- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
En masse DNA Electroporation for in vivo Transcriptional Assay in Ascidian Embryos
Published: Vol 11, Iss 18, Sep 20, 2021 DOI: 10.21769/BioProtoc.4160 Views: 3534
Reviewed by: Chiara AmbrogioAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
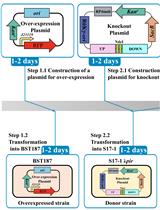
Efficient Genetic Transformation and Suicide Plasmid-mediated Genome Editing System for Non-model Microorganism Erwinia persicina
Tingfeng Cheng [...] Lei Zhao
Mar 20, 2024 2470 Views

Egg Microinjection for the Silkworm Bombyx mori
Hayato Yamada [...] Teruyuki Niimi
May 20, 2025 3345 Views
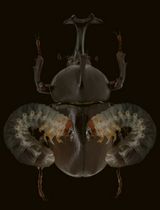
Introducing Exogenous DNA Vectors Directly into Trypoxylus dichotomus Larvae Via In Vivo Electroporation
Shinichi Morita and Teruyuki Niimi
Feb 20, 2026 210 Views
Abstract
Ascidian embryos are powerful models for functional genomics, in particular, due to the ease of generating a large number of transgenic embryos by electroporation. In addition, the small size of their genome makes them an attractive model for studying cis-regulatory elements that control gene expression during embryonic development. Here, I describe the adaptation of the seminal method developed 25 years ago in Ciona robusta for en masse DNA electroporation for in vivo transcription to additional species belonging to three genera. It is likely that similar optimizations would make electroporation successful in other ascidian species, where in vitro fertilization can be performed on a large number of eggs.
Keywords: AscidiansBackground
Ascidians are marine invertebrates that are the closest vertebrate relatives. Their fast, stereotyped, and external embryonic development gives rise to tadpole-like larvae closely resembling other chordates (vertebrates and amphioxus). Their compact genome, together with the availability of simple and efficient methods to manipulate gene expression and function, have made ascidians interesting model organisms for functional genomics (Lemaire, 2011; Satoh, 2014). In particular, the introduction of plasmid DNA into a fertilized egg by electric shock (electroporation) allows the generation of hundreds to thousands of transient transgenic embryos, which promoted Ciona robusta as the reference ascidian species almost 25 years ago (Corbo et al., 1997). DNA electroporation is a widely used method for in vivo transcriptional assays (for characterization of the activity of candidate cis-regulatory elements using reporter genes), over-expression/knockdown, and live imaging.
Ascidians are a diverse group of animals containing around 3,000 species that have been proven as interesting systems to study the evolution of developmental mechanisms since their genomes have been extensively rearranged, but their embryonic development is strongly conserved (Dardaillon et al., 2020). In particular, DNA electroporation has been applied to various species (Roure et al., 2014; Stolfi et al., 2014; Colgan et al., 2019; Coulcher et al., 2020). Here, I provide a detailed protocol to perform electroporation in four species: Phallusia mammillata, Phallusia fumigata, Ascidia mentula, and Molgula appendiculata (Figure 1), which cover almost 400 million years of evolution.
Materials and Reagents
Disposable scalpels
300- and 120-µm homemade sieves
Note: Cut a 50-ml plastic tube at ~4-5 cm from the opening. Cover a heating plate with aluminum foil. Fuse a piece of nylon mesh (Sefar Nitex, catalog numbers: 03-300/51 [300-µm opening], 03-120/49 [120-µm opening]) to the cut tube using the heated plate.
15-ml glass centrifuge tubes (Dutscher, catalog number: 092305)
15-ml plastic conical tubes (e.g., Sarstedt, catalog number: 62.554.502)
60-mm plastic Petri dishes (e.g., Sarstedt, catalog number: 82.1194.500)
92-mm plastic Petri dishes (e.g., Sarstedt, catalog number: 82.1473.001)
60-mm and 92-mm agarose-coated Petri dishes. Alternatively, gelatin-coated dishes (GF) can be used as in Sardet et al. (2011)
Note: Melt the agarose in sea water (1 g per 100 ml sea water). Fill a 92-mm dish with the hot solution. Make a thin agarose layer by pouring the agarose solution into the next dish; repeat until you have enough dishes (you will need two dishes per electroporation, plus a dozen for dechorionation, fertilization, and washing).
6-well plates
Note: Eggs tend to stick to new plastic material. To avoid this, reuse the same plates; simply wash with tap water and air dry.
Gloves (Phallusia blood is a powerful stain!)
Glass Pasteur pipets
Note: Soak them in tap water to avoid sticking of the embryos. Smooth the opening using a lighter. Prepare some with a larger opening (2-3 mm) using a diamond for egg collection in Phallusia.
100 ml glass beaker
Horizontal rotating shaker (e.g., VWR, catalog number: 444-2900)
1.5 ml microtubes
Low-binding 1.5 ml (e.g., VWR, catalog number: 525-0230) and 2 ml (e.g., VWR, catalog number: 525-0232) microtubes
4 mm electroporation cuvettes (e.g., Dutscher, catalog number: 38191)
Depression slides: 4 mm (Dutscher, catalog number: 065230) and 1.5 mm (Dutscher, catalog number: 65227) of thickness
Tungsten needle (Roboz Surgical Instrument Co., catalog number: RS-6065)
Adult ascidians are collected from the ocean by diving, using a trawl or a dredge and are provided by the French node of the European research infrastructure EMBRC (Station biologique de Roscoff: Phallusia mammillata and Ascidia mentula; Station marine de Banyuls-sur-mer: Phallusia mammillata, Phallusia fumigata, Ascidia mentula, and Molgula appendiculata)
Supercoiled circular plasmid DNA (20-100 µg) of the reporter construct
NaOH, 2.5 M
NaOH, 1 M
1 M Tris, pH 9.5
0.96 M D-mannitol solution (VWR, catalog number: 25311.297)
TE (10 mM Tris-HCl, 1 mM EDTA, pH 8.0)
25% glutaraldehyde solution (Sigma-Aldrich, catalog number: G6257)
Sodium thioglycolate (Sigma-Aldrich, catalog number: T0632)
Pronase (Sigma-Aldrich, catalog number: P5147)
0.2 µm filtered natural sea water or artificial sea water (BASWH: see Recipes)
Phallusia and Ascidia dechorionation solution (see Recipes)
Molgula dechorionation solution (see Recipes)
PBTw (see Recipes)
10× PBS (see Recipes)
X-gal staining solution (see Recipes)
X-gal stock solution (see Recipes)
Equipment
Scissors and tweezers
Temperature-controlled room set at 18°C
Dissecting scope for live embryo manipulation (e.g., Discovery V8 from Zeiss)
Square wave electroporator (Harvard Apparatus, BTX ECM830, catalog number: 45-0052)
Incubators
Dissecting scope equipped with a digital camera for staining analysis (e.g., Discovery V20+AxioCam Erc 5s from Zeiss)
Procedure
The in vivo transcriptional assay proceeds according to these successive steps: 1) gamete collection, 2) egg dechorionation, 3) fertilization, 4) DNA electroporation, 5) embryo fixation and staining, and 6) data collection and analysis. The first 4 steps differ between species. Here, I present two versions: one for P. mammillata, P. fumigate, and A. mentula (with minor changes among species); and one for M. appendiculata.

Figure 1. Adult ascidian species. (A) Phallusia mammillata (5-20 cm long). (B) Phallusia fumigata (5-20 cm long). (C) Ascidia mentula (4-10 cm long). (D) Molgula appendiculata (3-6 cm long).
Gamete collection
Ascidians are hermaphrodites. The aim is to collect both eggs and gametes, separately, from each individual.
Phallusia and Ascidia
P. mammillata, P. fumigate, and A. mentula belong to the same family, the Phlebobranchia. Gametes are collected from the gonoducts by dissection. To avoid unwanted self-fertilization, eggs are collected first. All species have transparent eggs.
Open the animals with a scalpel by cutting between the two siphons (Figure 2A). Phallusia tunic is thick and hard.
Incise the tunic toward the base of the animal on both sides. Do not cut too deep to avoid damaging the animal.
Open the tunic by pulling apart with your fingers.
Transfer the animal to a dish on its left side (you should see the heart beating at the base of the animal, opposite to the siphons). If the animal is on the right side, you should see a large, yellowish (Phallusia, Figure 2C) or reddish (Ascidia, Figure 2B) oviduct full of eggs.
Make a small cut in the oviduct and collect the eggs with a Pasteur pipet (Figure 2D).
Note: The eggs in the oviduct are very compact and embedded in a gelatinous substance. Use a wide-bore Pasteur pipet for Phallusia; otherwise, their aspiration through the small opening of a pipet may cause extreme deformation and egg death. Dead eggs are easily spotted since they turn opaque.

Figure 2. Gamete collection in Phallusia and Ascidia. (A) Schematic diagram illustrating the successive incisions (numbered red dotted lines) to open the tunic (left: lateral view with the two siphons pointing to the left; right: side view with the two siphons pointing toward the experimenter). (B) Dissected A. mentula individual where the gonoducts are exposed. (C) Dissected P. mammillata individual where the oviduct is clearly visible. (D) Egg collection in P. mammillata using a wide-bore Pasteur pipet. (E) P. mammillata egg. (F) A. mentula egg. Scale bar: 100 µm in D and E.Transfer the eggs to a 6-well plate filled with sea water. Phallusia individuals normally have a high number of eggs (0.5-2 ml eggs). Ascidia usually have fewer eggs, but the amount can reach 0.5-1 ml.
Check the egg quality (transparent and uniform egg cytoplasm, Figures 2E and 2F) and transfer them to a glass tube for several washes with sea water. Eggs can be kept overnight at 14°C, spread on a Petri dish, with no major decrease in quality.
You should see the spermiduct (sharp white duct) underneath the oviduct once you have removed the eggs. Make a small cut and collect the sperm into a 1.5-ml tube using a Pasteur pipet for Phallusia or a micropipet with a yellow tip. Avoid collecting sea water, debris, and blood. Dry sperm remains active for several days at 4°C.
Molgula
Contrary to Phallusia and Ascidia, very few eggs are present in the tiny oviduct (Figure 3D). Gametes are therefore collected by gonadal dissection. The oocytes collected from the gonad represent all stages of oogenesis. The procedure aims at collecting mainly fully grown oocytes that are not yet mature, as evidenced by the presence of a large germinal vesicle (GV). Fortunately, these oocytes mature spontaneously in sea water, and germinal vesicle breakdown (GVBD) occurs within 30-60 min.
Open the animals with scissors, starting from one siphon around the animal all the way to the other siphon (Figure 3A) (M. appendiculata are covered with debris, shells, and stones; use large scissors and make your way around these obstacles).
Pull open the tunic and drag the animal still attached by the siphons (Figure 3B).
Cut the body wall from siphon to siphon with fine scissors and pull open the animal (Figure 3C).
Under the dissecting scope, locate the gonads (one on each side of the animal, Figure 3C) under the pharyngeal basket with tweezers. You should see the short gonoducts pointing toward the siphons and the large ovary covered on one side by the testis (Figure 3D).
Cut out each gonad with scissors and place them in the same well of a 6-well plate filled with sea water (Figure 3E).

Figure 3. Gamete collection in Molgula appendiculata. (A) Schematic diagram illustrating the incision (red dotted line) to perform to open the tunic (left: lateral view with the two siphons pointing to the top; right: side view). (B) An individual (right side) after tunic (left side) removal. (C) Cut open the animal with a gonad visible at the center on each side. (D) Each gonad is composed of an ovary surrounded by a testis. The inlet shows a close-up view of the tiny spermiduct (top) and the oviduct (bottom), both openings pointing to the right. (E) A 6-well plate where pairs of gonads from 4 individuals have been collected. (F) A fertilizable egg is obtained after spontaneous maturation of a fully grown oocyte (scale bar: 50 µm).Under the dissecting scope, release the oocytes from the ovary using tweezers, and transfer the testis to a Petri dish filled with sea water (60-mm diameter). You should see a full range of oocytes (from tiny transparent oocytes to large, opaque, fully grown oocytes with GV).
Transfer all the oocytes from one individual (2 ovaries) through a 300 µm sieve placed in a 100 ml glass beaker (gonad debris will not go through).
Transfer the contents of the beaker to a 120 µm sieve and wash extensively with sea water to remove sperm and small oocytes.
Transfer the oocytes to a 60 mm agarose-coated dish.
Wait 30-60 min until GVBD occurs and produces fertilizable oocytes.
Release the sperm from the testis of all individuals that have been collected in the same Petri dish by dissociating each testis with tweezers (discard the testes afterwards). Collect the concentrated sperm solution into a 15-ml plastic tube and keep at 4°C until use.
Egg dechorionation
The chorion that protects the egg is made of an inner vitelline membrane consisting of extracellular material and an outer cellular layer of follicle cells. To perform electroporation, it is necessary to remove it; this is achieved using a mixture of sodium thioglycolate and pronase. The concentration of each compound and the duration of the dechorionation varies between species.
Note: Dechorionation is usually performed before fertilization, and the naked eggs can still be fertilized. For Molgula, since the procedure is rather quick, this can be performed after fertilization.
Collect the eggs in a glass tube and allow them to settle to the bottom.
Discard the excess sea water.
Activate the dechorionation solution by raising the pH (add 3 drops (around 100 µl) (Phallusia/Ascidia) or 6 drops (around 200 µl) (Molgula) 2.5 M NaOH to 7 ml dechorionation solution using a Pasteur pipet, and mix).
Proceed to dechorionation
In an agarose-coated dish for Phallusia/Ascidia, by adding the eggs to the dechorionation solution (~7 ml for a 60-mm dish and ~14 ml for a 92-mm dish). Mix well and place on a horizontal shaker at ~70 rpm.
In a glass tube for Molgula, by adding 3-4 ml dechorionation solution. Mix well by pipetting up and down.
Regularly check the dechorionation status under the scope. Dechorionation takes 30-45 min (Phallusia/Ascidia) and 7-15 min (Molgula) depending on the egg density.
Stop dechorionation when most eggs are clearly devoid of the chorion. Phallusia/Ascidia: gather the eggs at the center by swirling the dish and transfer them to a 15 ml glass tube. Add sea water to the top and gently mix by pipetting up and down. Molgula: discard as much dechorionation solution as possible. Add sea water to the top and gently mix by pipetting up and down.
Note: Dechorionation is more difficult to follow in Molgula: while follicle cells are quickly lost, the vitelline membrane is more difficult to see (look closely, change the scope mirror orientation; the presence of test cells on the egg’s surface is a good indication that the eggs are not fully dechorionated).
Wash extensively 2-4 times by allowing the eggs to settle down, replacing the sea water, and gently mixing with a Pasteur pipet. Debris (chorion and dead eggs) will float, while naked eggs rapidly sink to the bottom of the tube. Naked eggs are fragile and explode easily; be gentle and avoid air bubbles.
You can proceed to the next step using the same tube or collect the eggs in an agarose-coated dish for later use. Dechorionated, unfertilized eggs can be stored overnight at 4-14°C with no major decrease in quality.
Fertilization
(Phallusia/Ascidia only) Add 5 µl dry sperm to a 1.5-ml microtube containing 1 ml sea water.
(Optional) Fertilization is usually efficient with straight sperm solution; however, to achieve a complete fertilization rate and high synchrony, it is better to activate the sperm solution by raising the pH (in the ocean, sperm are activated by the slightly basic pH of the sea water). Add either 50 µl 1 M Tris pH 9.5 (Phallusia), 4 µl 1 M NaOH (Ascidia), or 8 µl 1 M NaOH (Molgula) to the sperm solution.
Note: Phallusia sperm can also be activated by incubating the 1 ml mixture for 15 min with 20 µl chorionated eggs as in Sardet et al. (2011).
Check the sperm motility by diluting an aliquot 50-100× in sea water and looking under the dissecting scope at high magnification. Sperm should be swimming frantically.
Add ~200 µl (60-mm dish) or 400-500 µl (92-mm dish or 15-ml glass tube) sperm solution.
Mix well, making the eggs float in the medium. You should see the sperm swimming intensively.
Check for egg deformation that occurs following fertilization and that should be visible within the first few minutes.
At 10-15 min post-fertilization, proceed to the washes to remove the sperm, since this protocol uses enormous quantities of sperm. (Fertilization in a Petri dish) Gather the eggs at the center by swirling the dish and transfer them to a 15-ml glass tube using a Pasteur pipet. Allow the eggs to settle to the bottom of the tube, discard most of the sea water using a Pasteur pipet, add clean sea water up to the top by gently pouring down the side of the tilted tube; resuspend the eggs thoroughly by flushing sea water with the pipet. Repeat this wash 1-3 ×.
Note: Embryos without their protective chorion are fragile and explode as soon that they come into contact with the water/air interface. Be cautious; gently manipulate them, avoiding air bubbles.
DNA electroporation
Prepare the DNA (20-100 µg) in 50 µl TE.
Add 200 µl 0.96 M D-mannitol and mix well.
Collect the fertilized eggs at the bottom of a glass tube.
Mark with a pen, the 100 µl level on the low-binding 1.5 ml tubes.
Transfer the fertilized eggs into the 1.5-ml low-binding tubes. Try to add the same number of eggs to each tube (one per construct to be tested).
Adjust the volume to the 100 µl mark with sea water.
To each tube, add 250 µl DNA/mannitol solution.
Note: Mannitol is used to reduce the salt concentration (and thus avoid electric arching during electroporation) while keeping the osmolarity at a sufficient level.
Transfer the total mixture to an electroporation cuvette (use a different Pasteur pipet for each tube to avoid mixing DNA).
Place the cuvette into the electroporator cuvette holder.
Apply a single electrical pulse of 37 V (Phallusia/Ascidia) or 20 V (Molgula) for 32 ms. The pulse should be done toward the end of the first cell cycle (but before cleavage): at 50-60 min post-fertilization.
Note: Electroporation can be performed any time after fertilization, and the timing does not affect the electroporation efficiency; however, the fertilized eggs are more robust to electroporation (but also to microinjection) during the last third of the first cell cycle.
Proceed to the electroporation of the next construct.
Once all electroporations are done, add some sea water to the cuvette and transfer the eggs to 92 mm agarose-coated Petri dishes (usually at least two dishes for one electroporation).
Note: A total of 4-6 electroporations are routinely done per fertilization round. With experience, 10-12 electroporations can be performed.
Check the dish under the dissecting scope. You should see intact embryos (since they are toward the end of the first cell cycle, you should see the myoplasm and/or deformations corresponding to the preparation of the first cleavage) and debris of exploded eggs.
Note: For species with transparent eggs (Phallusia/Ascidia), dead eggs are easily spotted since they become opaque. In most cases, there is a small proportion of eggs that do not survive the electroporation; these correspond to fragile or unfertilized eggs. In cases where you find no surviving embryos, there are two main explanations: the fertilization failed (unfertilized eggs systematically explode after the electrical pulse) or the voltage was too high. Try to make two experimental controls: non-dechorionated and non-electroporated embryos, and dechorionated and non-electroporated embryos.
Transfer the dishes to an incubator set at the desired temperature.
Note: 14-19°C is the common range of temperature for all species. They can also develop well at lower temperatures with a significant decrease in speed. Phallusia/Ascidia also develop well up to 22°C.
Spread the embryos by flushing sea water with a Pasteur pipet (they have a very high tendency to stick to each other if they are too close). Avoid moving the dishes before the 8-cell stage since blastomeres are very loosely attached to each other.
Embryo fixation and staining
Depending on the reporter gene, the course of action may differ. Here, I describe the use of LacZ as a reporter gene (detecting β-galactosidase activity using the chromogenic substrate X-gal).
Allow the embryos to develop until the desired stage.
Swirl the dish to gather the embryos at the center.
Collect the embryos using a glass Pasteur pipet and transfer them to a 2-ml low-binding microtube.
Allow the embryos to settle and discard the excess sea water.
Fill the tube with fixative (0.2% glutaraldehyde in sea water) and rotate the tube in your fingers for 30 s.
Fix for exactly 30 min at room temperature (over-fixing will abolish β-galactosidase enzymatic activity, too light a fixation may lead to embryo disintegration).
Remove half of the fixative and add 1 ml PBTw. Mix by inverting 2-3 ×.
Once the embryos have settled, replace the solution with 2 ml PBTw, mix by inversion, and wait 10 min. Repeat this wash 1 ×.
Rinse for 5 min in X-gal staining solution.
Replace with 250 µl staining solution containing 0.4 mg/ml X-gal (dilute the X-gal stock solution 100×).
Incubate at 37°C (an incubator is preferable to a water bath since no evaporation takes place in the tube).
Check the blue staining from time to time (it starts within a couple of hours depending on the strength of the tested regulatory region).
Once the staining is adequate (this may take several days, but this is a staining procedure that never produces any background), wash 2 × 5 min in PBTw. Typical examples of staining with a version of β-galactosidase targeted to the nucleus may be found in Roure et al. (2014) and Coulcher et al. (2020).
Post-fix for 1 h at room temperature or O/N at 4°C in PBTw containing 3.7% formaldehyde.
Samples can be stored long-term.
Data analysis
Data collection
Discard the fixative and wash 2 × 10 min in PBTw.
Transfer all the embryos to a thick depression slide.
Transfer part of the embryos to a thin depression slide.
Using a dissecting scope and a tungsten needle, score the number of stained embryos in the target tissue until you reach 100-200 scored embryos or until all the embryos in the tube have been scored (score only those embryos that have developed normally) (Roure et al., 2014; Coulcher et al., 2020).
Notes:
Embryos can be manipulated easily using a tungsten needle, a cat whisker, or an eyelash.
Depending on the aim, the approach can be refined by scoring not only the number of stained embryos, but also the number of stained cells per embryo; however, this may be time consuming.
Do not forget that this transgenesis method yields mosaic transgene expression (each cell has randomly inherited a variable amount of plasmid DNA) and that each embryo is an independent event; hence, the activity of a given region is determined by compiling the expression in all the embryos that are analyzed.
Take images of representative embryos.
Evaluation of cis-regulatory activity and comparisons
The assay provides qualitative data (i.e., the spatial domain of activity of a given piece of genomic DNA), but the percentage of stained embryos also reflects the strength of the regulatory region.
The score obtained may vary from experiment to experiment according to the ascidian batch, plasmid DNA preparation, and embryo concentration during electroporation; thus, we usually perform at least 3 biological replicates (different ascidian, different day of experimentation) and calculate an average score and standard deviation.
Note: Long (4-10 kb) genomic regions are usually strongly active (staining is visible within 1 h of incubation at 37°C, with over 90% of the embryos stained) and do not display much variation between experiments.
Often, the goal of this assay is to locate essential cis-regulatory elements; thus, one needs to compare the activity of various constructs. Obviously, the plasmid backbone should be the same. We also favor comparing the results of constructs that have been electroporated during the same experiment.
Recipes
Banyuls-sur-mer Artificial Sea Water HEPES (BASWH)
BASWH is a ‘Mediterranean type’ sea water with the following features: salinity of 37.9 g/L, osmolality of 1219 mOsmole, Na+/K+ ratio of 47, and a pH of ~8.0.
For 1 L, mix:
30.34 g NaCl
0.83 g KCl
1.47 g CaCl2·2H2O
4.98 g MgCl2·6H2O
6.29 g MgSO4·7H2O in MilliQ water
Adjust to 1 L
At this step, the solution can be stored at room temperature for months.
Before use, add 0.18 g NaHCO3 and 5 ml HEPES 1 M pH 8.0, and filtrate (0.2 µm). Can be stored for 1-2 months at 4°C.
Phallusia and Ascidia dechorionation solution
1% sodium thioglycolate
0.05% pronase in sea water
Mix thioglycolate in sea water by shaking, add pronase on top – no mixing – and keep at 4°C for a few hours before use.
Can be used for 1-2 weeks when kept at 4°C.
Bring aliquot to room temperature before use.
Molgula dechorionation solution
0.5% sodium thioglycolate
0.1% pronase in sea water
Mix thioglycolate in sea water by shaking, add pronase on top – no mixing – and keep at 4°C for a few hours before use.
Can be used for 1-2 weeks when kept at 4°C.
Bring aliquot to room temperature before use.
PBTw
PBS 1× containing 0.1% Tween-20
PBS 10×
For 1 L, add:
80.0 g NaCl
26.8 g Na2HPO4·7H2O
2.0 g KCl
2.4 g KH2PO4
Adjust pH to 7.4 with NaOH
Bring to 1 L, and autoclave
X-gal staining solution
1 mM MgCl2
3 mM K4Fe(CN)6
3 mM K3Fe(CN)6 in PBTw
X-gal stock solution
40 mg/ml X-gal in dimethylformamide
Acknowledgments
This protocol was used in the study ‘Conservation of peripheral nervous system formation mechanisms in divergent ascidian embryos’ by Coulcher et al. published in eLife in 2020 (doi: 10.7554/eLife.59157). This work was made possible by provision of adult ascidians thanks to the knowledge and generosity of G. Diaz (professional fisherman in Port-Vendres, France) and staff (divers and boat crew) at the marine stations of Banyuls-sur-mer and Roscoff (French node of the European research infrastructure EMBRC). I wish to thank members of my group. This work was supported by CNRS and Sorbonne Université, and by specific grants from the ANR (ANR-11-JSV2-007 and ANR-17-CE13-0027), the CNRS (DBM2020 from INSB), and the European project Assemble Plus (H2020-INFRAIA-1-2016–2017; Grant No. 730984).
Competing interests
There are no conflicts of interest or competing interests.
References
- Colgan, W., Leanza, A., Hwang, A., DeBiasse, M. B., Llosa, I., Rodrigues, D., Adhikari, H., Barreto Corona, G., Bock, S., Carillo-Perez, A., Currie, M., Darkoa-Larbi, S., Dellal, D., Gutow, H., Hokama, P., Kibby, E., Linhart, N., Moody, S., Naganuma, A., Nguyen, D., Stanton, R., Stark, S., Tumey, C., Velleca, A., Ryan, J. F. and Davidson, B. (2019). Variable levels of drift in tunicate cardiopharyngeal gene regulatory elements. Evodevo 10: 24.
- Corbo, J. C., Levine, M. and Zeller, R. W. (1997). Characterization of a notochord-specific enhancer from the Brachyury promoter region of the ascidian, Ciona intestinalis. Development 124(3): 589-602.
- Coulcher, J. F., Roure, A., Chowdhury, R., Robert, M., Lescat, L., Bouin, A., Carvajal Cadavid, J., Nishida, H. and Darras, S. (2020). Conservation of peripheral nervous system formation mechanisms in divergent ascidian embryos. Elife 9: e59157.
- Dardaillon, J., Dauga, D., Simion, P., Faure, E., Onuma, T. A., DeBiasse, M. B., Louis, A., Nitta, K. R., Naville, M., Besnardeau, L., Reeves, W., Wang, K., Fagotto, M., Gueroult-Bellone, M., Fujiwara, S., Dumollard, R., Veeman, M., Volff, J. N., Roest Crollius, H., Douzery, E., Ryan, J. F., Davidson, B., Nishida, H., Dantec, C. and Lemaire, P. (2020). ANISEED 2019: 4D exploration of genetic data for an extended range of tunicates. Nucleic Acids Res 48(D1): D668-D675.
- Lemaire, P. (2011). Evolutionary crossroads in developmental biology: the tunicates. Development 138(11): 2143-2152.
- Roure, A., Lemaire, P. and Darras, S. (2014). An otx/nodal regulatory signature for posterior neural development in ascidians. PLoS Genet 10(8): e1004548.
- Sardet, C., McDougall, A., Yasuo, H., Chenevert, J., Pruliere, G., Dumollard, R., Hudson, C., Hebras, C., Le Nguyen, N. and Paix, A. (2011). Embryological methods in ascidians: the Villefranche-sur-Mer protocols. Methods Mol Biol 770: 365-400.
- Satoh, N. (2014). Developmental genomics of ascidians. Hoboken, New Jersey, John Wiley & Sons, Inc.
- Stolfi, A., Lowe, E. K., Racioppi, C., Ristoratore, F., Brown, C. T., Swalla, B. J. and Christiaen, L. (2014). Divergent mechanisms regulate conserved cardiopharyngeal development and gene expression in distantly related ascidians. Elife 3: e03728.
Article Information
Copyright
Darras. This article is distributed under the terms of the Creative Commons Attribution License (CC BY 4.0).
How to cite
Readers should cite both the Bio-protocol article and the original research article where this protocol was used:
- Darras, S. (2021). En masse DNA Electroporation for in vivo Transcriptional Assay in Ascidian Embryos. Bio-protocol 11(18): e4160. DOI: 10.21769/BioProtoc.4160.
- Coulcher, J. F., Roure, A., Chowdhury, R., Robert, M., Lescat, L., Bouin, A., Carvajal Cadavid, J., Nishida, H. and Darras, S. (2020). Conservation of peripheral nervous system formation mechanisms in divergent ascidian embryos. Elife 9: e59157.
Category
Developmental Biology > Morphogenesis
Molecular Biology > DNA > Transformation
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link








