- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Imaging of Lipid Uptake in Arabidopsis Seedlings Utilizing Fluorescent Lipids and Confocal Microscopy
Published: Vol 11, Iss 22, Nov 20, 2021 DOI: 10.21769/BioProtoc.4228 Views: 4064
Reviewed by: Khyati Hitesh ShahAksiniya AsenovaAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

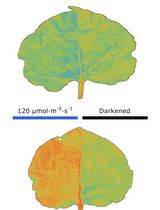
Live-Cell Monitoring of Piecemeal Chloroplast Autophagy
Masanori Izumi [...] Shinya Hagihara
Nov 5, 2025 1725 Views
Chloroplast Movement Imaging Under Different Light Regimes With a Hyperspectral Camera
Paweł Hermanowicz [...] Justyna Łabuz
Dec 20, 2025 794 Views
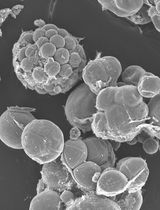
A Simple Protocol for Periodic Live Cell Observation of Flagellate Stages in the Lichen Alga Trebouxia
Enrico Boccato [...] Mauro Tretiach
Jan 20, 2026 228 Views
Abstract
Eukaryotic cells use a diverse set of transporters to control the movement of lipids across their plasma membrane, which drastically affects membrane properties. Various tools and techniques to analyze the activity of these transporters have been developed. Among them, assays based on fluorescent phospholipid probes are particularly suitable, allowing for imaging and quantification of lipid internalization in living cells. Classically, these assays have been applied to yeast and animal cells. Here, we describe the adaptation of this powerful approach to characterize lipid internalization in plant roots and aerial tissues using confocal imaging.
Graphic abstract:
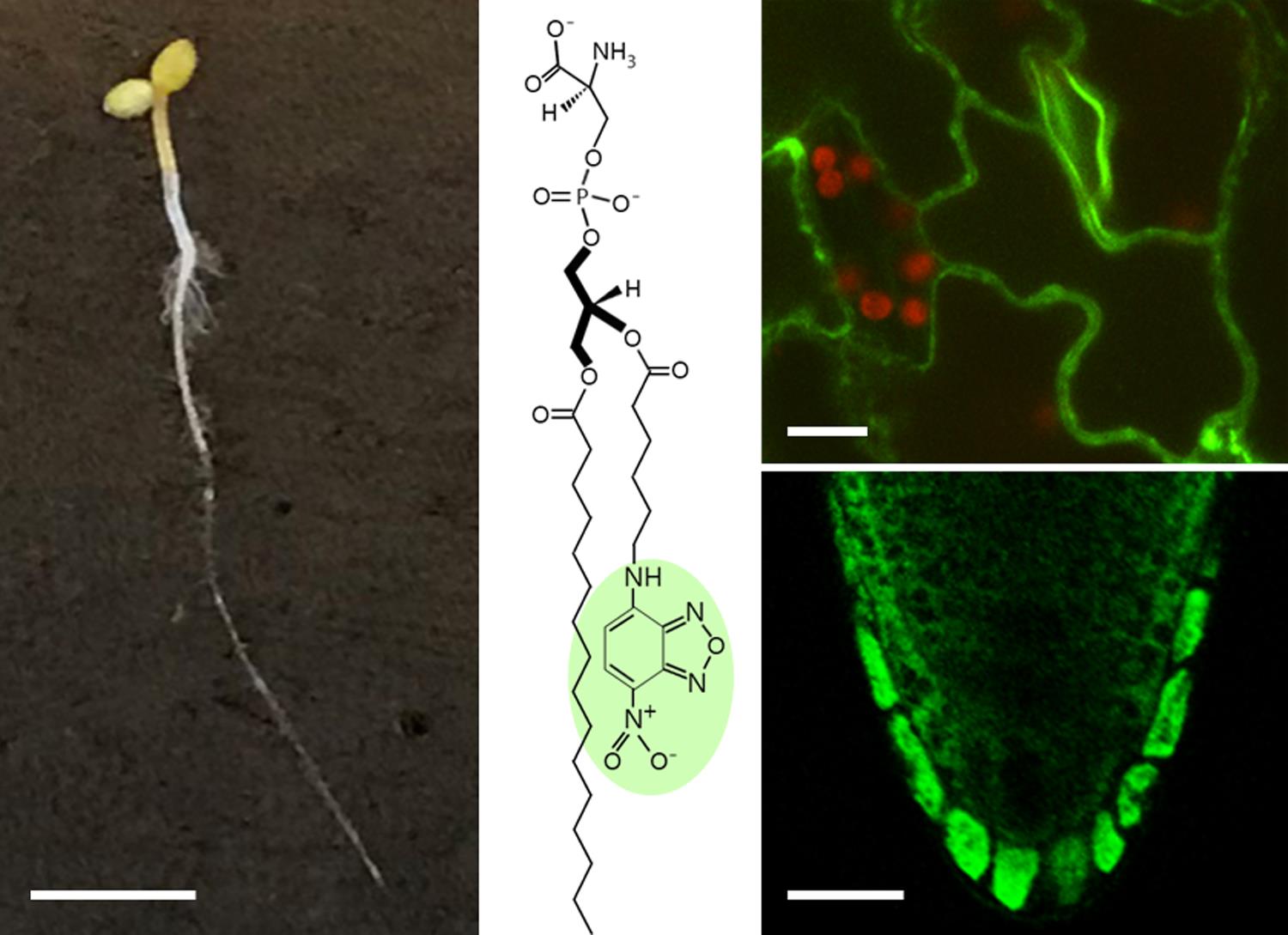
Fluorescent lipid uptake in Arabidopsis seedlings. Scale bars: seedling, 25 mm; leaf, 10 μm; root, 25 μm.
Background
In eukaryotic cells, movement of lipids across biological membranes (known as lipid flip-flop) is regulated by a diverse set of membrane transporters that can be classified into two categories: (i) ATP-independent transporters, also called scramblases, that facilitate a rapid bi-directional movement of lipids without metabolic energy input, and (ii) ATP-driven vectorial transporters that actively translocate lipids from one membrane leaflet to the other, often with high specificity. The latter group comprises ATP-dependent flippases and floppases, which catalyze the inward movement of lipids to the cytoplasmic membrane leaflet, and the outward movement to the extracellular/luminal side, respectively. A subgroup of P-type ATPases, the P4 ATPases, has emerged as a major group of lipid flippases that form heterodimeric complexes with members of the Cdc50 (cell division control 50) protein family (reviewed in Lopez-Marques et al., 2014; Andersen et al., 2016). While initially characterized as aminophospholipid flipases, recent studies of individual family members from fungi, plants, and animals show that P4 ATPases differ in their substrate specificities and mediate transport of a broader range of lipid substrates, including lysophospholipids, synthetic alkylphospholipids, and sugar-modified ceramides (Roland et al., 2019; Shin and Takatsu, 2019).
Quantitative assessment of P4 ATPase lipid transport activity is essential for the determination of substrate specificities and to establish whether and how this activity is regulated in the living cell. As P4-ATPases are trapped in an environment (cellular membranes) formed by their own substrate (lipids), analyzing their activity is not a trivial task, and most assays are based on the use of fluorescent lipid analogs, typically nitrobenzoxadiazol (NBD)-labeled lipids. These analogs have a fluorescent reporter group attached to a short-chain fatty acid (C6) and maintain most of the properties of endogenous phospholipids, except that they are more water-soluble, which facilitates incorporation from the medium into the outer monolayer of the plasma membrane.
Traditionally, lipid uptake assays employing NBD-lipids are carried out after heterologous expression of the plant P4 ATPases in yeast strains devoid of their endogenous lipid transporters (for a review, see Nintemann et al., 2019). However, many plant P4 ATPases express poorly, fail to fold, and/or traffic improperly when produced in heterologous systems. In addition, successful expression does not always result in active lipid translocation, probably due to the absence of plant-specific accessory proteins and/or cofactors required for the functioning of plant lipid transporters (McDowell et al., 2015).
The method presented here utilizes C6-NBD-lipids to study lipid internalization in intact plants, exemplified on Arabidopsis seedlings as a model. Small 5-day old seedlings are grown on plates under sterile conditions and then transferred to liquid growth medium supplemented with C6-NBD-lipids. After incubation for the desired time, the seedlings are washed with medium containing small amounts of a specific detergent to remove excess lipids attached to the cell walls. Finally, plants are visualized using confocal microscopy, and the data quantified using imaging software. This protocol can be used to characterize lipid internalization both in roots and aerial tissues and can be easily adapted to other plant species. For experiments in roots, growth of seedlings on agar plates is preferred to regular cultivation on soil, as removal from the soil causes damage to the root surface.
Materials and Reagents
Materials
Arabidopsis seeds (ecotype Col-0 or as desired)
Pipette tips PIPETMAN DIAMOND D10, D200, D1000 (Gilson, catalog numbers: F161630, F161930, F161670)
2-ml microcentrifuge snap-cap tubes with round bottom (e.g., BRAND, microcentrifuge tube, 2 ml with lid, PP; Merck, catalog number: BR780546-500EA)
Circular holder for 2-ml microcentrifuge tubes (e.g., PrepSafeTM microcentrifuge tube mini floating rack, clear; Merck, catalog number: Z756385)
50-ml glass beaker (e.g., BRAND, catalog number: 91217)
Square Petri dishes 120 × 120 × 17 mm, Greiner Bio-One (Fisher Scientific, catalog number: 07-000-330)
Micropore tape (3M, catalog number: 1530-1)
Aluminum foil (e.g., Sigma, catalog number: Z185140-1EA)
Centrifuge glasses DURAN® with conical bottom, 12 ml (Carl Roth, catalog number: K211.1)
25-μl calibrated glass syringe (model 702 N; Hamilton, catalog number: CAL80400)
1.5 ml screw amber glass vials with 8-mmTeflon-lined screw caps (e.g., VWR, catalog numbers: VWRI548-0019 and 548-0360)
Corning® Pasteur pipettes, non-sterile, 228 mm (Merck, catalog number: CLS7095B9)
PARAFILM® M (Merck, catalog number: P7793)
Clear glass jars with snap-cap, 11 ml, 22 × 45 mm (VWR, catalog number: 548-0625)
For aerial tissues, microscope slides 76 × 26 × 1 mm with cut edges (Histolab, catalog number: 06300)
For root tissues, diagnostic microscope slides 25 × 75 mm, 8 Wells of 6 mm, with black epoxy field around cavities (Histolab, catalog number: 06260)
Cover glass No. 1, 18 × 24 mm and 24 × 24 mm (Histolab, catalog numbers: 06602 and 06608)
Reagents
All reagents can be stored at room temperature, except NBD-phospholipid solutions, which should be kept at -20°C for long-term storage.
Sodium hypochlorite (14% Cl2) in aqueous solution, GPR RECTAPUR® (VWR Chemicals, catalog number: 27900.296)
37% hydrochloric acid solution (Sigma, catalog number: 320331)
Phytoagar (Duchefa Biochemie, catalog number: P1003)
2-(N-morpholino)ethanesulfonic acid (MES) (Merck-Millipore, catalog number: 1061261000; CAS number: 4432-31-9)
Potassium hydroxide (Sigma, catalog number: 484016-1KG; CAS number: 1310-58-3)
Dimethyl Sulfoxide (DMSO), sterile-filtered, BioPerformance Certified (Sigma, catalog number: D2438)
Methanol (Sigma, catalog number: 179337; CAS number: 67-56-1)
Chloroform, ethanol-stabilized and certified for absence of phosgene and HCl (Sigma, catalog number: 650471)
Fluorescent C6-NBD-phospholipids in chloroform
C6-NBD-phosphatidylethanolamine (Avanti Polar Lipids, catalog number: 810153)
C6-NBD-phosphatidylserine (Avanti Polar Lipids, catalog number: 810192)
C6-NBD-phosphatidylcholine (Avanti Polar Lipids, catalog number: 810130)
NBD-lysophosphatidylcholine (Avanti Polar Lipids, catalog number: 810128)
C6-NBD sphingomyelin (Avanti Polar Lipids, catalog number: 810218)
Tergitol solution type NP-40 (Sigma, catalog number: NP40S)
Low-melting-point agarose, analytical grade (Promega Corporation, catalog number: V2111)
Murashige and Skoog (MS) salts with vitamins (Phyto Technology Laboratories, catalog number: M519)
Half strength MS liquid medium (see Recipes)
Half strength MS plates (see Recipes)
Agarose solution (see Recipes)
C6-NBD-lipid stocks (see Recipes)
Equipment
Pipettes PIPETMAN Classic P2, P20, P200, P1000 (Gilson, models: F144801, F123600, F123601, F123602)
Precision tweezers Style #5, fine needle-sharp, anti-magnetic stainless steel (Merck, catalog number: T4537)
Scalpel (e.g., Sigma, catalog numbers: S2646 and S2896)
Glass dessicator (e.g., Boro 3.3 dessicator 20 cm with knob lid; BRAND, catalog number: 65038)
Shallow water bath (e.g., Precision GP 2S, 2L shallow water bath; ThermoFisher, catalog number: TSGP2S)
Incubator or heating block at 60°C (e.g., VWR, catalog number: 75838-270)
Analytical balance (e.g., Sartorius Entris-i II, 220 g/0.1 mg, Buch Holm, catalog number: 4669128)
Autoclave sterilizer (e.g., Presoclave III, 80 liters, Ø: 40 × 62 cm, Buch Holm, catalog number: 5083042)
Freezer (e.g., GRAM Bioline, model: BioCompact 210RF)
Refrigerator (e.g., GRAM Bioline, model: BioCompact 210RR)
Rotary evaporator equipped with vacuum pump or nitrogen gas supply [e.g., Büchi® Rotavapor® RII evaporator with jack and water bath (Sigma, catalog number: Z564036), equipped with a Vacuubrand diaphragm vacuum pump model MD1C (Sigma, catalog number: Z656194)]
Water purification systems (e.g., Milli-Q® Direct water purification system, Merck-Millipore, catalog number: ZR0Q008WW)
Fume hood (e.g., ErlabTM Captair 391 Smart Fume Hood, Fisher Scientific, catalog number: 15514360)
Microwave oven (e.g., H2100 Microwave Oven 220 Volt, Merck, catalog number: A9209)
Laminar flow cabinet (e.g., Fortuna Clean Bench, ScanLaf, Labogene)
Plant growth chamber (e.g., Sanyo Versatile Environmental Test Chamber, model: MLR-351H)
Confocal microscope, e.g., Leica TCS SP5 Confocal Laser Scanning Microscope equipped with an argon laser (Leica Microsystems A/S) and a C-Apochromat 63×/1.2 W autocorr M27 (CG=0.14-0.19 mm) (FWD=0.28 mm at CG=0.17 mm) objective (Carl Zeiss Microscopy, catalog number: 421787-9971-790)
Software
Leica LAS X software, version 3.5.7.23225 (Leica Microsystems A/S)
ImageJ 1.52n (National Institutes of Health, USA, http://imagej.nih.gov/ij) (Schneider et al., 2012)
Procedure
Prepare plant growth medium
Prepare ½ MS media plates (see Recipes) considering that one plate will be needed for every 20-25 seeds. For each lipid, uptake should be quantified for at least five seedlings per experiment. Allow some extra seeds to account for possible damaged seedlings along the procedure.
Note: Plates can be made in advance and kept at 4°C for about a month in a sterile bag.
Seed sterilization and plant growth conditions
We sterilize our seeds using chlorine vapor inside a desiccator, but the protocol can be used with any other sterilization procedure.
Transfer seeds to 2-ml microcentrifuge tubes (maximum of 50 seeds/tube) marked with a pencil or chlorine-resistant pen and place in a circular rack.
Place the rack with the seed-containing tubes (with lids open) and a 50-ml beaker with 25 ml of 14% hypochlorite solution into a desiccator jar placed in a fume hood (Figure 1A and 1B).
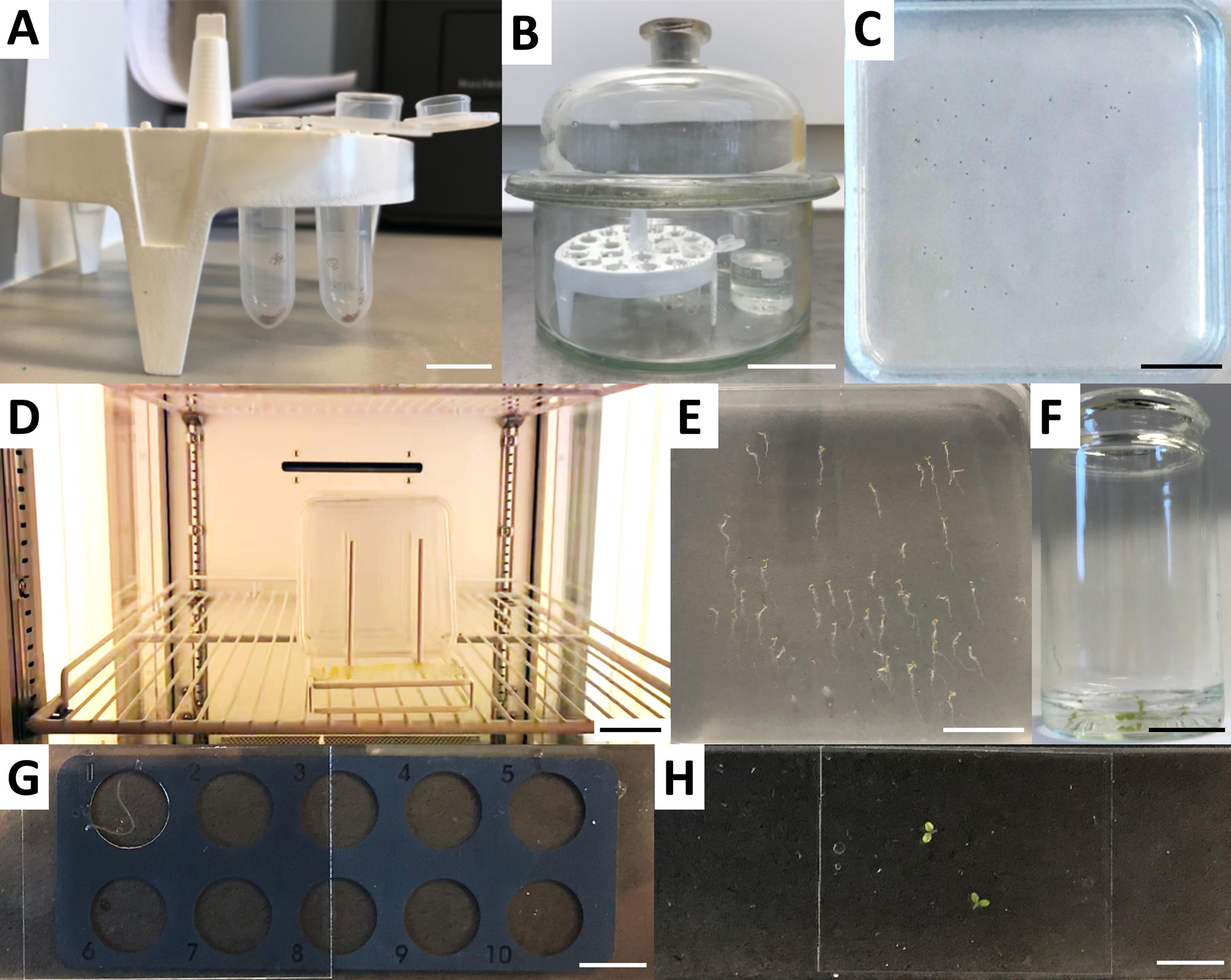

Figure 1. Processing of plant samples in lipid uptake assays. Seeds are placed into open 2-ml round bottom tubes (A) and taken to a desiccator jar together with 25 ml of sodium hypochlorite solution in a 50-ml glass beaker (B). Sterilization by chlorine gas is triggered by addition of 37% hydrochloric acid. (C) Sterile seeds are plated on square ½ MS plates and sealed with micropore 3M tape. (D) After stratification, plates are placed vertically in a growth chamber under a long-day regime. (E) Seedlings are ready for lipid uptake assays after five days of growth. (F) Seedlings are transferred to small glass jars containing ½ MS media and placed at 25°C. Lipid uptake assays start with addition of the desired NBD-lipid. (G-H) After washing, roots (G) or aerial parts (H) are transferred to microscope slides for visualization. Scale bars: A, F, and H, 1 cm; B and D, 5 cm; C and E, 2.5 cm.Carefully add 1 ml of 37% hydrochloric acid solution to the hypochlorite solution and immediately close the desiccator jar.
Allow sterilization by chlorine fumes to proceed for a period of approximately 3 to 4 h before opening the container and immediately closing the tube lids as fast as possible to preserve sterility.
Note: Sterilization periods longer than 4 h reduce seed viability.
Take the tubes with seeds to a sterile bench and plate on square ½ MS plates (see Recipes), trying to cover the whole surface so that seedlings do not touch each other as they grow (Figure 1C).
Close the plates with micropore tape and stratify for 2 to 4 days at 4°C before placing them vertically in a plant growth chamber at 22°C under a 16 h light/8 h dark light regime for 5 days (Figure 1D-1E).
Note: Lipid uptake seems to be poor in plants coming directly from the dark period. Allow the seedlings to stand in the light for at least 90-120 min before starting the experiment.
NBD-lipid uptake assay
For each lipid and plant line, prepare a glass jar with 250 µl of liquid ½ MS medium and incubate at 25°C for 5-15 min in a shallow water bath (Figure 2).
Using precision tweezers, transfer 2-4 seedlings to each glass jar and shake gently. Avoid touching the part of the seedling that will be visualized later (Figure 1F and Figure 2).
Start labeling by adding 1 µl of 10 mM C6-NBD-lipid (see Recipes) to each jar (40 µM final concentration in the medium) without touching the seedlings. Shake gently and incubate at 25°C for the desired time.
Note: Incubation time depends on the permeability of the cell wall to the specific lipid, which depends on both fatty acid tail length and head group charge. For C6-NBD-sphingomyelin, 15 min are sufficient, while lipids like C6-NBD-phosphatidylethanolamine and C6-NBD-phosphatidylserine are first detectable after 3 h.
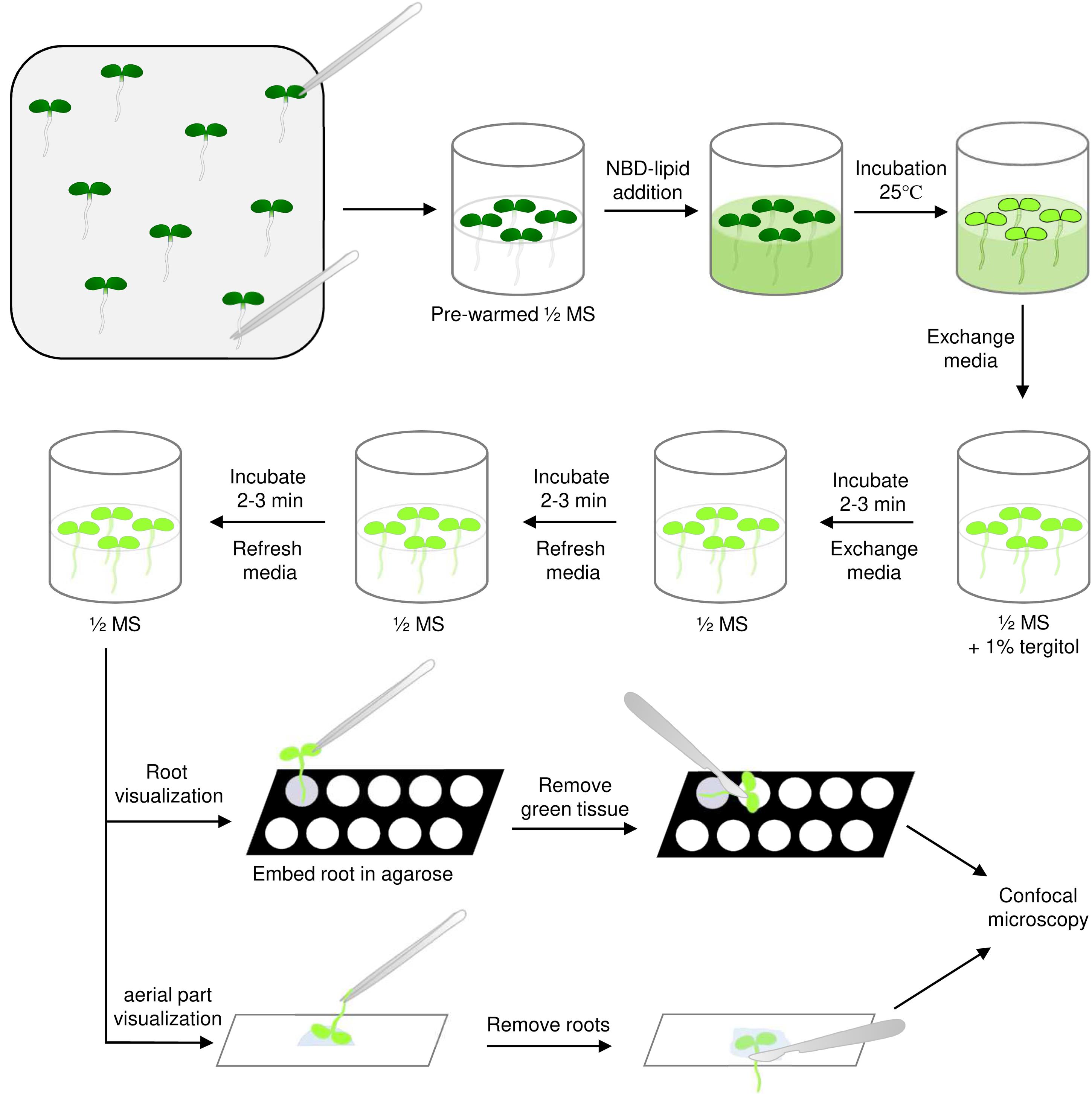
Figure 2. Lipid uptake assays in plants. Using precision tweezers, 5-day old seedlings are transferred to small glass jars containing pre-warmed ½ MS liquid medium without touching the part of the seedling that will be visualized later. After addition of the desired NBD-lipid, seedlings are incubated for different time periods. Subsequently, seedlings are washed once with detergent-containing ½ MS medium and twice with medium without detergent, before transfer to microscopic slides. Roots are immobilized using a low-concentration agarose solution, while aerial parts are simply placed in water. Plant material that will not be visualized is removed with the help of a scalpel before the sample is covered with an objective glass and visualized using confocal microscopy.Remove the supernatant, being careful not to touch the seedlings, and add 400 µl of ½ MS liquid medium with 1% tergitol solution at 25°C. Shake gently and incubate for 2-3 min at 25°C.
Repeat the wash step twice with 400 µl of ½ MS liquid medium without detergent at 25 °C, and keep the plants in ½ MS liquid medium at room temperature until visualization (not longer than 1 h).
Note: At this point, plants can be tested for metabolic conversion of the fluorescent lipid analogs using lipid extraction and thin layer chromatography analysis, as previously described (Poulsen et al., 2015).
Microscopic visualization
Visualization of roots
Taking care not to touch the areas to be visualized, place a seedling with the root inside a 6-mm well on a microscopic slide and add a 3-µl drop of 0.5% agarose solution on top (Figures 1G and Figure 2).
Use a scalpel to remove the aerial parts, and cover with an objective glass.
Mount the slide on the microscope and adjust the focus on the area of interest using bright field mode with a 63× objective.
Note: Exposure to laser light will cause bleaching. Therefore, it is recommended to adjust the focus in bright field mode before switching to fluorescent mode and immediately acquire an image.
Switch to confocal acquisition with excitation at 488 nm and emission recording at 490-508 nm. Pinhole diameter should be below two airy discs (about 100 μm).
Note: These settings are for lipids carrying an NBD group and will need to be adapted when using other fluorophores.
Set imaging parameters (laser power, detector gain, or exposure time) to get a bright signal without overexposure. We typically use 20-40% laser power and a 4-time line average scanning to prevent sample bleaching and compensate for the low signal intensity by increasing the detector gain and lowering the offset. Under these settings, the signal is stable for 25-45 s before significant bleaching occurs.
Acquire images without changing any of the set parameters and in a similar area and confocal plane for all plants (for an example, see Figure 3).
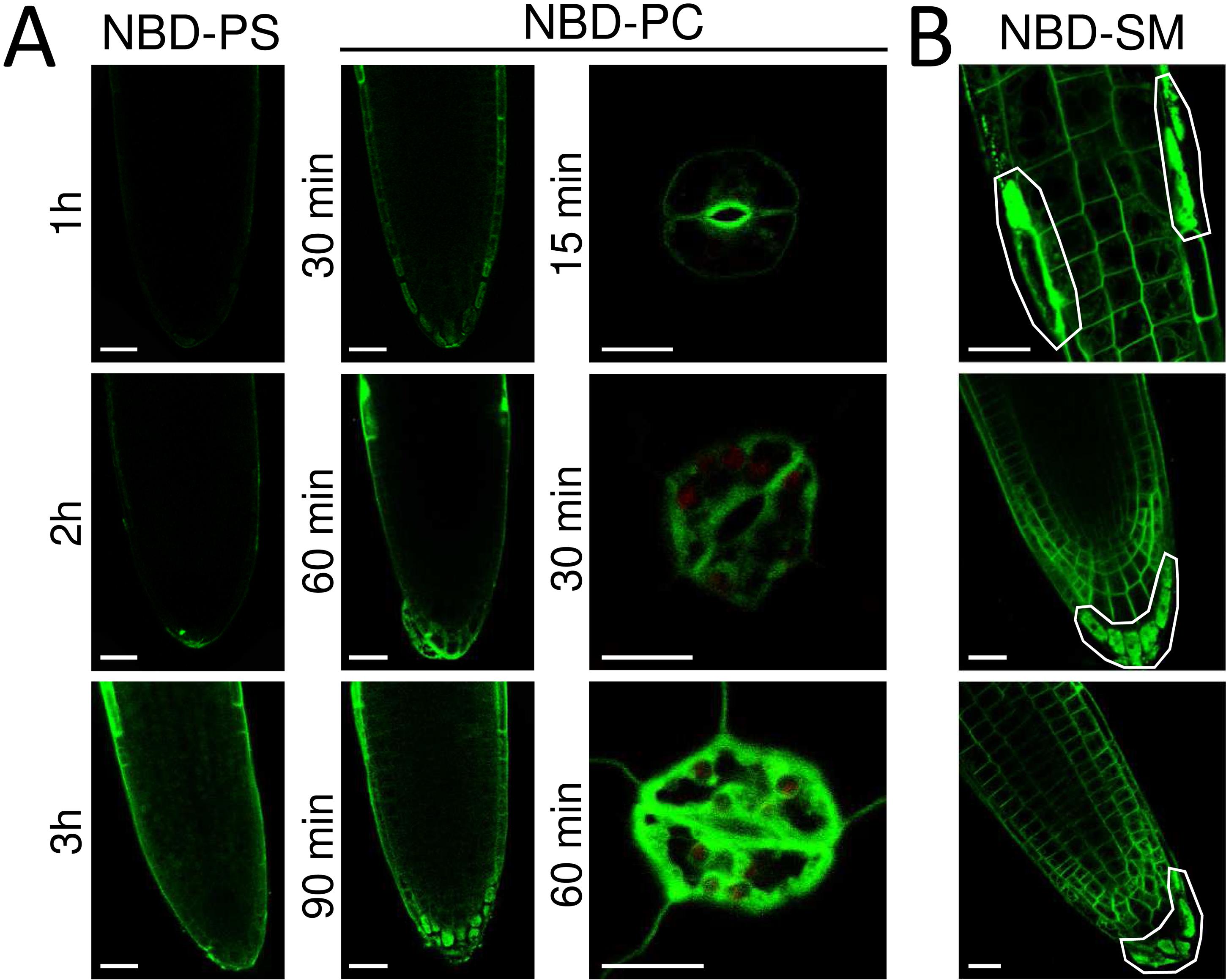
Figure 3. Examples of lipid uptake imaging in roots and aerial plant tissues. Seedlings (5-day old) were incubated with NBD-lipids for the indicated times and visualized by confocal microscopy after removal of excess lipid. (A) Time-course of a lipid uptake assay in roots and guard cells. Scale bars: roots, 25 μm; guard cells, 10 μm. (B) Examples of damaged roots. White polygons mark damaged areas. Scale bars: 20 μm.
Visualization of leaves
Drop 40-50 µl of water on a plain microscopy slide and use tweezers to place a seedling with the aerial parts inside the water (adaxial side up). Take care not to touch the areas to be visualized (Figure 1H and Figure 2).
Remove the root using a scalpel and cover with an objective glass.
Mount the slide on the microscope and locate cells using bright field mode with a 63× objective.
Note: Exposure to laser light will cause bleaching. Therefore, we recommend adjusting the focus in bright field mode before switching to fluorescent mode and acquiring an image immediately.
Switch to confocal acquisition with excitation at 488 nm and emission recording at 490-508 nm. Pinhole diameter should be below two airy discs (about 100 μm).
Note: These settings are for lipids carrying an NBD group and will need to be adapted when using other fluorophores.
Set imaging parameters (laser power, detector gain, or exposure time) to get a bright signal without overexposure. We typically use 20-40% laser power and 4-time line average scanning to prevent sample bleaching, and compensate for the low signal intensity by increasing the detector gain and lowering the offset. Under these settings, the signal is stable for 25-45 s before significant bleaching occurs.
Acquire images without changing any of the set parameters and in a similar area and confocal plane for all plants (for an example, see Figure 3).
Data analysis
Export the raw image data in a format compatible with ImageJ (e.g., tif) from the imaging system and import into ImageJ.
Go to Analyze > Set measurements and select mean gray value.
Draw a region of interest (ROI) in an area of the image where no fluorescence is present, then use Crtl + m to get the mean gray value of your background (Figure 4).
Deselect the ROI, go to Process > Math > Substract, type the value of the background in, and press OK to adjust the image.
Draw a new ROI covering an area that can be defined with clear parameters applicable to all images (e.g., the confocal image area covering from the root tip to the 8th epidermal cell above the tip, a region including five epidermal cells, a whole stoma, etc.) (Figure 4). Measure the mean gray value using Ctrl + m. Values appear sequentially in a table that can be exported to Excel.
Note: Damaged tissue will show an abnormal fluorescence compared to the rest of the sample (see Figure 3B). Be sure not to include this in the quantification.
For each plant line and lipid, use the quantification of at least three independent experiments with five seedlings each to calculate the average values for the whole population, and carry out a statistical analysis using an appropriate t-test.
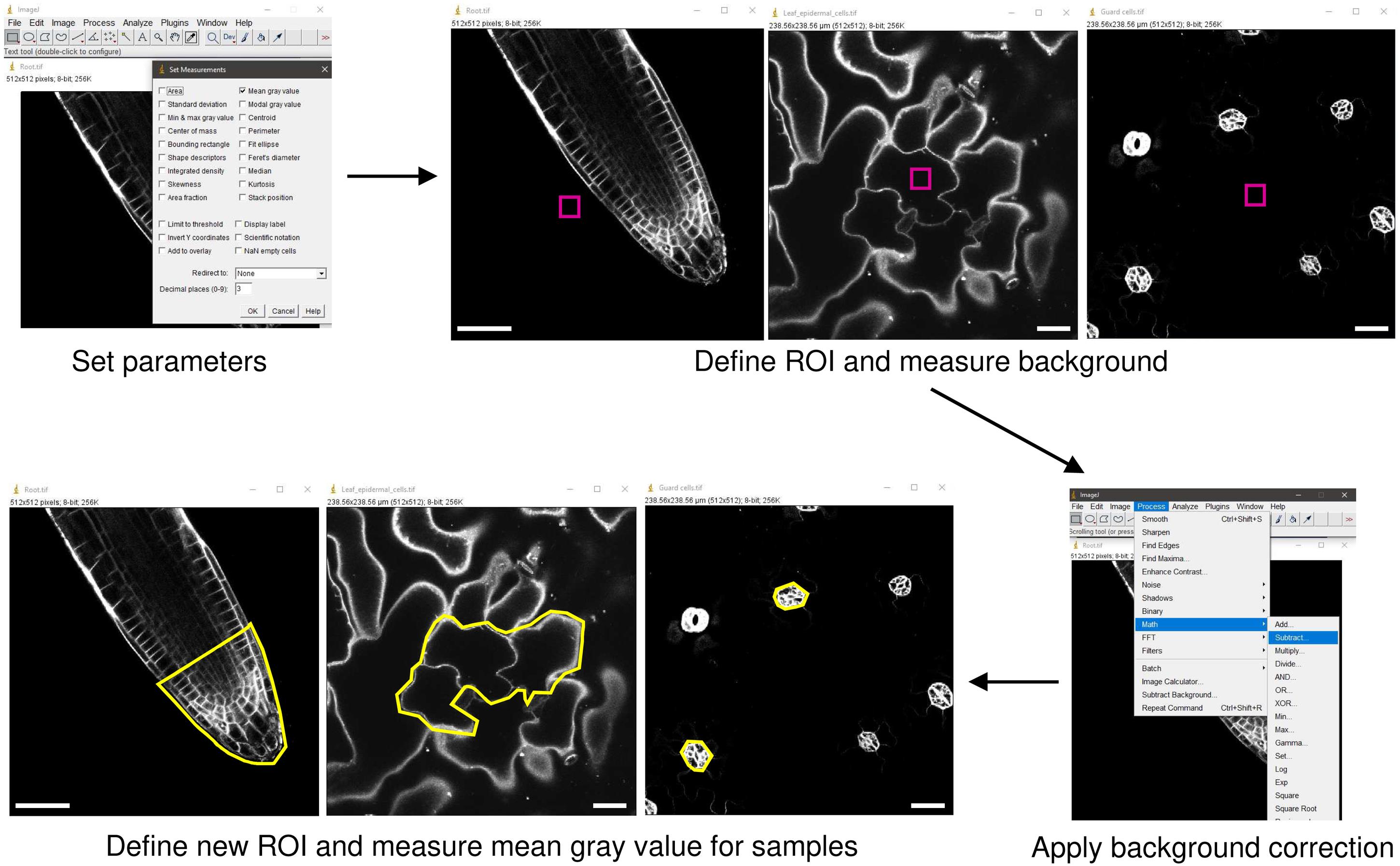
Figure 4. Image analysis using ImageJ. After setting the system to measure mean gray value (top right), define a convenient region of interest (ROI) for each image to quantify the background (top left). Apply a background correction by using the Process > Math > Substract command (bottom left). Finally, define a new ROI and measure the mean gray values of the samples. Scale bars: 25 μm.
Recipes
Note: Prepare all media using ultrapure water with purification sensitivity of 18 MΩ.cm at 25°C.
Half strength MS liquid medium
Mix 2.21 g L-1 Murashige and Skoog (MS) salts with vitamins and 0.5 g/L 2-(N-morpholino)ethanesulfonic acid (MES) in the desired water volume.
Adjust the pH to 5.7 with 2 M KOH.
Autoclave at 121°C for 20 min and store at 4°C.
Half strength MS plates
Mix MS salts and MES and adjust the pH, as above.
Add 0.7% phytoagar.
Autoclave at 121°C for 20 min.
Let the media cool down to approximately 60°C and pour onto square Petri dishes inside a laminar flow bench.
Let the plates cool down and solidify, place them in a sterile bag, and store at 4°C.
Agarose solution
Mix low-melting-point agarose with the desired amount of water to obtain a 0.5% (w/v) solution.
Heat up in the microwave until the solution starts boiling, then take out of the microwave and mix well.
If necessary, repeat the boiling step until all the agarose is fully melted.
Keep in a heating block or an incubator at 60°C to prevent solidification.
C6-NBD-lipid stocks
All steps must be performed in glass tubes in order to prevent nonspecific binding of lipids.
Note: Chloroform is a chemical hazard. Do not breathe gas/fumes/vapor/spray. Wear suitable protective clothing. Work in a fume hood.
Use a glass syringe to transfer the desired amount of C6-NBD-lipid into a 12-ml glass tube.
Dry the lipids under a gentle stream of nitrogen gas so that a dried lipid film is formed at the bottom of the tube.
Resuspend the C6-NBD-lipids in DMSO to a final concentration of 10 mM.
Note: DMSO lipid suspensions can be stored at -20°C in a well-covered glass container and used for up to 2 weeks. However, they are prone to precipitation owing their hygroscopic nature.
Acknowledgments
This protocol was adapted from our previous work (Poulsen et al., 2015). This work was supported by the Danish National Research Foundation (DNRF85) and the Danish Council for Independent Research|Natural Sciences (FNU, project number 10-083406). Current work in RLLM's group is supported by the Novo Nordisk Foundation (NovoCrops; Project Number NNF19OC0056580) and the Independent Research Fund Denmark | Nature and Universe (Project Number 1026-00024B). Imaging data were collected at the Center for Advanced Bioimaging Denmark (CAB), University of Copenhagen.
Competing interests
The authors declare no competing interests.
Ethics
No human or animal subjects are used in this protocol.
References
- Andersen, J. P., Vestergaard, A. L., Mikkelsen, S. A., Mogensen, L. S., Chalat, M. and Molday, R. S. (2016). P4-ATPases as Phospholipid Flippases-Structure, Function, and Enigmas. Front Physiol 7275.
- Lopez-Marques, R. L., Theorin, L., Palmgren, M. G. and Pomorski, T. G. (2014). P4-ATPases: lipid flippases in cell membranes. Pflugers Arch 466(7): 1227-1240.
- Poulsen, L. R., Lopez-Marques, R. L., Pedas, P. R., McDowell, S. C., Brown, E., Kunze, R., Harper, J. F., Pomorski, T. G. and Palmgren, M. (2015). A phospholipid uptake system in the model plant Arabidopsis thaliana. Nat Commun 67649.
- Roland, B. P., Naito, T., Best, J. T., Arnaiz-Yépez, C., Takatsu, H., Yu, R. J., Shin, H.-W. W. and Graham, T. R. (2019). Yeast and human P4-ATPases transport glycosphingolipids using conserved structural motifs. J Biol Chem 2941794-1806.
- Nintemann, S. J., Palmgren, M. and López-Marqués, R. L. (2019). Catch you on the flip side: A critical review of flippase mutant phenotypes. Trends Plant Sci 24468-478.
- McDowell, S. C., López-Marqués, R. L., Cohen, T., Brown, E., Rosenberg, A., Palmgren, M. G. and Harper, J. F(2015). Loss of the Arabidopsis thaliana P4-ATPases ALA6 and ALA7 impairs pollen fitness and alters the pollen tube plasma membrane. Front Plant Sci 6197.
- Schneider, C. A., Rasband, W. S. and Eliceiri, K. W. (2012). NIH Image to ImageJ: 25 years of image analysis. Nat Methods 9671-675.
- Shin, H. W. and Takatsu, H. (2019). Substrates of P4-ATPases: beyond aminophospholipids(phosphatidylserine and phosphatidylethanolamine). FASEB J 333087-3096.
Article Information
Copyright
© 2021 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
López-Marqués, R. L. and Pomorski, T. G. (2021). Imaging of Lipid Uptake in Arabidopsis Seedlings Utilizing Fluorescent Lipids and Confocal Microscopy. Bio-protocol 11(22): e4228. DOI: 10.21769/BioProtoc.4228.
Category
Plant Science > Plant cell biology > Cell imaging
Cell Biology > Cell imaging > Live-cell imaging
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










