.
Centrifuge the lysate at full speed on a tabletop centrifuge set to 4°C, for 15 min.
Transfer the supernatant, containing extracted proteins, to a new tube.
Perform a second extraction on the pellet from Step B.1.e.
Add 300 μL of extraction buffer and 3 μL of HALT protease inhibitor to the pellet (no NP-40 is used for the second extraction) and proceed immediately to the sonication step without incubating on ice for 15 min.
Sonicate and centrifuge as described in Steps B.1.a-e.
Combine the supernatant with the extracted proteins from Step B.1.f.
Transfer 60 μL of the combined supernatants to a clean tube as the “Input” sample and store at -20°C.
Immunoprecipitate HA-tagged proteins
Prepare Pierce anti-HA magnetic beads.
Transfer 25 μL of the bead slurry to a clean 1.5 mL centrifuge tube.
Add 175 μL of binding buffer to the beads, flick the tube to mix, and briefly centrifuge to collect all of the liquid at the bottom of the tube.
Insert the tube into a magnetic rack and remove the buffer after the beads have collected to the side of the tube.
Remove the tube and add 1 mL of binding buffer to the beads.
Rotate the tube in your hand to mix beads and buffer, for 1 min.
Use the magnetic rack to remove the buffer from the beads.
Add the extracted proteins from Step B.1.h to the beads.
Incubate with end-over-end rotation for 30 min at room temperature (or overnight at 4°C).
Briefly centrifuge the tube to collect all of the liquid to the bottom, then use the magnetic rack to remove the unbound protein sample from the beads; save this sample in a tube labeled “Unbound”.
Wash the beads three times with binding buffer, and save the buffer from each wash for downstream analysis if necessary.
Resuspend the beads in 300 μL of binding buffer and use the magnetic rack to remove the buffer from the beads.
Repeat Step B.2.e.i two more times, for a total of three washes.
Elute proteins from the beads using the HA peptide.
Resuspend the beads in 30 μL of 2 mg/mL HA peptide.
Incubate at 37°C, shaking, for 5 min.
Use the magnetic rack to separate the eluted proteins from the beads, and save the eluted proteins in a tube labeled “Elution”.
Extract cross-linked proteins from acrylamide gels for protein identification
Load 5 μL of protein ladder and the “Elution” samples of the immunoprecipitated proteins, collected in Step B.2.g above, into the wells of a polyacrylamide gel.
Separate the proteins in the sample by SDS-PAGE, at 180 V, for approximately 45 min, or until the protein ladder has sufficiently resolved.
Stain the proteins in the gel with SimplyBlue SafeStain, or another Coomassie stain (Figure 2).
Rinse the gel three times with 100 mL of deionized water.
Add approximately 20 mL of SimplyBlue SafeStain, or enough stain to completely cover the gel, and incubate at room temperature for 1 h, with gentle shaking.
Remove the stain and wash the gel with 100 mL of deionized water for 1 h, at room temperature, with gentle rocking.
After 1 h, add 20 mL of 20% NaCl (w/v, prepared in deionized water) and continue incubating for 2 h, or overnight, at room temperature with gentle shaking.
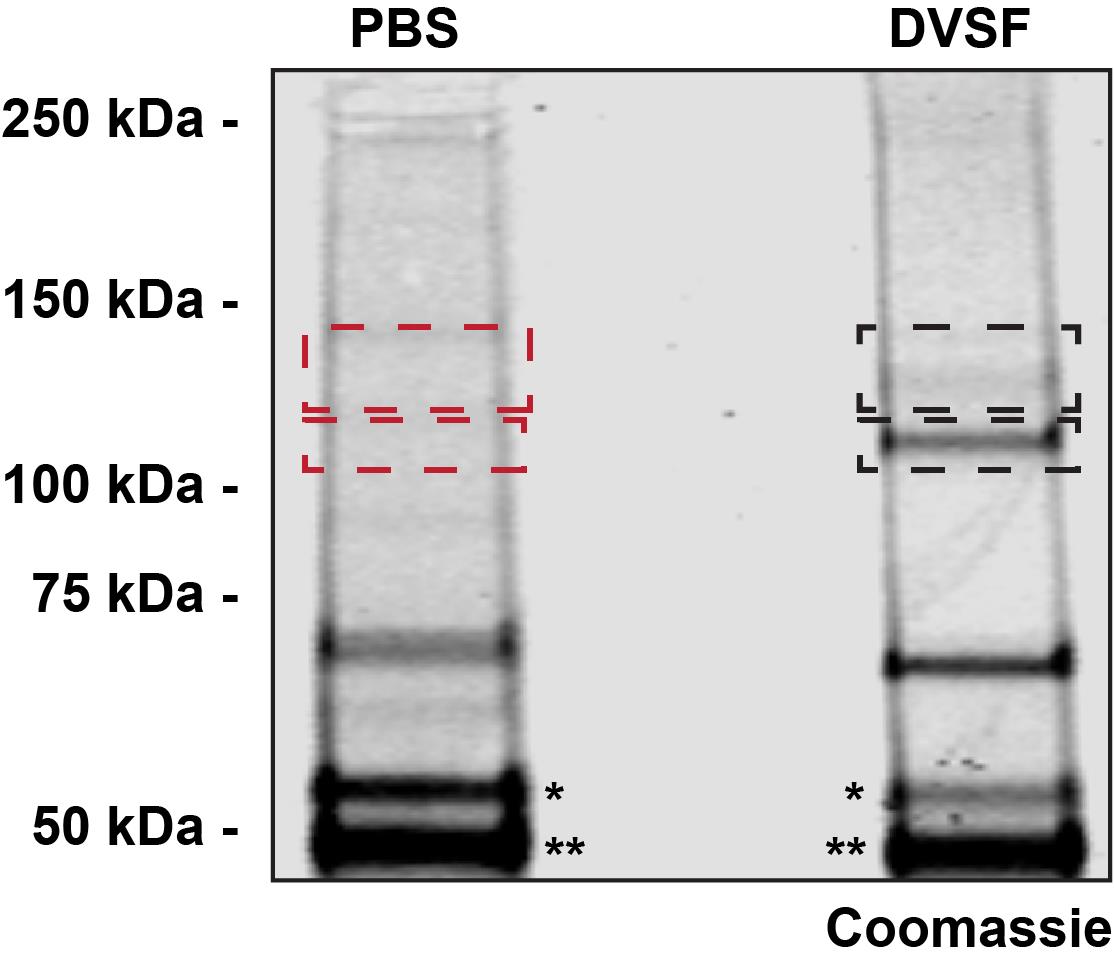
Figure 2. Coomassie staining of DVSF-crosslinked proteins prior to extraction for mass spectroscopy protein identification. Parasites expressing epitope-tagged PfJ2 were incubated with 3 mM DVSF for 30 min at 37°C, then PfJ2 was immunoprecipitated from parasite lysates. The immunoprecipitated proteins were separated using SDS-PAGE and stained with SimplyBlue SafeStain. The black, dashed boxes indicate areas of the gel in the DVSF-treated lane extracted for protein identification. The red, dashed boxes indicate areas of the gel in the PBS-treated lane extracted as controls for the protein identification.
Using a clean razor blade, extract the cross-linked protein bands from the gel (i.e., extract the bands that run higher in the gel than the non-crosslinked protein).
Extract slices from equivalent areas of the gel in the PBS-only control sample (e.g., extract a cross-linked protein band that runs at 100 kDa, and extract an equivalently sized piece of gel at 100 kDa in the PBS-only sample).
Transfer the gel slices into 1.5 mL centrifuge tubes.
Destain the proteins in the gel slices.
Incubate with 30% ethanol (v/v, prepared in deionized water) for 15 min, or until the gel slice is no longer stained, at room temperature, gently rotating.
Identify the proteins present in each gel slice, either in-house or through a mass-spectroscopy service.
Identify the potential Trx-domain substrates, by eliminating the proteins identified in the PBS-only gel slices from those identified in the DVSF-crosslinked samples.
Notes
We describe the use of DVSF for trapping, extracting, and identifying Trx-domain substrates in P. falciparum parasites, but the components of this protocol are easily adaptable for other organisms. Accordingly, the specifics of protein extraction, immunoprecipitation, and Western blotting should be adapted to the workflow that individual researchers have for their specific organisms.
Recipes
Immunoprecipitation extraction buffer
40 mM Tris-HCl pH 7.4
150 mM KCl
1 M EDTA
1× HALT protease inhibitor
0.5% NP-40 (v/v)
Immunoprecipitation binding buffer
20 mM Tris-HCl pH 7.4
150 mM KCl
1 mM EDTA
1× HALT protease inhibitor
0.1% NP-40 (v/v)
Phosphate buffered saline (PBS), pH 7.4
3.2 mM Na2HPO4
0.5 mM KH2PO4
1.3 mM KCl
135 mM NaCl
Protein loading dye
120 μL of 4× Li-Cor Protein Sample Loading Buffer
20 μL of 2-mercaptoethanol
10× Transfer buffer
30.3 g of TRIS base
144.1 g glycine
Bring to 1 L with Milli-Q or deionized water.
1× Transfer buffer
100 mL of 10× Transfer Buffer
200 mL of methanol
700 mL of Milli-Q or DI water
0.1% Tween-20, in 1× PBS
Add 1 mL of Tween-20 to 1 L of 1× PBS
10× Tris-buffered saline (TBS)
87.7 g NaCl
100 mL of 1 M Tris-HCl, pH 7.5
Milli-Q or deionized water to 1 L
Dilute to 1× before using
2 mg/mL HA Peptide
HA Synthetic Peptide was reconstituted in 1× TBS to a concentration of 2 mg/mL.
Acknowledgments
We thank Dan Goldberg for anti-PMV antibodies. This work was supported by awards from the ARCS Foundation to D.W.C., and the US National Institutes of Health to D.W.C. (T32AI060546) and to V.M. (R01AI130139).
Competing interests
The authors declare no conflicts of interest.
References
- Andricopulo, A. D., Akoachere, M. B., Krogh, R., Nickel, C., McLeish, M. J., Kenyon, G. L., Arscott, L. D., Williams, C. H., Jr., Davioud-Charvet, E. and Becker, K. (2006). Specific inhibitors of Plasmodium falciparum thioredoxin reductase as potential antimalarial agents. Bioorg Med Chem Lett 16(8): 2283-2292.
- Allan, K. M., Loberg, M. A., Chepngeno, J., Hurtig, J. E., Tripathi, S., Kang, M. G., Allotey, J. K., Widdershins, A. H., Pilat, J. M., Sizek, H. J., et al. (2016). Trapping redox partnerships in oxidant-sensitive proteins with a small, thiol-reactive cross-linker. Free Radic Biol Med 101: 356-366.
- Araki, K., Ushioda, R., Kusano, H., Tanaka, R., Hatta, T., Fukui, K., Nagata, K. and Natsume, T. (2017). A crosslinker-based identification of redox relay targets. Anal Biochem 520: 22-26.
- Becker, K., Gromer, S., Schirmer, R. H. and Müller, S. (2000). Thioredoxin reductase as a pathophysiological factor and drug target: Thioredoxin reductase in medicine and parasitology. Eur J Biochem 267 (20): 6118-25.
- Biddau, M., Bouchut, A., Major, J., Saveria, T., Tottey, J., Oka, O., van-Lith, M., Jennings, K. E., Ovciarikova, J., DeRocher, A., et al. (2018). Two essential Thioredoxins mediate apicoplast biogenesis, protein import, and gene expression in Toxoplasma gondii. PLoS Pathog 14(2): e1006836.
- Biddau, M. and Sheiner, L. (2019). Targeting the apicoplast in malaria. Biochem Soc Trans 47(4): 973-983.
- Cobb, D. W., Kudyba, H. M., Villegas, A., Hoopmann, M. R., Baptista, R. P., Bruton, B., Krakowiak, M., Moritz, R. L. and Muralidharan, V. (2021). A redox-active crosslinker reveals an essential and inhibitable oxidative folding network in the endoplasmic reticulum of malaria parasites. PLoS Pathog 17(2): e1009293.
- Jessop, C. E., Watkins, R. H., Simmons, J. J., Tasab, M. and Bulleid, N. J. (2009). Protein disulphide isomerase family members show distinct substrate specificity: P5 is targeted to BiP client proteins. J Cell Sci 122(Pt 23): 4287-4295.
- Krnajski, Z., Gilberger, T. W., Walter, R. D. and Muller, S. (2001). The malaria parasite Plasmodium falciparum possesses a functional thioredoxin system. Mol Biochem Parasitol 112(2): 219-228.
- Kehr, S., Sturm, N., Rahlfs, S., Przyborski, J. M. and Becker, K. (2010). Compartmentation of redox metabolism in malaria parasites. PLoS Pathog 6(12): e1001242.
- Lu, J. and Holmgren, A. (2014). The thioredoxin superfamily in oxidative protein folding. Antioxid Redox Signal 21(3): 457-470.
- Naticchia, M. R., Brown, H. A., Garcia, F. J., Lamade, A. M., Justice, S. L., Herrin, R. P., Morano, K. A. and West, J. D. (2013). Bifunctional electrophiles cross-link thioredoxins with redox relay partners in cells. Chem Res Toxicol 26(3): 490-497.
- Oka, O. B., Pringle, M. A., Schopp, I. M., Braakman, I. and Bulleid, N. J. (2013). ERdj5 is the ER reductase that catalyzes the removal of non-native disulfides and correct folding of the LDL receptor. Mol Cell 50(6): 793-804.
- West, J. D., Stamm, C. E., Brown, H. A., Justice, S. L. and Morano, K. A. (2011). Enhanced toxicity of the protein cross-linkers divinyl sulfone and diethyl acetylenedicarboxylate in comparison to related monofunctional electrophiles. Chem Res Toxicol 24(9): 1457-1459.
Article Information
Copyright
© 2022 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Cobb, D. W., Woods, G. S. and Muralidharan, V. (2022). Activity-based Crosslinking to Identify Substrates of Thioredoxin-domain Proteins in Malaria Parasites. Bio-protocol 12(4): e4322. DOI: 10.21769/BioProtoc.4322.
Category
Microbiology > Microbial biochemistry > Protein > Activity
Biochemistry > Protein > Interaction > Crosslinking
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










