- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Purification of Mitochondrial Ribosomes with the Translocase Oxa1L from HEK Cells
(*contributed equally to this work) Published: Vol 11, Iss 15, Aug 5, 2021 DOI: 10.21769/BioProtoc.4110 Views: 5285
Reviewed by: Khyati Hitesh ShahAndrea SaponaroAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

Workflow for Fluorescence-Targeted Lamella Milling From Vitrified Cells With a Coincident Fluorescence, Electron, and Ion Beam Microscope
Elise G. Perton [...] Jacob P. Hoogenboom
Jul 20, 2025 2866 Views

Protocol for the Preparation of a Recombinant Treacle Fragment for Liquid–Liquid Phase Separation (LLPS) Assays
Nadezhda V. Petrova [...] Artem K. Velichko
Sep 20, 2025 1860 Views

Optimized Secretome Sample Preparation From High Volume Cell Culture Media for LC–MS/MS Proteomic Analysis
Basil Baby Mattamana [...] Peter Allen Faull
Dec 20, 2025 1323 Views
Abstract
Mitochondrial ribosomes (mitoribosomes) perform protein synthesis inside mitochondria, the organelles responsible for energy conversion and adenosine triphosphate (ATP) production in eukaryotic cells. To investigate their functions and structures, large-scale purification of intact mitoribosomes from mitochondria-rich animal tissues or HEK cells have been developed. However, the fast purification of mitoribosomes anchored to the mitochondrial inner membrane in complex with the Oxa1L translocase remains particularly challenging. Herein, we present a protocol recently developed and modified in our lab that provides details for the efficient isolation of intact mitoribosomes with its translocase Oxa1L. We combined the cell culture of PDE12-/- or wild-type HEK293 cell lines with the isolation of mitochondria and the purification steps used for the biochemical and structural studies of mitoribosomes and Oxa1L.
Graphic abstract:

Schematic procedure for the purification of mitoribosomes from HEK cells. The protocol described herein includes two main sections: 1) isolation of mitochondria from HEK cells; and 2) purification of mitoribosome-Oxa1L from mitochondria. RB: Resuspension Buffer (see Recipes) (Created with BioRender.com).
Background
Human mitochondria possess their own genome and house all the components necessary for transcription, RNA maturation, and translation of the encoded genetic information (Dennerlein et al., 2017). The mitoribosome, one of the most important machineries within mitochondria, is responsible for the translation of essential mitochondrial mRNAs encoding components of oxidative phosphorylation (OXPHOS) complexes (Ott et al., 2016). Since all proteins translated by the human mitoribosome are membrane proteins, knowledge of the mechanism of co-translational membrane protein insertion is central to understanding mitoribosome translation (Hildenbeutel et al., 2012; Stiller et al., 2016). In human mitochondria, the oxidase assembly 1-like (Oxa1L) translocase plays a central role in the membrane insertion of mitochondrially encoded products (Haque et al., 2010; Stiller et al., 2016); however, due to the unique features of mitochondria, little is known regarding the molecular and quality control mechanisms underlying mitochondrial translation.
Owing to the structures obtained by single-particle electron cryo-microscopy (cryo-EM), opportunities now arise to comprehensively study the molecular mechanisms underlying the translational action and quality control of the human mitoribosome (Rathore, 2020). Although the existence of quality control in mitochondria has been predicted based on analogy with bacterial and eukaryotic cytosols (Ayyub et al., 2020), biochemical or structural evidence of ribosome-associated quality control in mitochondria remain elusive. We reasoned that any attempt to induce translational stalling may trap mitoribosomes in various stages of the translation cycle during the act of nascent chain insertion into the inner mitochondrial membrane, and generate intermediates suitable for structural analysis by cryo-EM. To this end, we purified mitoribosomes from a genetically engineered human cell line lacking the 2’-5’ phosphodiesterase exonuclease 12 (PDE12). PDE12 facilitates the maturation of mt-tRNA; therefore, a PDE12 knockout can lead to aberrant polyadenylation of the 3’ ends of mt-tRNAs and, consequently, mitoribosome stalling (Rorbach et al., 2011; Pearce et al., 2017). As expected, we observed a substantial proportion of stalled mitoribosomes and discovered a mitoribosome-associated quality control pathway (mtRQC) to rescue the elongational stalling ribosomes (Desai et al., 2020). Our report of the first mitoribosome-associated quality control pathway provides novel insights into the regulation of mitochondrial translation. Since the cell needs to detect and respond rapidly even to subtle changes in translation rates, quality control is intimately coupled to elongation. During purification, we included GMPPCP, a non-hydrolysable GTP analog, to prevent dissociation of GTPases from mitoribosomes, and successfully captured five additional structures of virtually every elongating ribosomal state. Purifying the mitoribosome in complex with its translocase Oxa1L was challenging since membrane proteins are unstable and tend to dissociate during complicated isolation steps. Accordingly, we added n-dodecyl β-D-maltoside (β-DDM) along with cardiolipin to solubilize mitoribosomes anchored to the mitochondrial inner membrane via the Oxa1L translocase. We also trapped Oxa1L on all our active and stalled ribosomes, giving us a first glimpse at the co-translational insertion of mitochondrial inner membrane proteins.
This protocol was modified from prior methods developed in our lab (Amunts et al., 2015; Brown et al., 2017) and by others (Greber et al., 2014 and 2015; Aibara et al., 2018), as well as in our current work (Desai et al., 2020). The method described here can be used to obtain mitoribosomes in complex with Oxa1L, which has advantages such as the applicability for mitochondria isolation from large volumes of suspension cell culture, the use of common media and buffers, ease of handling, and absence of additional affinity purification steps. In our previous publication and experiments, we observed a good reproducibility between HEK293 cell batches using this methodology.
Materials and Reagents
T75/T175 flasks (Thermo ScientificTM)
1 L/2 L Tissue Culture-treated roller bottles (CorningTM)
50 ml/250 ml/5 L polycarbonate Erlenmeyer flasks (CorningTM)
Teflon/glass Dounce homogenizers (Cambridge Glassblowing Limited)
Miracloth (Merck Millipore Calbiochem, catalog number: 475855)
SW40 tubes for mitochondria gradient (Beckman, catalog number: 344060)
Polycarbonate bottle with cap assembly for type 70Ti rotor, for cushion (Beckman, catalog number: 355618)
TLS-55 tubes for mitoribosome gradient (Beckman, catalog number: 347356)
Dulbecco’s Modified Eagle Medium (GibcoTM, Life technologies)
Tetracycline-free fetal bovine serum (GibcoTM, Life technologies)
Freestyle 293 Expression medium (GibcoTM, Life technologies)
Dnase (RNA-free) (HT Biotechnology, N401a)
n-Dodecyl β-D-maltoside (Anatrace, catalog number: D310)
18:1 Cardiolipin (TOCL; Avanti, catalog number: 710335C)
β, γ-Methyleneguanosine 5′-triphosphate sodium salt (GMPPCP; Sigma, catalog number: M3509)
Protease inhibitor (cOmpleteTM, Roche)
Rabbit anti-mS27 (Abcam, catalog number: ab67157)
Mouse anti-OXA1L (Abcam, catalog number: ab88975)
Goat anti-mouse Alexa Fluor Plus 800 (ThermoFisher, catalog number: A32730)
Goat anti-rabbit Alexa Fluor Plus 680 (ThermoFisher, catalog number: A32734)
MIB Buffer (Mitochondria isolation buffer, see Recipes)
SM4 Buffer (Sucrose/mannitol buffer, see Recipes)
MIBSM Buffer (MIB+SM4: Experimental buffer, see Recipes)
SEM Buffer (Sucrose EDTA MOPS buffer, see Recipes)
60%, 32%, 23%, and 15% Sucrose Gradient Buffer (see Recipes)
Lysis Buffer (see Recipes)
Sucrose Cushion Buffer (see Recipes)
Resuspension Buffer (see Recipes)
15-30% Linear Sucrose Gradient Buffer (see Recipes)
Equipment
Automated cell counter (Countess® II FL)
Avanti-J26 XP centrifuge (Beckman Coulter)
High speed centrifuge (Beckman Coulter, model: Avanti J-26XP)
Ultracentrifuge (Beckman Coulter, model: Optima L-80 XP)
Tabletop ultracentrifuge (Beckman Coulter, model: OptimaTM MAX)
Centrifuge (Eppendorf, model: 5430 R)
NanoDrop® 2000 (Thermo)
Odyssey® CLx Infrared Imaging System (LI-COR Biosciences)
Vitrobot Mark IV (Thermo Fisher Scientific)
Titan Krios microscope (Thermo Fisher Scientific FEI)
Procedure
Cell Culture
Maintain PDE12-/- HEK293 cells in T75 flasks in DMEM supplemented with 10% FBS, at 37°C and 5% CO2. Cells should be ready for transfer to a T175 flask in ~2-3 days.
Scale up by splitting every 2-3 days until the target cell mass is reached. At least 8 flasks with 30 ml cells per flask is recommended as a starting point for adapting to suspension culture.
At 90% confluence, detach cells with 0.05% trypsin-EDTA for 10-15 min at 37°C, and monitor the cell rounding. Centrifuge at 200 × g to remove trypsin, then resuspend the pelleted cells 2-3 times in 10-20 ml Freestyle medium with 1% FBS to wash out old media.
Resuspend the detached cells in 30-50 ml Freestyle media with 1% FBS, adjust cell density with media to around 0.5 × 106 cells/ml, and grow at 37°C, 8% CO2, 120 × g.
Scale up by diluting and splitting with pre-warmed fresh Freestyle media with 1% FBS. Count cells using an automated cell counter and adjust cell concentration to 0.5 × 106 cells/ml.
Observe and count the cells every other day, and proceed to split/dilute the cells if the cell density is above 1.5 × 106 cells/ml.
The final cell density can range from 0.8 to 1.5 × 106 cells/ml, and the isolation of mitochondria requires at least 10 L as a starting point (Please see Notes for starting volume of wild-type).
Harvest cells by centrifugation at 1,000 × g for 7 min at 4°C. Decant the supernatant carefully and resuspend the pelleted cells quickly in pre-cooled PBS to wash out the media.
Centrifuge the resuspended cells at 1,200 × g for 10 min at 4°C. Decant the supernatant carefully and weigh the pellet. Continue to isolate the mitochondria from harvested cells, or snap-freeze the cells in liquid nitrogen and store at -80°C.
Mitochondria isolation
Pre-cool the Teflon/glass Dounce homogenizer O/N prior to use.
Resuspend the pellet (e.g., 10 g) in 60 ml MIB buffer.
Allow cells to swell by gently stirring in a 4°C cold room for 15 min.
Add 20 ml SM4 buffer (1/3 of MIB buffer volume).
Dounce homogenize the samples (> 60 up-and-down passes).
Centrifuge the homogenized sample at 800 × g for 15 min at 4°C. Collect supernatant (Supernatant-1) through a Miracloth (or other cheese cloth with a typical pore size of ~22-25 µm). Keep the pellet.
Resuspend the pellet in 20 ml MIBSM (3:1 MIB:SM) buffer, homogenize manually (~15 up-and-down passes), then centrifuge at 4°C, 800 × g for 15 min.
Collect supernatant (Supernantant-2) through a Miracloth and combine with Supernatant-1 from B-6. Centrifuge at 1,000 × g for 15 min at 4°C. Collect supernatant.
Centrifuge the supernatant from the previous step at 10,000 × g for 15 min at 4°C.
Keep pellet, but carefully wash out loose pellet without disturbing the tight portion.
Resuspend in 10 ml MIBSM buffer, add 100 U RNase-free DNase I per 10 g cells, rotate on a roller in a cold room for 20 min.
Centrifuge at 10,000 × g for 15 min at 4°C.
Resuspend the pellet in ~2 ml SEM buffer and homogenize gently with a small Dounce homogenizer. Perform no more than five up-and-down passes.
Prepare a 15-23-32-60% sucrose gradient in SW40 tubes. Place 1.5 ml of the 60% sucrose stock buffer into the Beckman Ultra-Clear centrifuge tube. Carefully overlay 4.5 ml of the 32% sucrose stock buffer. Repeat with 1.5 ml of the 23% sucrose stock buffer and then with 1.5 ml of the 15% sucrose stock buffer.
Load resuspended crude mitochondrial sample from Step B13 on top of the 15% sucrose in the density gradient prepared in Step B14. Centrifuge in an SW40 rotor at 28, 000 × g for 1 h at 4°C.
The intact mitochondria form a brown band at the 60%-32% sucrose interface. Carefully remove the sucrose from the top until reaching the mitochondrial band, then take the brown band with a pipette at the interface of 32% and 60% sucrose.
Snap-freeze the purified mitochondria in liquid nitrogen and store at -80°C.
Mitoribosome-Oxa1L purification
Defrost the frozen mitochondria on ice.
Add 2 volumes of lysis buffer to mitochondria (e.g., Add 8 ml lysis buffer to 4 ml mitochondria). Mix immediately by inverting the tube several times.
Homogenize with a small Teflon/glass Dounce homogenizer to assist lysis.
Rotate on a roller in a cold room for 20 min to complete the lysis.
Centrifuge the lysed material (approximately 9 ml) at 30,000 × g for 20 min at 4°C to remove the insoluble material. Decant the supernatant carefully from the pellet and discard the pellet.
Prepare the sucrose cushion in Ti70 (tube vol approx. 24 ml): underlay 7 ml 1 M (34%, w/v) Sucrose Cushion Buffer at the bottom of the tube.
Layer approximately 17 ml lysed mitochondrial sample on the sucrose cushions carefully, resulting in a lysate:cushion ratio of 2.5:1.
Centrifuge the sample at ~231,550 × g for 60 min at 4°C.
Discard the supernatant and rinse the tubes sequentially with resuspension buffer to remove residual sucrose (without disturbing the pellet).
Resuspend pellets in a total of 100 µl resuspension buffer.
Measure mitoribosome absorption at A260.
Load the entire sample onto a linear 15-30% sucrose gradient tube. Centrifuge in a TLS-55 rotor at 213, 626 × g for 60-90 min at 4°C.
Fractionate the gradient, determine the optical density at A260, and pool the fraction corresponding to the nucleic acid peak together. The typical A260:A280 ratio of the peak is >1.6.
Exchange the buffer if necessary and dilute the sample with at least 5-10 volumes of final buffer; concentration or dialysis are recommended. Calculate the final concentration using a NanoDrop and the conversion: 1 A260 = 0.1 mg/ml.
Use the purified mitoribosome sample for subsequent experiments or snap-freeze in resuspension buffer and store at -80°C.
Data analysis
The mitoribosome-Oxa1L purification protocol described here employs PDE12-/- HEK293 cells, but is applicable to wild-type or other HEK293-derived cells. The high-quality purified sample enables biochemical and structural analysis, which can be applied to study human mitoribosomal translation. Representative results for structural studies are presented as figures below; these analyses are just briefly described to work as quality controls for the sample preparation in this protocol.
Following subsequent centrifugation steps, the intact mitochondria are separated on the sucrose gradient (Figure 1); special care is needed to collect the brown band at the 60%-32% interface with minimal contamination from the surrounding buffer. Preparation of high-quality intact mitochondria can be extrapolated to purify or study other mitochondrial macromolecules.
Mitochondria that have been successfully isolated are lysed, and mitoribosome-Oxa1L are finally purified with a sucrose cushion followed by a continuous linear 15-30% sucrose gradient. The sucrose gradient can then be fractionated, either automatically or manually, by carefully taking fractions with a pipette or punching the tube bottom with a 21 G needle and collecting the drops. Two major mitoribosomal populations are identified in the gradient: the 55S monosome and the large 39S subunit, as illustrated in Figure 2. An additional peak located closer to the bottom may appear in preparations contaminated with 80S cytoplasmic ribosomes. Oxa1L was present in our samples, as confirmed by western blotting (Figure 2B) and cryo-EM maps (Figure 3). To investigate nascent chain translocation, we performed focused classification with signal subtraction (FCwSS) on the mitoribosomal exit tunnel. For further information or other in silico processing, please see Figure S3, S12A in our published paper (Desai et al., 2020).
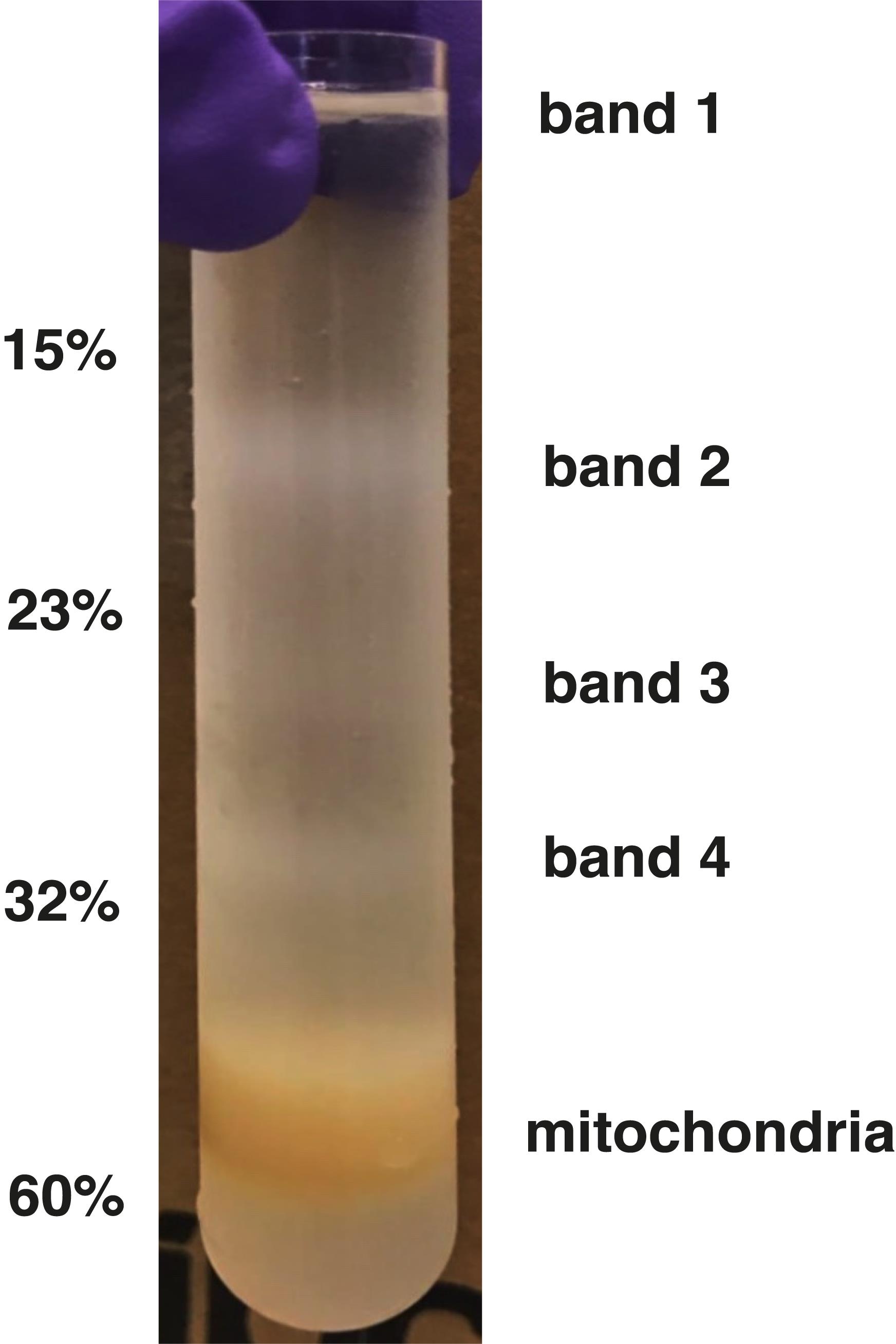

Figure 1. Isolation of mitochondria on a sucrose gradient. Mitochondria were separated on a discontinuous sucrose gradient. The purified mitochondria are found at the 32%/60% interface.
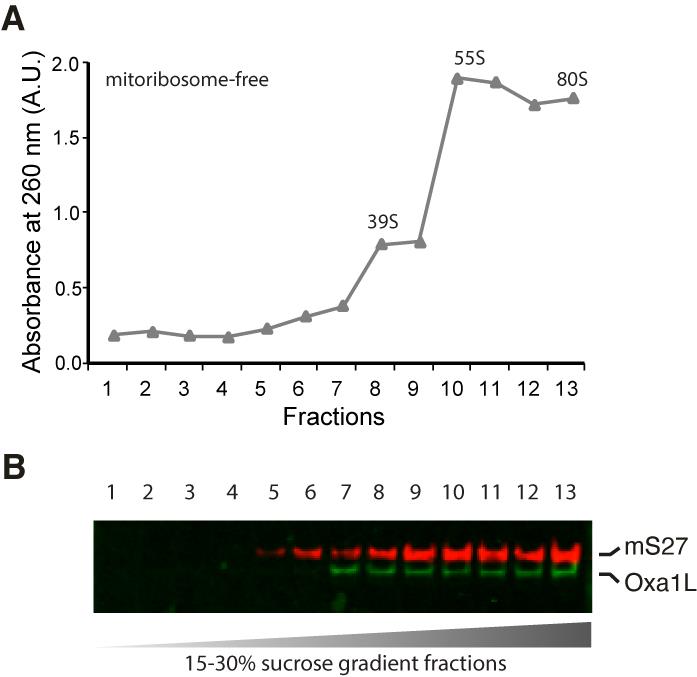
Figure 2. Purification of mitoribosomes with Oxa1L on a sucrose gradient. A. Absorbance at 260 nm of the sucrose gradient fractions. B. Western blotting of mitoribosomal protein mS27 and OXA1L in fractions, showing that Oxa1L co-pellets with mitoribosomes in our preparation.
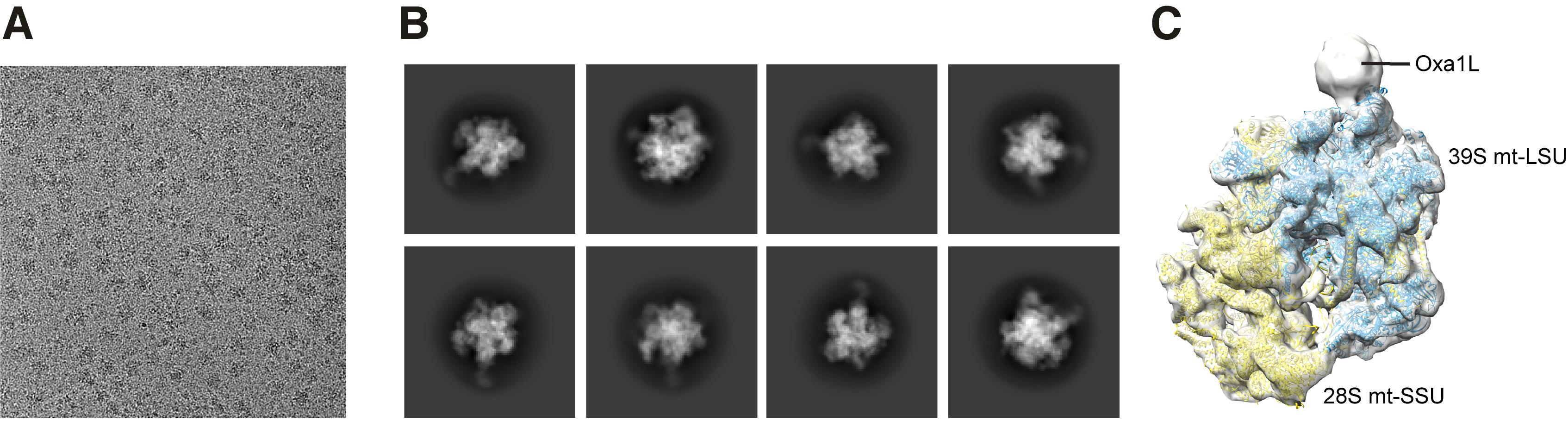

Figure 3. cryo-EM micrograph, 2D classes, and 3D initial reconstruction. A. A micrograph of the mitoribosome-Oxa1L sample. B. Representative processing data (2D classes). C. 3D reconstruction of the mitoribosome-Oxa1L complex.
Notes
Notes for cell culture:
The cell viability should be maintained at >90%, and for the final culture that is harvested >95%. The cell culture procedure is specifically for PDE12-/- cells used in this manuscript.
For wild-type cells, all steps are the same except two:
Grow cells in FreeStyle media + 5% FBS in suspension culture. Split when the cell density reaches 2 × 106 cells/ml (compared with A3-4).
Harvest cells at a concentration of ~3-4 × 106 cells/ml.
Once you have 10-20 g cell pellets (from 10-15 L PDE12-/- HEK cells or 2 L wild-type HEK293 cells) as a starting point, there are no differences in the following steps for mitochondria isolation and mitoribosome purification.
Notes for mitochondria isolation:
It is important to work quickly and keep everything on ice throughout the procedure.
Besides a Dounce homogenizer in Step B5; alternatively, you could use nitrogen cavitation at a pressure of 500 psi for 20 min (Aibara et al., 2018). Pre-cool the nitrogen cavitation chamber O/N at 4°C.
Notes for mitoribosome-Oxa purification:
You can always start with a smaller volume of cells and subsequently prepare a smaller volume of buffers to keep the ratio the same.
The small sucrose gradient with the TLS-55 rotor in this protocol allows for the rapid purification of mitoribosome-Oxa1L; however, there is normally an overlap between 55S monosome and 39S large subunit peaks. Separation of the 55S and 39S populations is not necessary for cryo-EM studies, given that further processing will be performed. If a better separation is needed in other situations, larger tubes (e.g., SW40) and a longer running time (e.g., 16 h) are recommended.
In our dataset, among all the mitoribosome samples, the proportion of complexes of mitoribosome with Oxa1L is around 12%.
Recipes
Buffers for mitochondria isolation
Mitochondrial isolation buffer (MIB)
50 mM HEPES-KOH, pH 7.4
10 mM KCl
1.5 mM MgCl2
1 mM EDTA
1 mM EGTA
1 mM DTT
Protease inhibitors (1 tablet/50 ml)
Sucrose/mannitol buffer (SM4)
280 mM sucrose
840 mM mannitol
50 mM HEPES-KOH, pH 7.5
10 mM KCl
1.5 mM MgCl2
1 mM EDTA
1 mM EGTA
1m M DTT
Protease inhibitors (1 tablet/50 ml)
Experimental buffer (MIBSM)
3 volumes MIB mix with 1 volume SM4 (e.g., 120 ml MIB buffer + 40 ml SM4 buffer)
Sucrose-EDTA MOPS (SEM) buffer
250 mM sucrose
1 mM EDTA
20 mM MOPS-KOH (alternative: HEPES-KOH, pH 7.4)
Sucrose gradient (SG) buffer
20 mM HEPES-KOH, pH 7.4
1 mM EDTA and 60%/32%/23% and 15% sucrose, respectively (make separate stock solutions of 4 different sucrose concentrations for the preparation of the stepwise sucrose gradient).
Buffers for mitoribosome purification
Lysis buffer
25 mM HEPES-KOH, pH 7.4
150 mM KCl
50 mM MgOAc
1.5% β-DDM
0.15 mg/ml Cardiolipin
500 mM GMPPCP
2 M DTT
Protease inhibitors (1 tablet/50 ml)
Sucrose cushion (SC)
1 M sucrose (34% w/v)
20 mM HEPES-KOH, pH 7.4
100 mM KCl
20 mM MgOAc
0.6% β-DDM
0.06 mg/ml Cardiolipin
250 mM GMPPCP
2 mM DTT
Resuspension buffer (RB)
20 mM HEPES-KOH, pH 7.4
100 mM KCl
5 mM MgOAc
0.15% β-DDM
0.015 mg/ml Cardiolipin
250 mM GMPPCP
2 mM DTT
15-30% linear sucrose gradients
15-30% linear sucrose in 20 mM HEPES-KOH, pH 7.4
100 mM KCl
5 mM MgOAc
0.05% β-DDM
0.005 mg/ml Cardiolipin
250 mM GMPPCP
2 mM DTT
Acknowledgments
This work received funding from the following sources: UK Medical Research Council (MC_U105184332); Wellcome Trust Senior Investigator award (WT096570); Agouron Institute; and the Louis-Jeantet Foundation. H.Y. is funded by an EMBO Long-Term Fellowship (EMBO ALTF 806-2018), and N.D. is funded by a Wellcome Trust Clinical PhD Fellowship (110301/Z/15/Z). This protocol was taken from Desai et al. (2020) with minor modifications.
Competing interests
The authors declare no competing financial or non-financial interests.
References
- Aibara, S., Andrell, J., Singh, V. and Amunts, A. (2018). Rapid Isolation of the Mitoribosome from HEK Cells. J Vis Exp(140).
- Amunts, A., Brown, A., Toots, J., Scheres, S. H. W. and Ramakrishnan, V. (2015). Ribosome. The structure of the human mitochondrial ribosome. Science 348(6230): 95-98.
- Ayyub, S. A., Gao, F., Lightowlers, R. N. and Chrzanowska-Lightowlers, Z. M. (2020). Rescuing stalled mammalian mitoribosomes - what can we learn from bacteria? J Cell Sci 133(1).
- Brown, A., Rathore, S., Kimanius, D., Aibara, S., Bai, X. C., Rorbach, J., Amunts, A. and Ramakrishnan, V. (2017). Structures of the human mitochondrial ribosome in native states of assembly. Nat Struct Mol Biol 24(10): 866-869.
- Dennerlein, S., Wang, C. and Rehling, P. (2017). Plasticity of Mitochondrial Translation. Trends Cell Biol 27(10): 712-721.
- Desai, N., Yang, H., Chandrasekaran, V., Kazi, R., Minczuk, M. and Ramakrishnan, V. (2020). Elongational stalling activates mitoribosome-associated quality control. Science 370(6520): 1105-1110.
- Greber, B. J., Boehringer, D., Leitner, A., Bieri, P., Voigts-Hoffmann, F., Erzberger, J. P., Leibundgut, M., Aebersold, R. and Ban, N. (2014). Architecture of the large subunit of the mammalian mitochondrial ribosome. Nature 505(7484): 515-519.
- Greber, B. J., Bieri, P., Leibundgut, M., Leitner, A., Aebersold, R., Boehringer, D. and Ban, N. (2015). Ribosome. The complete structure of the 55S mammalian mitochondrial ribosome. Science 348(6232): 303-308.
- Haque, M. E., Elmore, K. B., Tripathy, A., Koc, H., Koc, E. C. and Spremulli, L. L. (2010). Properties of the C-terminal tail of human mitochondrial inner membrane protein Oxa1L and its interactions with mammalian mitochondrial ribosomes. J Biol Chem 285(36): 28353-28362.
- Hildenbeutel, M., Theis, M., Geier, M., Haferkamp, I., Neuhaus, H. E., Herrmann, J. M. and Ott, M. (2012). The membrane insertase Oxa1 is required for efficient import of carrier proteins into mitochondria. J Mol Biol 423(4): 590-599.
- Ott, M., Amunts, A. and Brown, A. (2016). Organization and Regulation of Mitochondrial Protein Synthesis. Annu Rev Biochem 85: 77-101.
- Pearce, S. F., Rorbach, J., Van Haute, L., D'Souza, A. R., Rebelo-Guiomar, P., Powell, C. A., Brierley, I., Firth, A. E. and Minczuk, M. (2017). Maturation of selected human mitochondrial tRNAs requires deadenylation. Elife 6: e27596.
- Rathore, S. (2020). Structural characterisation of mitochondrial macromolecular complexes using cryo-EM: Mitoribosome biogenesis and respiratory chain supercomplex Doctoral dissertation. Stockholm University.
- Rorbach, J., Nicholls, T. J. and Minczuk, M. (2011). PDE12 removes mitochondrial RNA poly(A) tails and controls translation in human mitochondria. Nucleic Acids Res 39(17): 7750-7763.
- Stiller, S. B., Hopker, J., Oeljeklaus, S., Schutze, C., Schrempp, S. G., Vent-Schmidt, J., Horvath, S. E., Frazier, A. E., Gebert, N., van der Laan, M., Bohnert, M., Warscheid, B., Pfanner, N. and Wiedemann, N. (2016). Mitochondrial OXA Translocase Plays a Major Role in Biogenesis of Inner-Membrane Proteins. Cell Metab 23(5): 901-908.
Article Information
Copyright
© 2021 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Readers should cite both the Bio-protocol article and the original research article where this protocol was used:
- Yang, H. and Desai, N. (2021). Purification of Mitochondrial Ribosomes with the Translocase Oxa1L from HEK Cells. Bio-protocol 11(15): e4110. DOI: 10.21769/BioProtoc.4110.
- Desai, N., Yang, H., Chandrasekaran, V., Kazi, R., Minczuk, M. and Ramakrishnan, V. (2020). Elongational stalling activates mitoribosome-associated quality control. Science 370(6520): 1105-1110.
Category
Biochemistry > Protein > Isolation and purification
Biophysics > Microscopy > Cryogenic microscopy
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link









