- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Split-luciferase Complementation Imaging Assay to Study Protein-protein Interactions in Nicotiana benthamiana
(*contributed equally to this work) Published: Vol 11, Iss 23, Dec 5, 2021 DOI: 10.21769/BioProtoc.4237 Views: 12196
Reviewed by: José A O’BrienMalgorzata LichockaAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
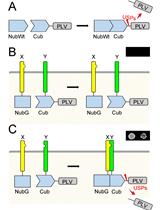
Mating Based Split-ubiquitin Assay for Detection of Protein Interactions
Wijitra Horaruang and Ben Zhang
May 5, 2017 17373 Views
MicroScale Thermophoresis as a Tool to Study Protein-peptide Interactions in the Context of Large Eukaryotic Protein Complexes
Maximilian G. Plach [...] Thomas Schubert
Dec 5, 2017 24662 Views
In vitro Auto- and Substrate-Ubiquitination Assays
Hye Lin Park [...] Gyeong Mee Yoon
Apr 5, 2022 3568 Views
Abstract
The experimental identification of protein-protein interactions (PPIs) is critical to understand protein function. Thus, a plethora of sensitive and versatile approaches have been developed to detect PPIs in vitro or in vivo, such as protein pull-down, yeast two-hybrid (Y2H), co-immunoprecipitation (co-IP), and bimolecular fluorescence complementation (BiFC) assays. The recently established split-luciferase complementation (Split-LUC) imaging assay has several advantages compared to other approaches to detect PPIs in planta: it is a relatively simple and fast method to detect PPIs in vivo; the results are quantitative, with high sensitivity and low background; it measures dynamic PPIs in real-time; and it requires limited experimental materials and instrumentation. In this assay, the amino-terminal and carboxyl-terminal halves of the luciferase enzyme are fused to two proteins of interest (POIs), respectively; the luciferase protein is reconstituted when two POIs interact with each other, giving rise to a measurable activity. Here, we describe a protocol for the Split-LUC imaging assay using a pair of modified gateway-compatible vectors upon Agrobacterium-mediated transient expression in Nicotiana benthamiana. With this setup, we have successfully confirmed a series of interactions among virus-plant proteins, virus-virus proteins, plant-plant proteins, or bacteria-plant proteins in N. benthamiana.
Background
The dynamic formation of complexes is fundamental for proteins to achieve their biological functions in most cellular processes. Therefore, identifying and characterizing protein-protein interactions (PPIs) is crucial to understanding the biological roles of proteins of interest (POIs) in their molecular contexts. Several approaches have been established to detect PPIs (reviewed in Morsy et al., 2008; Xing et al., 2016), each with its intrinsic advantages and disadvantages. For example, the yeast two-hybrid (Y2H) assay is widely used for large-scale screening of interacting proteins, since it is easy to perform, affordable, fast, and has very low space requirements. In addition, the split ubiquitin two-hybrid assay identifies interactions between non-soluble membrane proteins (Thaminy et al., 2004). However, when applied to non-yeast proteins, the Y2H assay detects PPIs in a decontextualized manner, expressing the fusion proteins in a surrogate system. Therefore, interactions involving plant proteins identified in this system need to be further confirmed with other techniques in plant cells and co-existence of the POIs in natural conditions needs to be assessed. Co-immunoprecipitation (co-IP) is used to determine PPIs in vivo and can identify both direct and indirect interactions, but it requires the use of specific antibodies and/or fusion proteins and is a relatively labor-intensive and costly procedure. Bimolecular fluorescence complementation (BiFC) and Förster resonance energy transfer by fluorescence lifetime imaging (FRET-FLIM) assays are useful imaging approaches to identify PPIs in vivo, with the advantage that spatial information regarding PPIs is generated. Nevertheless, both methods require the availability of sophisticated microscopy equipment, which can frequently be a limiting factor.
The split-luciferase complementation (Split-LUC) imaging assay was developed as an alternative method to detect dynamic and likely direct PPIs in plant cells in a fast and sensitive manner (Chen et al., 2008). In this assay, POIs (protein A and protein B) are fused to different halves of the luciferase enzyme (N- or C-terminal fragments, NLuc or CLuc) and co-expressed in plant cells. The luciferase protein is reconstituted only when protein A interacts with protein B, driving the NLuc and CLuc fragments into close spatial proximity, hence restoring enzymatic activity. This enzymatic activity on the substrate luciferin results in emission of light, which can be detected and quantified by a charge-coupled device (CCD) imaging system or a luminometer. Therefore, this assay can not only qualitatively detect PPIs but also allows to measure the dynamic intensity of PPIs in a quantitative manner, enabling comparative studies.
In the initial description of the protocol, the translational fusion of POIs to NLuc or CLuc was performed by traditional restriction and ligation (Chen et al., 2008). To simplify and speed up the cloning procedure, we have generated a Gateway-compatible version of the original Split-LUC vectors (Chen et al., 2008), which we named pGWB-NLuc and pGWB-CLuc destination vectors. Capitalizing on these vectors, Agrobacterium tumefaciens-mediated transient expression in the model Solanaceae species Nicotiana benthamiana, and the Split-LUC method, we have successfully identified multiple PPIs in planta, including interactions between plant proteins, viral proteins and plant proteins, and bacterial effectors and plant proteins (Wang, Y. et al., 2019; Derkacheva et al., 2020; Wang, L. et al., 2020 and 2021; Yu et al., 2020).
This protocol includes five major sections: 1) preparation of plants (N. benthamiana), 2) plasmid construction, 3) Agrobacterium-mediated transient expression, 4) measurement of luciferase activity, and 5) data analysis.
Materials and Reagents
Conical centrifuge tubes (15 ml) (e.g., Thermo ScientificTM NuncTM 15 ml, catalog number: 339650, or similar).
Microcentrifuge tubes (1.5 ml) (e.g., Thermo ScientificTM NuncTM, catalog number: 3451, or similar).
Needleless syringe (1 ml) (e.g., BD® 1 ml Insulin Syringe U-100 Slip Tip with BD PrecisionGlideTM Needle 25G × 5/8 (0.5 mm × 16 mm, catalog number: 329651, or similar)
Plastic Petri dishes (90 mm diameter) (e.g., Thermo ScientificTM SterilinTM Petri dishes 90 mm, catalog number: 101R20, or similar)
Syringe filter (0.45 μm) (e.g., Millex, Millex®-HV, catalog number: SLHV033RB, or similar)
Plastic square Petri dishes (10 × 10 cm) (e.g., Thermo ScientificTM NuncTM Lab-TekTM, catalog number: 4021, or similar)
Pipette tips (e.g., AIBIO, catalog number: T1040000, or similar)
Paper towels
Spectrophotometer plastic cuvettes (e.g., BRAND, catalog number: 759015, or similar)
Standard potting soil (e.g., Pindstrup, catalog number: 1034593214, or similar)
Vermiculite (e.g., Shanghai Chunying Horticulture Co., Ltd, vermiculite 2-4 mm, or https://www.amazon.com/s?k=Professional+Grade+Vermiculite+by+Plantation+Products&ref=nb_sb_noss_2, or similar)
Pots (e.g., https://www.amazon.com/HOMENOTE-Plastic-Outdoor-Planters-Multiple/ dp/B08HCX2D9V/ref=sr_1_42?dchild=1&keywords=flowerpot&qid=1621586733&sr=8-42, or similar)
N. benthamiana plants (see Figure 1)
Escherichia coli (strain DH5α) home-made or purchased competent cells (for commercial competent cells, e.g., Beijing TransGen Biotech, catalog number: CD501, or similar)
A. tumefaciens (strain GV3101) home-made or purchased competent cells (for commercial competent cells, e.g., Weidi Bio, catalog number: AC1001, or similar)
Potassium hydroxide (KOH) (e.g., Sinopharm Chemical Reagent Co, SCR®, catalog number: 10017018, or similar)
pGWB-NLuc vector (Figure 2A, GenBank accession number: MW557526; Addgene: 174050)
pGWB-CLuc vector (Figure 2B, GenBank accession number: MW557527; Addgene: 174051)
pDONRTM 207 (Thermo Scientific, catalog number: PVT11146) or equivalent cloning vector
Gateway® BP reaction kit (Thermo Scientific, catalog number: 11789-013) or equivalent cloning enzymes
Gateway® LR reaction kit (Thermo Scientific, catalog number: 11791100) or equivalent cloning enzymes
Note: Traditional cloning is also possible. The appropriate restriction enzymes and T4 DNA ligase (e.g., Thermo Scientific, catalog number: EL0011, or similar) are required in this case.
Rabbit polyclonal anti-Luciferase (Sigma, catalog number: L0159)
Antibiotics (kanamycin, gentamicin, and rifampicin)
1 mM XenoLight D-Luciferin (PerkinElmer, catalog number: 122799)
75% ethanol (e.g., Sinopharm Chemical Reagent Co, SCR®, catalog number: 801769610, or similar)
Yeast extract (e.g., OXOID, catalog number: LP0021, or similar)
Agar (e.g., Sinopharm Chemical Reagent Co, SCR®, catalog number: 10000561, or similar)
Tryptone (e.g., OXOID, catalog number: LP0042, or similar)
Sodium chloride (NaCl) (e.g., Sinopharm Chemical Reagent Co, SCR®, catalog number: 10019318, or similar)
3’,5’-Dimethoxy-4’-hydroxyacetophenone (AS) (e.g., Sigma-Aldrich, catalog number: 2478-38-8, or similar)
Magnesium chloride hexahydrate (MgCl2·6H2O) (e.g., Sinopharm Chemical Reagent Co, SCR®, catalog number: 10012818, or similar)
MES free acid monohydrate (MES) (e.g., Amresco, catalog number: Amresco E169, or similar)
Dimethyl sulfoxide (DMSO) (e.g., Diamond, catalog number: A100231-0500, or similar)
Distilled sterile water (e.g., Thermo ScientificTM GibcoTM, catalog number: 15230204, or similar)
Luria-Bertani (LB) medium (see Recipes)
Agrobacterium infiltration buffer (see Recipes)
Equipment
Scissors
Nanodrop spectrophotometer (e.g., Thermo Scientific, model: NanoDrop 2000c, or similar)
Note: This is not essential – any other method to quantify DNA (e.g., in a regular spectrophotometer or by visualization in an agarose gel) is also valid.
Vortexer (e.g., Scientific Industries, model: Vortex-Genie 2, catalog number: S1-0246, or similar)
Petri dish incubator at 28°C (e.g., Panasonic, model: MIR-262-PC, or similar)
Petri dish incubator at 37°C (e.g., Panasonic, model: MIR-154-PC, or similar)
Autoclave (e.g., SANYO, model: MLS-3780, or similar)
Flow hood (clean bench) (e.g., Shanghai Shangjing Co., mode: CA-1390-1, or similar)
Plant growth space (e.g., Percival Growth Chamber, model: I-36VL; 24°C, 16-h-light/8-h-dark, or similar)
Electronic balance (e.g., Sartorius, model: BSA224S, or similar)
Centrifuge (e.g., Eppendorf, model: centrifuge 5424, or similar)
pH meter (e.g., Sartorius, model: PB-10, or similar)
Tube incubator (shaker) at 28°C (e.g., Eppendorf, New BrunswickTM, catalog number: I26 M1324-0006, or similar)
Tube incubator (shaker) at 37°C (e.g., Eppendorf, New BrunswickTM, catalog number: I26 M1324-0006, or similar)
CCD imaging apparatus (e.g., NightShade LB985, Berthold Technologies, or similar)
Software
IndiGO software
Graphpad Prism 7 (GraphPad, http://www.graphpad.com), or equivalent program for statistical analysis
Text processor (e.g., Microsoft Word, or similar)
Spreadsheet (e.g., Microsoft Excel, or similar)
Procedure
Preparation of N. benthamiana plants (4 weeks)
Sow N. benthamiana seeds in a 1:1 mix of potting soil and vermiculite, cover the pot with plastic film, and then plant them in a controlled growth chamber under long day conditions (16-h light/ 8-h dark) at 25°C with 70% humidity and a light intensity of 130 mE m-2s-1 (Figure 1).
Transfer 10-day-old seedlings into individual pots (e.g., 10 × 10 cm), and let them grow until they are 4 weeks old (Figure 1).
Note: Water plants and control pests as needed to keep plants healthy. Fertilizer can be added if necessary. Healthy plant material is essential for efficient transient expression of the POIs, and hence for the success of the experiment.

Figure 1. Cultivation of N. benthamiana plants for transient expression experiments. Arrows indicate leaves suitable for infiltration. The age of the plants used is indicated.
Vector construction
Gateway-compatible binary vectors (pGWB-NLuc and pGWB-CLuc) were generated from the original pCAMBIA1300-NLuc/CLuc vectors (Chen et al., 2008) by integrating the Gateway cassette (Figure 2) (Wang, Y. et al., 2019; Yu et al., 2020). To generate the constructs that encode the fusion POIs, these vectors can be used in combination with Gateway cloning (alternatively, the original vectors can be used in combination with traditional cloning). The Gateway cloning procedure is outlined below.
Clone the genes encoding the POIs into a Gateway entry vector (e.g., pDONRTM 207, Thermo Scientific) through a BP reaction (Thermo Scientific, GatewayTM BP ClonaseTM Enzyme mix) following the manufacturer’s instructions (www.thermofisher.cn) and transform each reaction solution into E. coli competent cells. Perform colony PCR to select positive clones and confirm them by sequencing.
Subclone the cloned genes into the destination vectors, pGWB-NLuc and pGWB-CLuc, through an LR reaction (Thermo Scientific, GatewayTM LR ClonaseTM Enzyme mix), following the manufacturer’s instructions (www.thermofisher.cn), and transform each reaction solution into E. coli competent cells. Perform colony PCR to select positive clones and confirm them by sequencing.
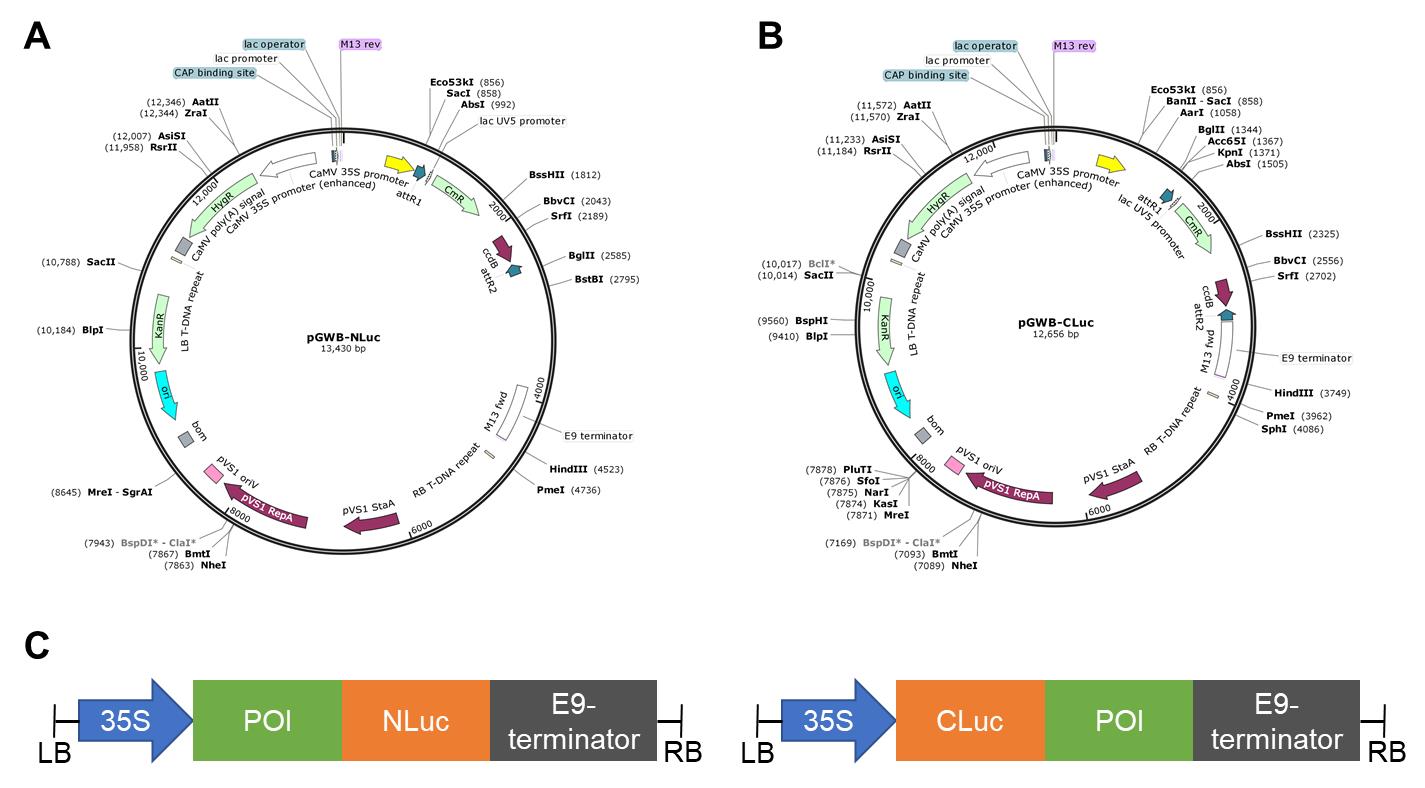
Figure 2. Gateway-compatible vectors for Split-LUC imaging assay in plants. Schematic diagrams showing pGWB-NLuc (A) and pGWB-CLuc (B) vectors (visualized in Snapgene). The size of the plasmids in bp is indicated. Promoters, terminators, open reading frames, attR1, attR2, and T-DNA are shown in the maps. CaMV: Cauliflower mosaic virus; LB: left border; RB: right border; ccdB: Control of Cell Death B (gene); HygR: Hygromycin resistance (gene); KanR: Kanamycin resistance (gene); CmR: Chloramphenicol resistance (gene). (C) Schematic representation of the cassettes to express the fusion proteins. POI, protein of interest.
Agrobacterium-mediated transient expression in N. benthamiana
Prepare and autoclave LB medium (103 kPa/121°C/15 min). Cool down LB medium with agar until the bottle can be touched with bare hands (but is still in liquid form), add the appropriate antibiotics (50 μg/ml kanamycin, 50 μg/ml gentamicin, and 25 μg/ml rifampicin), and pour the medium into Petri dishes.
Transform the plasmids generated in B (Vector construction), as well as appropriate controls, into A. tumefaciens cells by chemical transformation. Competent A. tumefaciens cells (GV3101) can be purchased (see Materials and Reagents above) or prepared using standard protocols (e.g., Kaman-Toth et al., 2018).
Perform colony PCR with specific primers to detect the positive clones. Inoculate one positive single A. tumefaciens colony from each plate into 5 ml of LB liquid medium containing antibiotics (50 µg/ml of kanamycin, 25 µg/ml of rifampicin, and 50 µg/ml of gentamicin) in a 15 ml tube, with shaking at 220 × g at 28°C for 16-24 h until OD600 = 0.5-1.5.
Note: The volume depends on the number of leaves to be inoculated. Usually, A. tumefaciens cells cultured from 5 ml LB medium should be enough for the later inoculation of at least three N. benthamiana leaves.
Optional step: Make a glycerol stock for individual A. tumefaciens clones (to a final 30% glycerol concentration) in a tube (e.g., 1.5 ml microcentrifuge tube) and store at -80°C. Before using, streak out the A. tumefaciens clones carrying the desired plasmids on solid LB medium with the corresponding antibiotics (50 µg/ ml of kanamycin, 25 µg/ml of rifampicin, and 50 µg/ml of gentamicin), and incubate them at 28°C for 24-48 h. Grow a liquid culture of each A. tumefaciens clone at 28°C for 12-16 h to an OD600 = 0.5-1.5. If this step is not needed, proceed to step 5.
Centrifuge the bacteria at 4,000 × g, room temperature, for 10 min and discard the supernatant.
Note: The supernatant should be removed completely, so the use of a micropipette is advised.
Resuspend the A. tumefaciens cells in the infiltration buffer (see Recipe 1) to a final concentration of OD600 = 0.1-0.5.
Note: The final OD600 is important for transient transformation. The optimal OD should be experimentally determined.
Mix equal volumes of A. tumefaciens clones carrying the desired plasmids to express protein A-NLuc or CLuc-protein B to prepare the infiltration solution (Figure 3A). Appropriate controls (both positive and negative) are required.
Note: To rule out nonspecific interactions and artifactual signals, the protein used as negative interaction control should have the same subcellular localization as the POIs.

Figure 3. Illustration of the infiltration process.Keep the infiltration solution at room temperature and in the dark for 2-6 h.
Note: During the incubation, acetosyringone from the infiltration buffer induces the expression of the virulence genes of A. tumefaciens, thus facilitating the transfer and incorporation of the T-DNA region into the host plant genome. This step, therefore, is advisable for effective transformation.
Select fully expanded leaves of 4-week-old N. benthamiana plants for infiltration (Figure 1, Figure 3B).
Infiltrate the A. tumefaciens inoculum carefully into the selected leaves using a 1 ml needleless syringe. Infiltration is performed on the abaxial side (Figure 3B), and pressure is applied with the fingertip on the adaxial side (Figure 3C).
Notes:
Infiltration of N. benthamiana works better in dry leaves without water droplets from condensation or contact with other leaves. On the abaxial side of the leaf, pricking softly with a needle (e.g., from a 1 ml syringe) helps ease infiltration.
To avoid damaging plant leaves, apply the infiltration slowly without too much pressure.
Choose healthy leaves of similar size. At least three independent leaves from different plants need to be used for biological replicates.
Infiltrate the A. tumefaciens inoculum carrying POIs and negative control side-by-side in the same leaf (Figure 3D) to avoid leaf-to-leaf variation.
Note: Use 75% ethanol to clean your gloves before moving to another inoculum during infiltration to avoid cross-contamination.
Use a permanent marker pen to label the infiltrated area (Figure 3E). Gently wipe the wet surface of the leaf with tissue paper.
Place the plants in the growth chamber for 36-48 h (e.g., 2 days) before measuring luciferase activity. Accumulation of the transiently expressed proteins can usually be detected at 36-60 h post inoculation (hpi).
Measurement of luciferase activity with a CCD imaging system
Infiltrate 1 mM luciferin solution (in H2O) into the inoculated leaves 2 days after A. tumefaciens infiltration.
Detach the whole leaves and keep them in the dark for 10 min before imaging to avoid interference by chlorophyll luminescence.
Capture luminescence images using a CCD imaging apparatus (e.g., NightShade LB985) with an IndiGO software and measure the relative luciferase activity (luminescence intensity/cm2 leaf area) following the manufacturer’s instructions as shown in Figure 4.
Notes:
Since the luminescence will be low, the camera must be cooled down to the default temperature, and luminescence images must be captured in the dark. Adjust the exposure time (Figure 4E) according to the signal intensity.
Since the detection of luminescence relies on the abundance of reconstituted luciferase protein, the accumulation of all tested proteins and controls must be assessed by western blotting with anti-luciferase antibodies (Rabbit polyclonal anti-Luciferase, Sigma, Cat# L0159), which recognize both NLuc and CLuc.
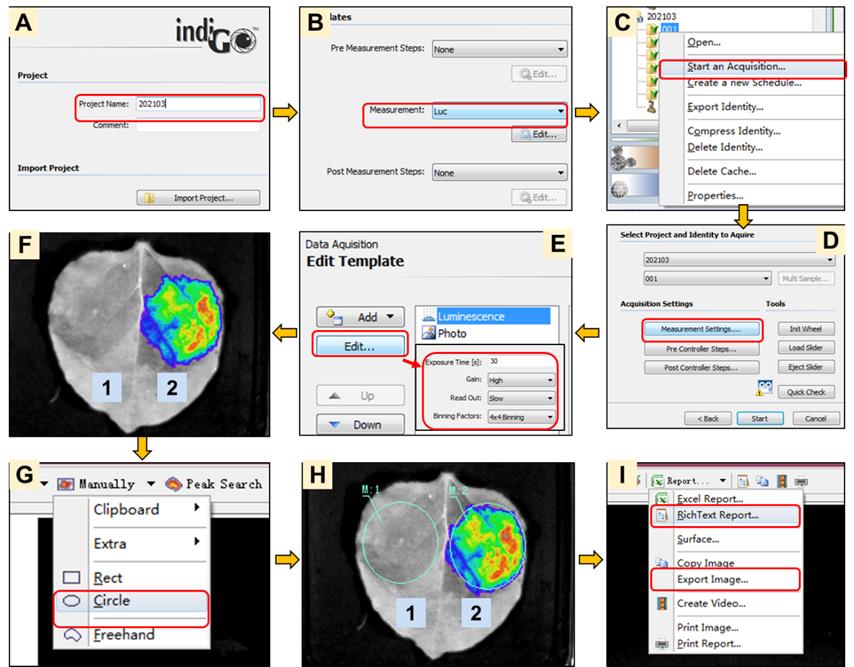
Figure 4. Illustration of the measurement of the relative luciferase activity in Split-LUC imaging assays. The example shows the self-interaction of the viral protein V2, encoded by the geminivirus tomato yellow leaf curl virus (TYLCV, GenBank accession number: AJ489258). “1” indicates the area co-expressing V2-NLuc and CLuc-SlWRKY75 (negative control); “2” indicates the area co-expressing V2-NLuc and CLuc-V2 (experiment).
Export the images (.png format) of the Split-LUC imaging assay from the IndiGO software by clicking “Export Image” (Figure 4I).
Export the values of the relative luciferase activity in the defined regions of interest (ROI) for all samples and controls from the IndiGO software by clicking “RichText Report” (Figure 4I). Open the RichText Report (.rtf) file with a text processor (e.g., the Microsoft Word program) and copy the analysis details of the relative luciferase activity to a spreadsheet (e.g., Excel) (Table 1).
Note: The average value of the relative luciferase activity in the ROI is equal to the total signal value of the ROI (Overall) divided by the number of pixels.
Table 1. Example of the values of relative luciferase activity per ROI from the experiment depicted in Figure 4H
Cps: counts per secondArea Details Area Label Area [mm2] Border [mm] Width [mm] Height [mm] Pixel Background ROI 1 208.89 51.29 15.68 16.96 9642 No ROI 2 221.94 53 15.68 18.02 10213 No Analysis Details Area Label Overall [cps] Average [cps] Min [cps] Max [cps] Variance [cps] Deviation [cps] ROI 1 48669 5.05 2 9 729.2 27 ROI 2 1242420 121.65 1 377 5191470.29 2278.48
Data analysis
An example of PPI detected by Split-LUC imaging assay in N. benthamiana is shown in Figure 5.
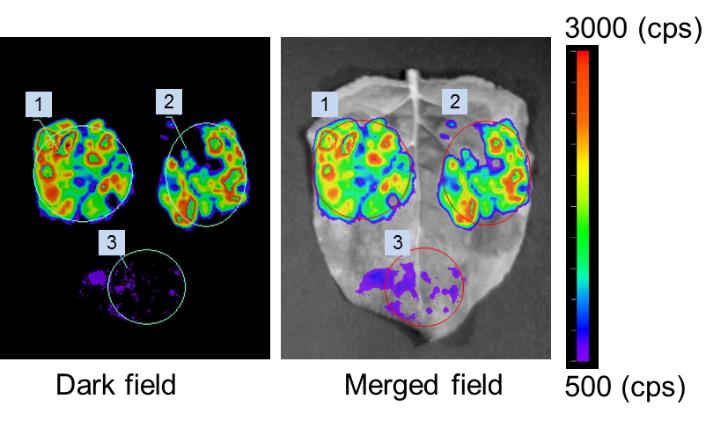
Figure 5. Example PPI detected by Split-LUC imaging assay: the viral protein V2 from TYLCV interacts with Argonaute 4 (AGO4) proteins from N. benthamiana. “1” indicates the area co-expressing V2-NLuc and CLuc-NbAGO4-1 (experiment); “2” indicates the area co-expressing V2-NLuc and CLuc-NbAGO4-2 (experiment); “3” indicates the area co-expressing V2-NLuc and CLuc-SlWRKY75 (negative control). The luciferase bioluminescence from N. benthamiana leaves was imaged 2 days after infiltration. An image of a representative leaf is shown. Cps: counts per second.Table 2. Example of average values (cps) of relative luciferase activity of ROIs from the experiment depicted in Figure 5 with all biological replicates in N. benthamiana leaves
V2-NLuc + CLuc-NbAGO4-1 V2-NLuc + CLuc-NbAGO4-2 V2-NLuc + CLuc-SlWRKY75 Replicate 1 6.47 5.62 0.85 Replicate 2 3.90 3.29 0.85 Replicate 3 6.90 6.61 0.59 Replicate 4 3.87 4.06 0.58 Copy the average values (cps) of relative luciferase activity of ROIs from the experiment (Figure 5) with all biological replicates (Table 2) and transfer them to an appropriate program (e.g., GraphPad Prism7) for data representation and statistical analysis (Figure 6). Perform the appropriate statistical analyses to compare POIs to their negative control (Figure 6).
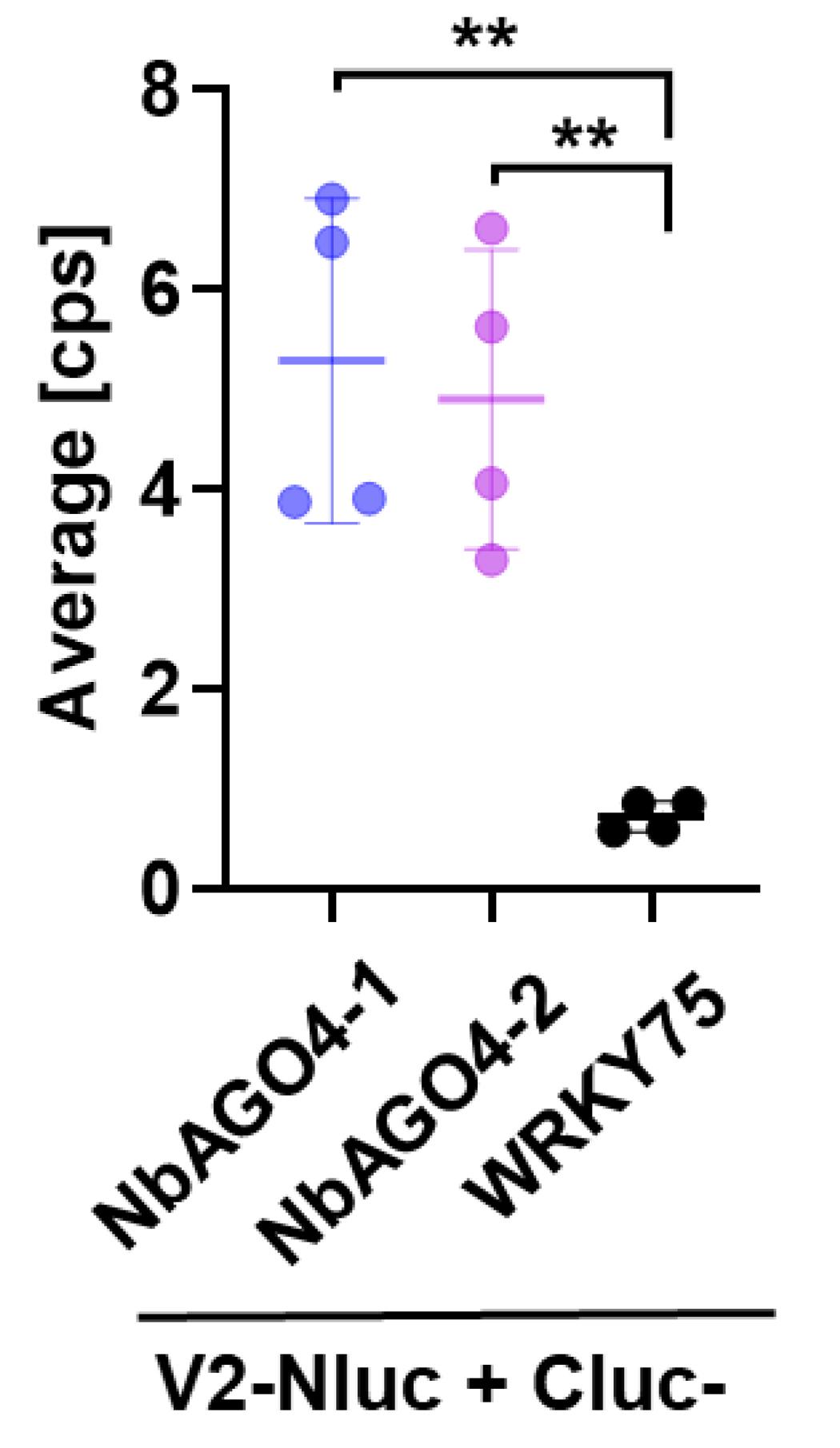
Figure 6. NbAGO4-1 and NbAGO4-2 interact with V2 in Split-LUC assays. Average values of relative luciferase activity of ROIs in N. benthamiana leaves co-expressing V2-NLuc and CLuc-NbAGO4-1 (experiment), or CLuc-NbAGO4-2 (experiment), or CLuc-SlWRKY75 (negative control) are shown. Error bars represent the standard deviation from the mean of four biological replicates. Asterisks indicate a statistically significant difference (according to One Way ANOVA with Dunnett's multiple comparison, **: P < 0.01) between experiment samples and the negative control.
Recipes
Agrobacterium infiltration buffer
1 M MES (pH 5.7) 100 μl
1 M MgCl2 100 μl
150 mM acetosyringone 10 μl
Add 9.890 ml of water to reach 10 ml
Adjust MES to pH 5.7 with KOH.
Dissolve MES and MgCl2 in sterile water and filter each solution with a 0.2 μm syringe filter to avoid contamination. Store this stock solution at 4°C. Dissolve acetosyringone in DMSO, aliquot, and store this stock solution at -20°C.
LB medium
10 g Tryptone
5 g NaCl
5 g Yeast extract
Adjust pH to 7.5 with NaOH, and add water to reach 1 L. To prepare solid medium, add 15 g agar to 1 L of medium and autoclave.
Acknowledgments
We thank Yuanzheng Wang for his help on generation of the Gateway-compatible vectors (pGWB-NLuc and pGWB-CLuc), Xinyu Jian for technical and administrative assistance, Aurora Luque for technical assistance during this work, and all the members of the Lozano-Duran and Macho laboratories for the constant improvement of this protocol and helpful discussions. This work is supported by the Strategic Priority Research Program of the Chinese Academy of Sciences (grants XDB27040204 and XDB27040206) and the Shanghai Center for Plant Stress Biology (Chinese Academy of Sciences). The original paper in which this protocol was used is Wang, L. et al. (2020; DOI: 10.7554/eLife.55542).
Competing interests
The authors have no competing interests to declare.
Ethics
No human or animal subjects are used in this protocol.
References
- Chen, H., Zou, Y., Shang, Y., Lin, H., Wang, Y., Cai, R., Tang, X. and Zhou, J. M. (2008). Firefly luciferase complementation imaging assay for protein-protein interactions in plants. Plant Physiol 146(2): 368-376.
- Derkacheva, M., Yu, G., Rufian, J. S., Jiang, S., Derbyshire, P., Morcillo, R. J. L., Stransfeld, L., Wei, Y., Menke, F. L. H., Zipfel, C. and Macho, A. P. (2020). The Arabidopsis E3 ubiquitin ligase PUB4 regulates BIK1 homeostasis and is targeted by a bacterial type-III effector. bioRxiv DOI. 10.1101/2020.10.25.354514.
- Kaman-Toth, E., Pogany, M., Danko, T., Szatmari, A. and Bozso, Z. (2018). A simplified and efficient Agrobacterium tumefaciens electroporation method. 3 Biotech 8(3): 148.
- Morsy, M., Gouthu, S., Orchard, S., Thorneycroft, D., Harper, J. F., Mittler, R. and Cushman, J. C. (2008). Charting plant interactomes: possibilities and challenges. Trends Plant Sci 13(4): 183-191.
- Thaminy, S., Miller, J., and Stagljar, I. (2004). The split-ubiquitin membrane-based yeast two-hybrid system. In: Fu, H. (Ed.). Protein-Protein Interactions. Methods Mol Biol 261: 297-312.
- Wang, L., Ding, Y., He, L., Zhang, G., Zhu, J. K. and Lozano-Duran, R. (2020). A virus-encoded protein suppresses methylation of the viral genome through its interaction with AGO4 in the Cajal body. Elife 9: e55542.
- Wang L., Tan H., Medina-Puche L., Wu M., Gómez B. G., Gao M., Shi C. N., Jimenez-Góngora T., Fan P., Ding X., Zhang D., Yi D., Rosas-Diaz T., Liu Y., Aguilar E., Fu X. and Lozano-Durán R. (2021). Combinatorial interactions between viral proteins expand the functional landscape of the viral proteome. bioRxiv: DOI.10.1101/2021.04.07.438767.
- Wang, Y., Li, Y., Rosas-Diaz, T., Caceres-Moreno, C., Lozano-Duran, R. and Macho, A. P. (2019). The IMMUNE-ASSOCIATED NUCLEOTIDE-BINDING 9 protein is a regulator of basal immunity in Arabidopsis thaliana. Mol Plant Microbe Interact 32(1): 65-75.
- Xing, S., Wallmeroth, N., Berendzen, K. W. and Grefen, C. (2016). Techniques for the analysis of protein-protein interactions in vivo. Plant Physiol 171(2): 727-758.
- Yu, G., Xian, L., Xue, H., Yu, W., Rufian, J. S., Sang, Y., Morcillo, R. J. L., Wang, Y. and Macho, A. P. (2020). A bacterial effector protein prevents MAPK-mediated phosphorylation of SGT1 to suppress plant immunity. PLoS Pathog 16(9): e1008933.
Article Information
Copyright
Wang et al. This article is distributed under the terms of the Creative Commons Attribution License (CC BY 4.0).
How to cite
Readers should cite both the Bio-protocol article and the original research article where this protocol was used:
- Wang, L., Yu, G., Macho, A. P. and Lozano-Durán, R. (2021). Split-luciferase Complementation Imaging Assay to Study Protein-protein Interactions in Nicotiana benthamiana . Bio-protocol 11(23): e4237. DOI: 10.21769/BioProtoc.4237.
- Wang, L., Ding, Y., He, L., Zhang, G., Zhu, J. K. and Lozano-Duran, R. (2020). A virus-encoded protein suppresses methylation of the viral genome through its interaction with AGO4 in the Cajal body. Elife 9: e55542.
Category
Plant Science > Plant molecular biology > Protein
Biochemistry > Protein > Interaction > Protein-protein interaction
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link











