- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
A One-enzyme RT-qPCR Assay for SARS-CoV-2, and Procedures for Reagent Production
Published: Vol 11, Iss 2, Jan 20, 2021 DOI: 10.21769/BioProtoc.3898 Views: 6199
Reviewed by: Anonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
Visualization, Quantification, and Modeling of Endogenous RNA Polymerase II Phosphorylation at a Single-copy Gene in Living Cells
Linda S. Forero-Quintero [...] Timothy J. Stasevich
Aug 5, 2022 2975 Views
Binding Affinity Measurements Between DNA Aptamers and their Virus Targets Using ELONA and MST
Gregory T. Pawel [...] Ana Sol Peinetti
Nov 5, 2022 3437 Views
Detection of Cytoplasmic and Nuclear Circular RNA via RT-qPCR
Ke-En Tan [...] Yat-Yuen Lim
Sep 5, 2023 3453 Views
Abstract
Given the scale of the ongoing COVID-19 pandemic, the need for reliable, scalable testing, and the likelihood of reagent shortages, especially in resource-poor settings, we have developed an RT-qPCR assay that relies on an alternative to conventional viral reverse transcriptases, a thermostable reverse transcriptase/DNA polymerase (RTX) (Ellefson et al., 2016). Here we show that RTX performs comparably to the other assays sanctioned by the CDC and validated in kit format. We demonstrate two modes of RTX use – (i) dye-based RT-qPCR assays that require only RTX polymerase, and (ii) TaqMan RT-qPCR assays that use a combination of RTX and Taq DNA polymerases (as the RTX exonuclease does not degrade a TaqMan probe). We also provide straightforward recipes for the purification of this alternative reagent RTX. We anticipate that in low resource or point-of-need settings researchers could obtain the available constructs and begin to develop their own assays, within whatever regulatory framework exists for them.
Keywords: CoronavirusBackground
While various virus detection methods have been implemented for the detection of SARS-CoV-2 infection, including a variety of molecular diagnostics and immunodiagnostic tests, the Reverse Transcriptase quantitative Polymerase Chain Reaction (RT-qPCR) remains the primary and most sensitive test for SARS-CoV-2 detection (D'Cruz et al., 2020; Tang et al., 2020). The primacy of RT-qPCR is in large measure because antibody-based tests as well as rapid nucleic acid diagnostic platforms, such as Abbott IDNow, often suffer from poor sensitivity, especially during early infection when viral loads are generally low in patients (Basu et al., 2020; D'Cruz et al., 2020). Given the importance of the early diagnosis in containing COVID-19 outbreak (Peck, 2020), the need for a rapid and scalable RT-qPCR setup is imminent. Unfortunately, there are increasing shortages of a wide variety of reagents necessary to scale RT-qPCR-based tests (Pettit et al., 2020). The main manufacturers of PCR platforms cannot scale-up the production of tests in sufficient quantities to supply resource-poor settings. Even were resource-poor settings to attempt to develop their own solutions, researchers, clinicians, and public health officials often lack the necessary reagents, including enzymes, to develop testing programs (Kavanagh et al., 2020).
Herein, we layout protocols for dye-based and TaqMan probe-based RT-qPCR assays for CDC-designed SARS-CoV-2 N gene tests (https://www.fda.gov/media/134922/download). We detail a one-enzyme RTX-based RT-qPCR protocol, as well as a two-enzyme RTX/Taq DNA polymerase-based RT-qPCR protocol. While the dye-based one enzyme and the TaqMan probe-based two enzyme RT-qPCR both exhibit comparable performances in the detection of viral RNA, the TaqMan probe-based assay exhibited lower background. In these, we compare our RTX enzyme preparations, for which we provide a detailed purification protocol, with commercially available reagents.
Our RTX-mediated protocols are as sensitive as those that rely on enzyme combinations or commercial enzymes, and can expedite SARS-CoV-2 tests by reducing the number of kits or key enzymes that must be purchased. These protocols can also form the basis for further assay simplification and democratization via the use of so-called ‘cellular reagents’: polymerase-overexpressing cells that can be directly added to molecular diagnostics assays without loss of sensitivity or specificity (Bhadra et al., 2018).
Materials and Reagents
For RTX polymerase-based RT-qPCR analysis
Barrier tips for pipets
Tris-HCl pH7.5 and 8.0 (Sigma-Aldrich, catalog number: T1503 )
(NH4)2SO4 (Sigma-Aldrich, catalog number: A4413 )
KCl (Sigma-Aldrich, catalog number: P3911 )
MgSO4 (Sigma-Aldrich, catalog number: M7506 )
D-glucose (Sigma-Aldrich, catalog number: 47829 )
1% Triton® X-100 (New England Biolabs, B9004S ), store at -20 °C
Forward PCR primers (Integrated DNA Technologies, Table 1) (Note a), store at -20 °C, 100 µM Stock
Reverse PCR primers (Integrated DNA Technologies, Table 1) (Note a), store at -20 °C, 100 µM Stock
TaqMan probes (Integrated DNA Technologies, Table 1) (Note a), store at -20 °C
20x EvaGreen solution (Biotium, #31000), store at 4 °C
4 mM deoxyribonucleotides (dNTP) (New England Biolabs, N0446S ), store at -20 °C
5 M Betaine (Sigma-Aldrich, B0300 ), store at 4 °C
20 U/µl SUPERase·In RNase Inhibitor (Thermo Fisher Scientific, AM2694 ), store at -20 °C
5 U/µl Taq DNA polymerase (New England Biolabs, M0267L ), store at -20 °C
0.2 mg/ml RTX polymerase (Note b) (purified by Ellington lab), store at -20 °C
Nuclease-free water (store at room temperature)
qPCR tubes or plates with optical covers(Note c) (Roche, 04729692001 ), store at room temperature
Cold-block or ice
10x RTX buffer (see Recipes)
10x ThermoPol buffer (see Recipes)
Notes:
Individual primer and probe stocks or pre-made primer-probe mixes for the CDC N1, N2, and N3 assays (Table 1) available from Integrated DNA Technologies may be used.
RTX polymerase with proofreading capability (RTX), and RTX polymerase without proofreading capability (RTX Exo-) have been compared.
LightCycler 96 real-time PCR machine and 96-well plates with optical plastic film cover designed for use with LightCycler platform were used for these experiments.
Table 1. CDC TaqMan RT-qPCR primers and probes for SARS-CoV-2a
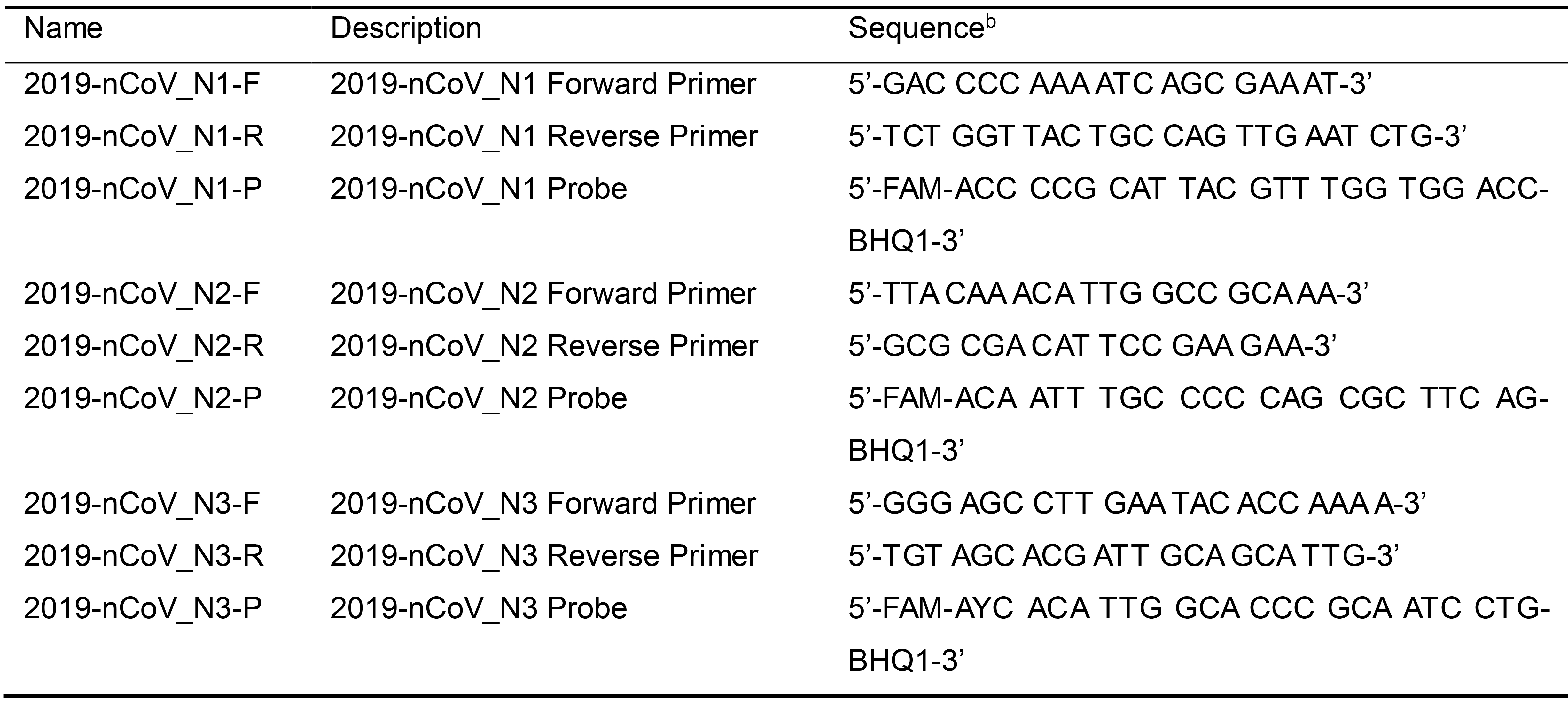
aPrimer and probe sequences adapted from https://www.cdc.gov/coronavirus/2019-ncov/lab/rt-pcr-panel-primer-probes.html; last accessed on May 6th, 2020. According to the CDC, oligonucleotide sequences are subject to future changes as the 2019-Novel Coronavirus evolves. Refer to the CDC website for the latest updates.
bFAM: 6-carboxyfluorescein; BHQ-1: Black Hole Quencher 1
For purification of RTX DNA polymerase
Sterile culture tubes and flasks
Centrifugation tubes (Nalgene, catalog number: 3115-0050 )
Sterile syringe filter 0.2 µm (PES, VWR, catalog number: 28145-501 )
RTX or RTX Exo- polymerase expression plasmid (Addgene, Plasmid #102787 and #102786 )
Competent E. coli T7 RNA polymerase-based protein expression strain, such as BL21 (DE3) (New England Biolabs, C2527H ), store at -80 °C
Antibiotics such as ampicillin or carbenicillin (Goldbio, catalog number: 4800-94-6 )
Superior BrothTM (Athena Enzyme SystemsTM , catalog number: 0105 ), store at room temperature
1 M Isopropyl-β-D-thiogalactoside (IPTG) (Sigma Aldrich), store at -20 °C
Ni-NTA agarose (Thermo Fisher Scientific, catalog number: 88223 ), store at 4 °C
Disposable protein purification columns (Thermo Fisher Scientific, catalog number: 29924 )
20,000 MWCO Dialysis cassette (Thermo Fisher Scientific, catalog number: 66012 )
HiTrapTM Heparin HP column (GE Healthcare, catalog number: 17-0406-01 )
Bis-Tris mini protein gel (Thermo Fisher, catalog number: NP0321BOX )
EDTA-free protease inhibitor tablet (Thermo Scientific, catalog number: A32965 )
Note: All chemicals were molecular biology or analytical grade and purchased from Sigma-Aldrich (see below for catalog number) unless otherwise indicated.
NaCl (Sigma-Aldrich, catalog number: S7653 )
Imidazole (Sigma-Aldrich, catalog number: I5513 )
Igepal CO-630 (Sigma-Aldrich, catalog number: 542334 )
MgSO4 (Sigma-Aldrich, catalog number: M7506 )
HEW Lysozyme (Sigma-Aldrich, catalog number: L6876 )
Tris (Sigma-Aldrich, catalog number: T1503 )
DTT (Sigma-Aldrich, catalog number: D9779 )
KCl (Sigma-Aldrich, catalog number: P3911 )
Tween-20 (Sigma-Aldrich, catalog number: P9416 )
Purification buffers for RTX DNA polymerase purification, store at 4 °C (see Recipes)
Resuspension Buffer
Equilibration Buffer
Lysis Buffer
Wash Buffer
Elution Buffer
Heparin Buffer A
Heparin Buffer B
Dialysis buffers for dialysis of RTX DNA polymerase, store at 4 °C (see Recipes)
Ni-NTA Dialysis Buffer
Heparin Dialysis Buffer
Final Dialysis Buffer
Equipment
Magnetic stir bar
Magnetic stirrer
Real-time PCR machine
Note: LightCycler96 real-time PCR machine from Roche was used for these experiments.
Refrigerated shaker incubator (New Brunswick Scientific, model: Innova44 )
Sonicator (Fisher Scientific, Sonic Dismembrator Model 500)
Centrifuge (Beckman Coulter, model: Avanti JXN-26 , Fixed angle)
Thermomixer (Eppendorf, model: Thermomixer C )
Fast protein liquid chromatography (FPLC) machine (GE Healthcare, model: AKTA pure )
Fraction collector (GE Healthcare, model: Fraction Collector F9-R )
Software
LightCycler96 software (Roche; available for download from https://lifescience.roche.com/en_us/brands/realtime-pcr-overview.html#software)
Procedure
Dye-based RT-qPCR reaction set up using only RTX DNA polymerase
Set up EvaGreen dye-containing RT-qPCR reactions for CDC SARS-CoV-2 N1, N2, and N3 assays as shown in Table 2. The three assays target three different regions within the nucleocapsid phosphoprotein (N) gene near the 3’-end of the ~30 kb viral genome. In particular, N1 primers (Table 1) amplify a 72 nt long region from position 28287 to 28358 of the genome (GenBank Accession No. MN985325.1). The N2 primers produce a 67 bp amplicon from the genomic position 29164 to 29230. Meanwhile, the N3 primers amplify a 72 nt long region from genomic position 28681 to 28752. RNA isolation should be performed before RT-qPCR.
Table 2. EvaGreen RT-qPCR reaction mix using only RTX DNA polymerase
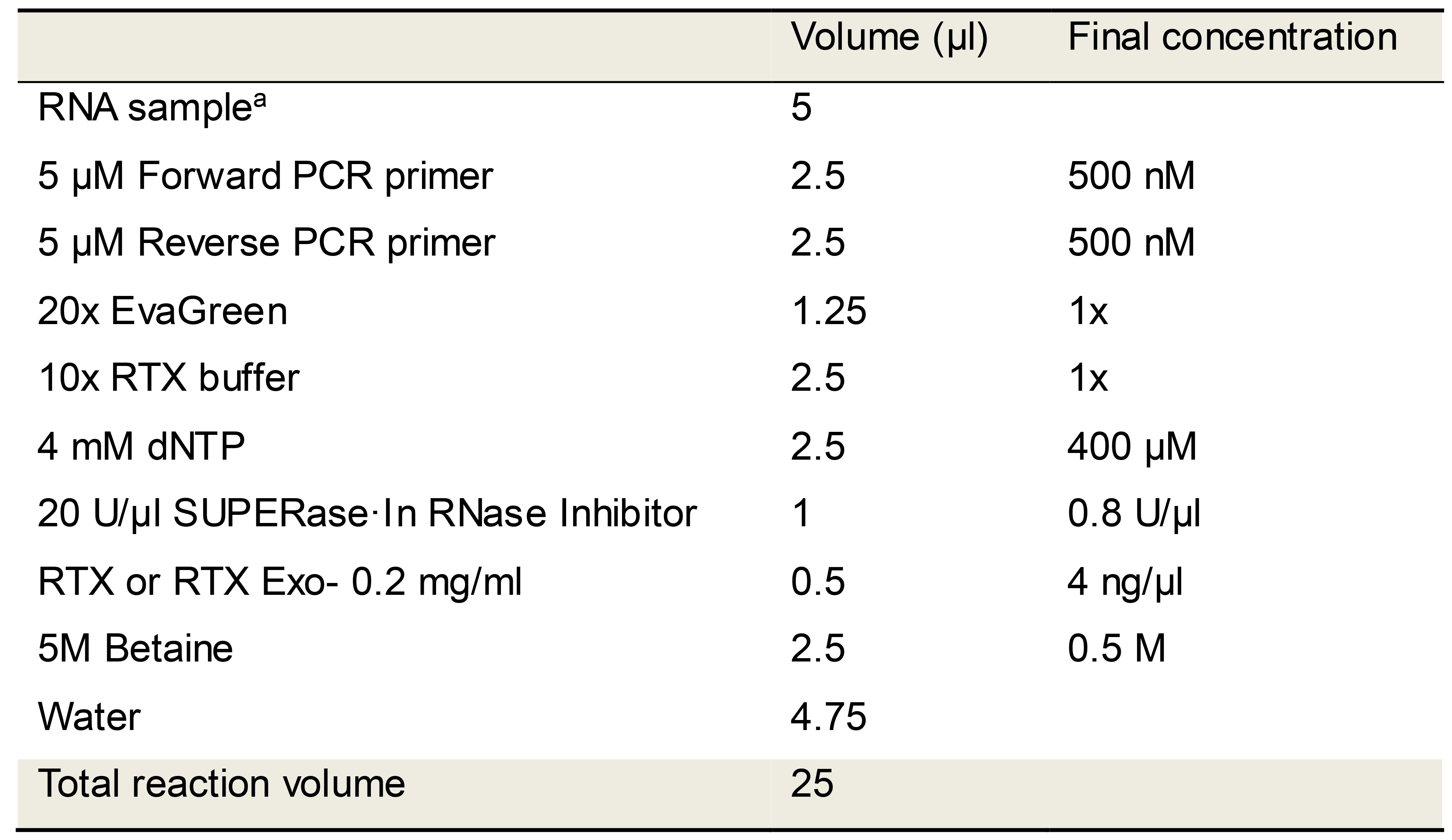
aFull length SARS-CoV-2 or MERS-CoV N synthetic RNAs were used in these studies. The RNA copies per reaction (100 to 100,000 copies) are indicated in the relevant figures. Patient saliva is reported to have median SARS-CoV-2 viral loads of 3.3 × 106 copies/ml (To et al., 2020).Assemble all reaction mixes on a cold-block and directly transfer to a real-time PCR machine programmed to cycle through the steps depicted in Table 3. Include Melt curve analysis steps after qPCR amplification in order to distinguish target amplicons from spurious products.
Table 3. RTX polymerase-based EvaGreen RT-qPCR cycling conditions
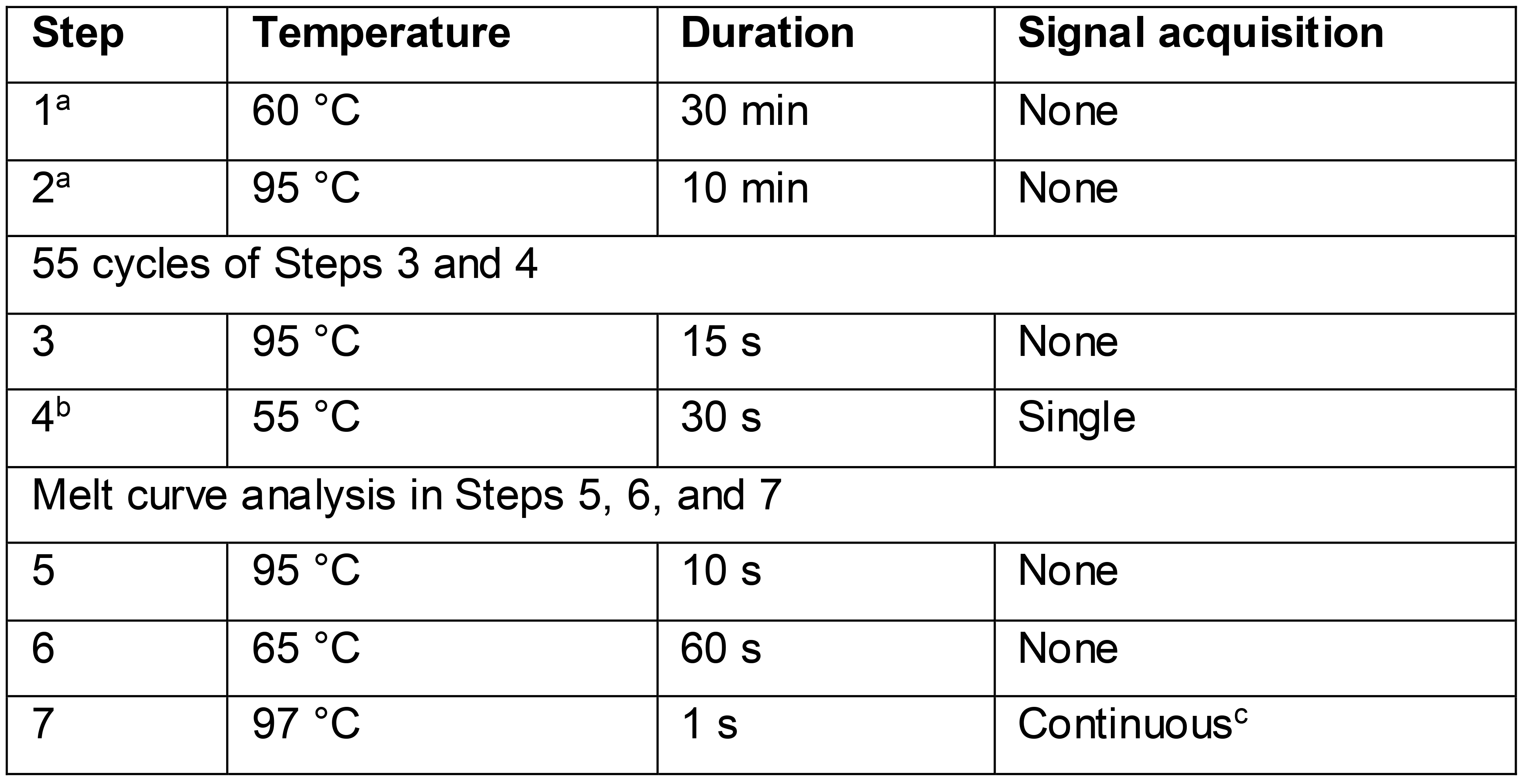
aAssay time may be reduced by shortening the reverse transcription step (step 1) to 15 min and the following denaturation step (step 2) to 2 min.
bSignal acquisition occurs in the FAM (fluorescein) channel during steps 4 and 7.
c20 measurements/°C temperature change
RTX polymerase-based TaqMan RT-qPCR reaction set up
In order to perform TaqMan RT-qPCR we developed a one-pot two enzyme system comprising RTX and Taq DNA polymerases. Set up RTX-based TaqMan RT-qPCR reactions for CDC SARS-CoV-2 N1, N2, and N3 assays as shown in Table 4.
Assemble all reaction mixes on a cold-block and directly transfer to a real-time PCR machine programmed to cycle through the steps depicted in Table 5.
Table 4. RTX polymerase-based TaqMan RT-qPCR reaction mix
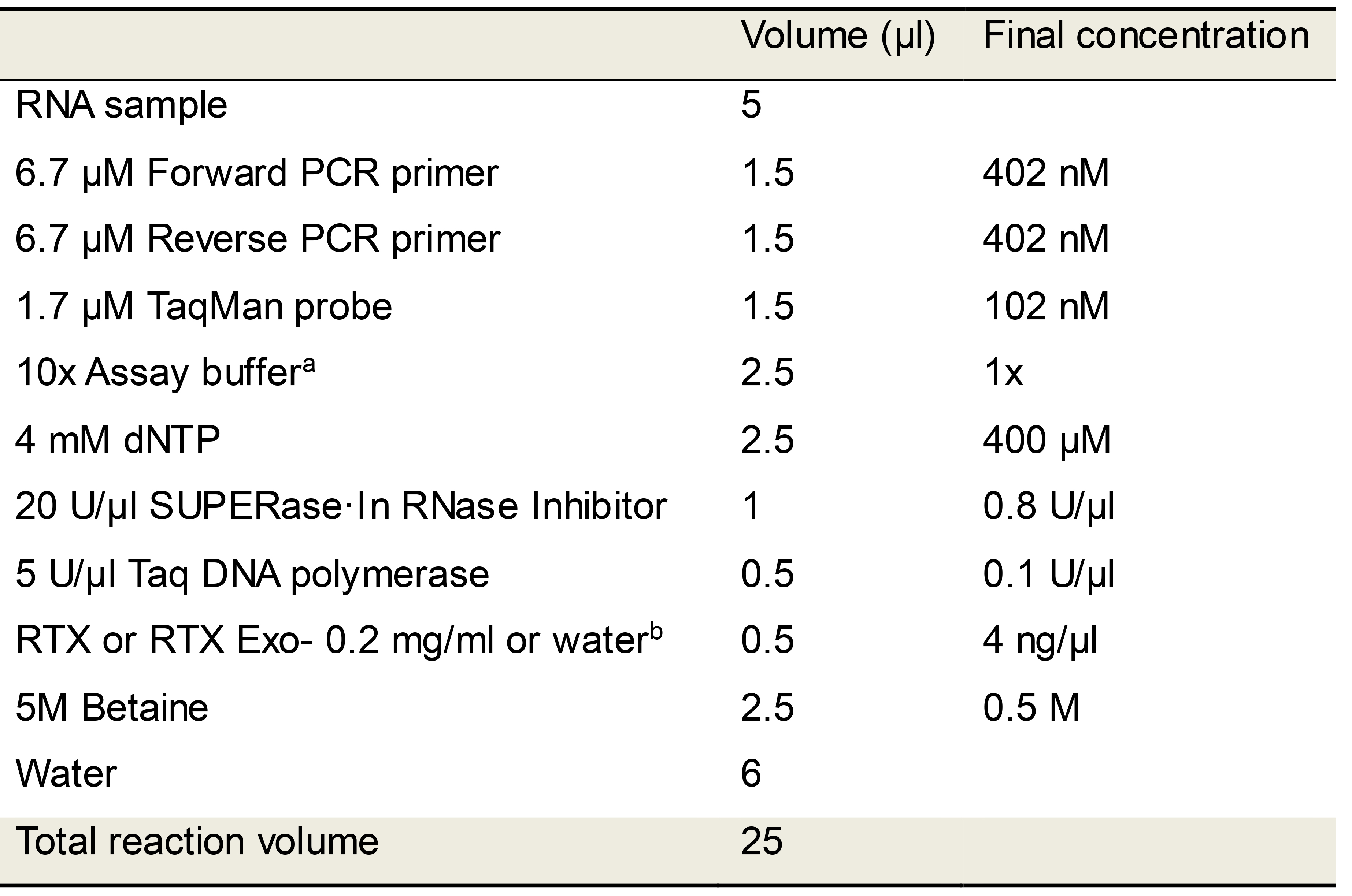
aAssay buffer: Duplicate TaqMan RT-qPCR reactions were performed either in 1x ThermoPol buffer or in 1x RTX buffer as indicated to determine optimum buffer conditions.
bControl reactions lacking RTX polymerase were set up using the same recipe except RTX was replaced with water.
Table 5. RTX polymerase-based TaqMan RT-qPCR cycling conditions
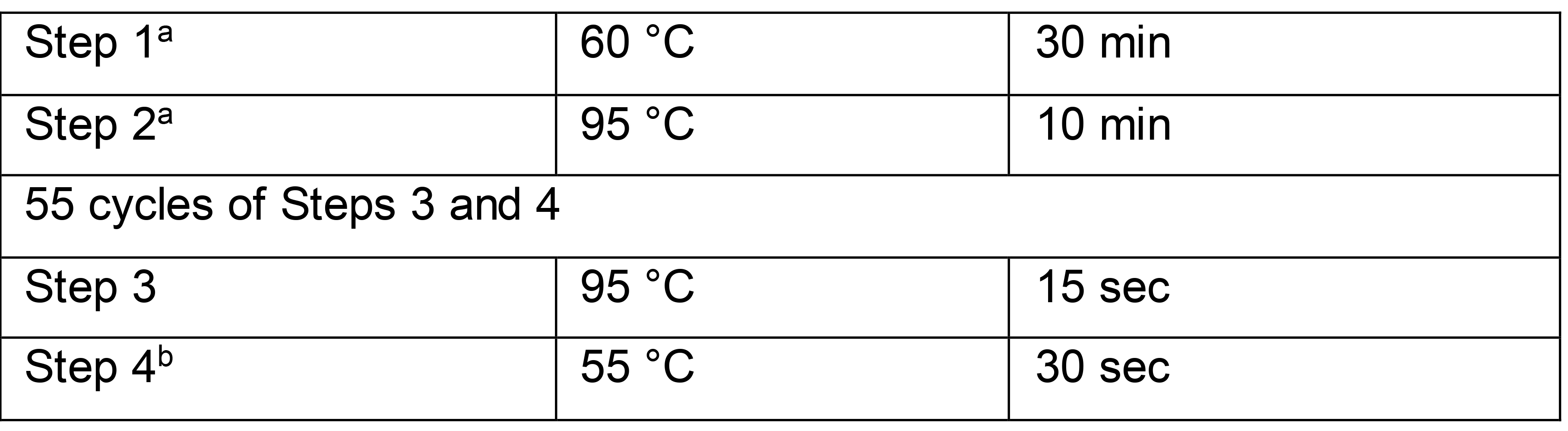
aAssay time may be reduced by shortening the reverse transcription step (step 1) to 15 min and the following denaturation step (step 2) to 2 min without compromising results.bSignal acquisition occurs in the FAM (fluorescein) channel during step 4.
SARS-CoV-2 RT-qPCR using commercial TaqPathTM 1-Step RT-qPCR Master Mix, CG
To perform CDC N1, N2, and N3 SARS-CoV-2 TaqMan RT-qPCR assays using commercial assay master mix, set up reactions using TaqPathTM 1-Step RT-qPCR Master Mix, CG (Thermo Fisher Scientific, Waltham, MA, USA) as shown in Table 6.
Assemble all reactions on a cold-block and then incubate at room temperature (25 °C) for 2 min prior to cycling on a real-time PCR machine using the steps indicated in Table 7.
Table 6. CDC N1, N2, N3 TaqMan RT-qPCR set up using commercial RT-qPCR master mix
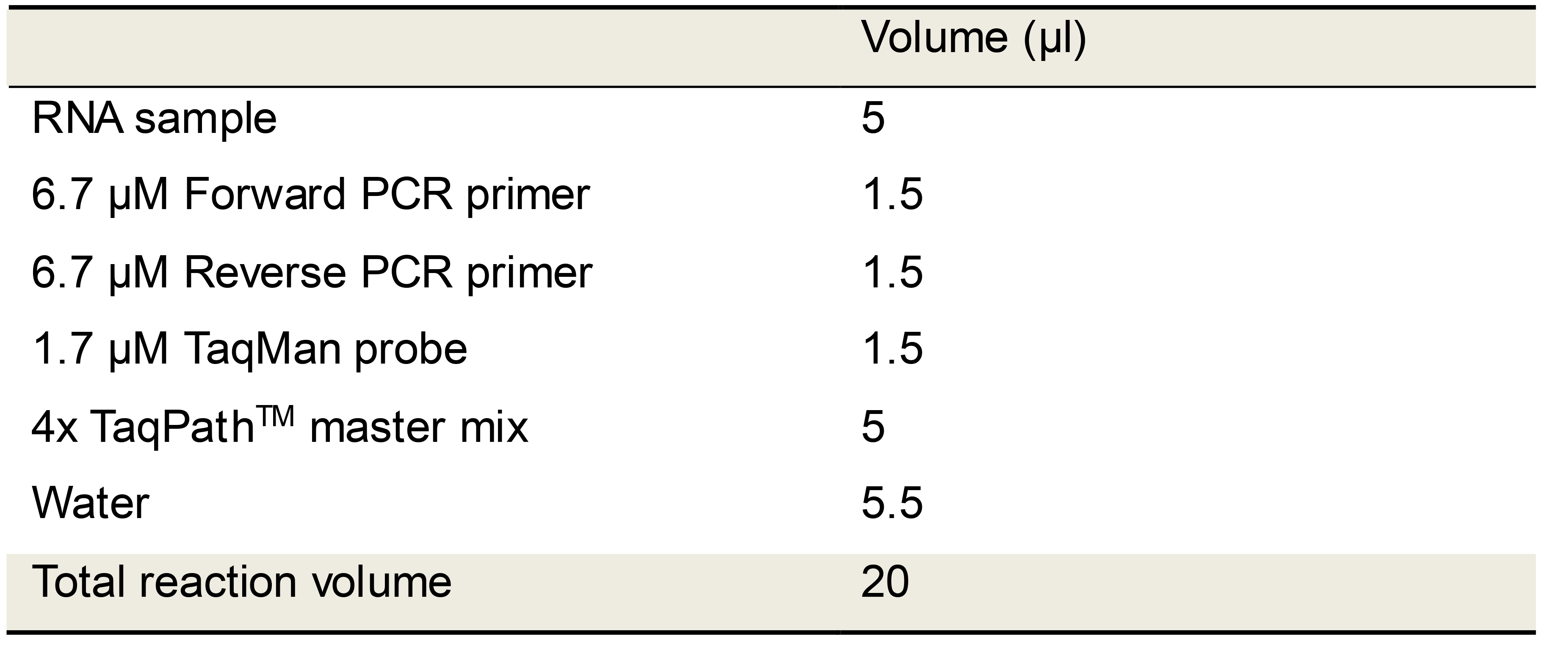
Table 7. TaqPathTM TaqMan RT-qPCR cycling conditions
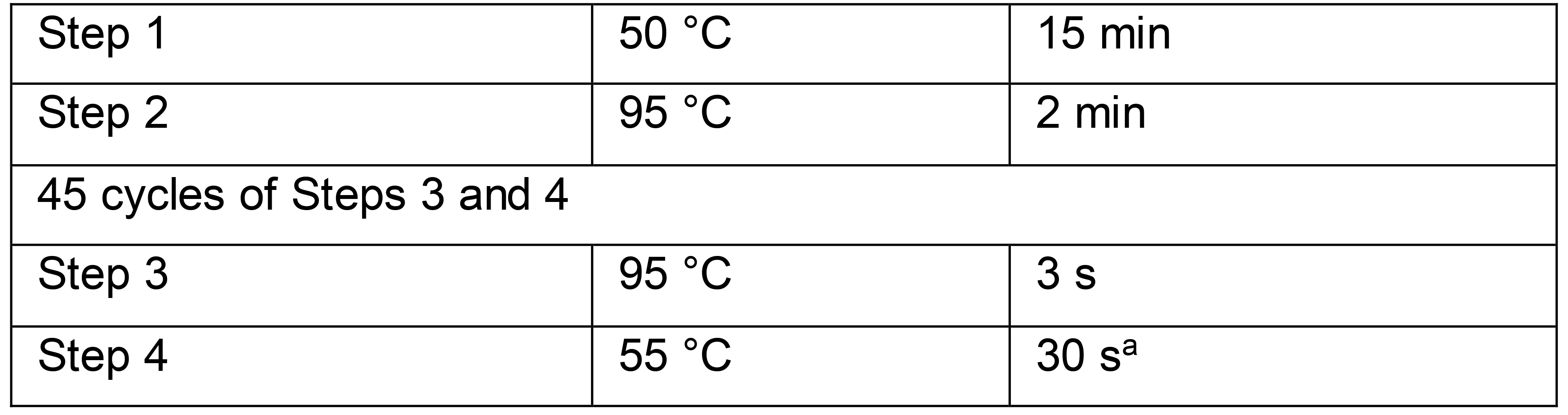
aSignal acquisition occurs in the FAM (fluorescein) channel during step 4.
Purification of RTX
Transform a T7-based E. coli expression strain, such as BL21(DE3), with a RTX expression plasmid that contains a carboxyl terminal six histidine tag for purification.
On the next day, pick an individual transformant colony and inoculate it into desired sterile media (10.Superior BrothTM) with appropriate antibiotics (e.g., Carbenicillin or Ampicillin 100 µg/ml) and 1-2% glucose Grow this starter culture overnight in shaking incubator at 37 °C.
On day three, inoculate 1 L sterile Superior BrothTM (Athena Enzyme SystemsTM) or other rich media with appropriate antibiotics (e.g., Carbenicillin or Ampicillin 100 µg/ml) in a 4 L Erlenmeyer flask with 250-350 μl of the overnight starter culture. Incubate at 37 °C, 220-250 rpm until the culture reaches OD600 of 0.4-0.7 (0.5-0.6 is ideal).
When the desired OD600 is reached, place flask(s) in 4 °C cold room for 30-45 min and set shaking incubator to 18 °C. Following cold incubation, induce expression of RTX by adding 1 ml of 1 M IPTG to each 1 L of culture. Place flask(s) in the 18 °C shaking incubator for 16-18 h.
Following overnight expression, harvest cells by centrifuging cultures at 4 °C, 5,000 x g for 20 min.
On the ice, resuspend cell pellet in 30 ml cold Lysis Buffer. Transfer the resuspended cell pellet to a small 50 ml beaker with a clean stir bar and place securely in an ice bath. With moderate stirring, sonicate the sample using 40% amplitude and 1 s ON/4 s OFF for 4 min total sonication time.
Centrifuge the resulting lysate at 4 °C, 35,000 x g for 30 min. Carefully transfer the supernatant to a clean ultracentrifugation tube, shake in thermomixer at 400 rpm, 85 °C for 10 min, and then place on ice for 10 min.
Centrifuge the heat-treated lysate at 4 °C, 35,000 x g for 30 min. Carefully transfer the supernatant to a clean tube and filter the clarified lysate using a 0.2 μm filter.
Prepare Ni-NTA agarose columns with a final column volume (CV) of 1 ml using 10 ml PierceTM disposable columns. Equilibrate column with 20 CV Equilibration Buffer.
Apply the clarified lysate from 1 L of expressed culture to a previously equilibrated column and collect the flow-through.
Wash column with 20 CV Equilibration Buffer and collect the flow-through. Wash column with 5 CV Wash Buffer and collect flow-through.
Elute RTX from a column with 5 ml Elution Buffer and transfer to a dialysis cassette with appropriate molecular weight cut-off. Dialyze eluate into 2 L of Ni-NTA Dialysis Buffer for 3-4 hours at 4 °C. Then dialyze eluate into a second 2 L of Ni-NTA Dialysis Buffer overnight at 4 °C.
Pass the dialyzed eluate over an equilibrated 5 ml heparin column (HiTrapTM Heparin HP) and elute along a sodium chloride linear gradient (100 mM to 2 M NaCl) generated from Heparin buffer A and Heparin buffer B mixed by an FPLC machine. The peak corresponding to RTX or its variants could be expected between 40-60% Buffer B.
Collect fraction tubes containing RTX peaks from the fraction collector. Pool the fractions and dialyze into 2 L Heparin Dialysis Buffer for 3-4 h. Then transfer dialysis cassettes into 2 L Final Dialysis Buffer overnight where it is expected that the protein sample volume will decrease significantly. Following the final dialysis, recover the protein sample and determine protein concentration before storing at -20 °C. The RTX stock is typically kept concentrated (5-10 mg/ml) in the Final Dialysis Buffer. If a lower concentration is required (e.g., 0.2 mg/ml working solution), Final Dialysis Buffer was used to dilute the RTX stock to the desired final concentration. An example of RTX purification is shown in Figure 7.
A similar protocol should work for Taq and other thermostable DNA polymerases, but may require different dialysis and storage buffers.
Data analysis
Analyze RT-qPCR data using LightCycler96 software.
Example data for dye-based RTX-only RT-qPCR
Representative results of SARS-CoV-2 RT-qPCR tests performed using synthetic RNA templates and only RTX or RTX Exo- DNA polymerases are depicted in Figures 1 and 2. These results demonstrate that RTX DNA polymerase alone, with or without proofreading capability, can support dye-based RT-qPCR analyses. In our hands, the full-length RTX DNA polymerase was slightly better than the Exo- version, especially in the N3 assay.
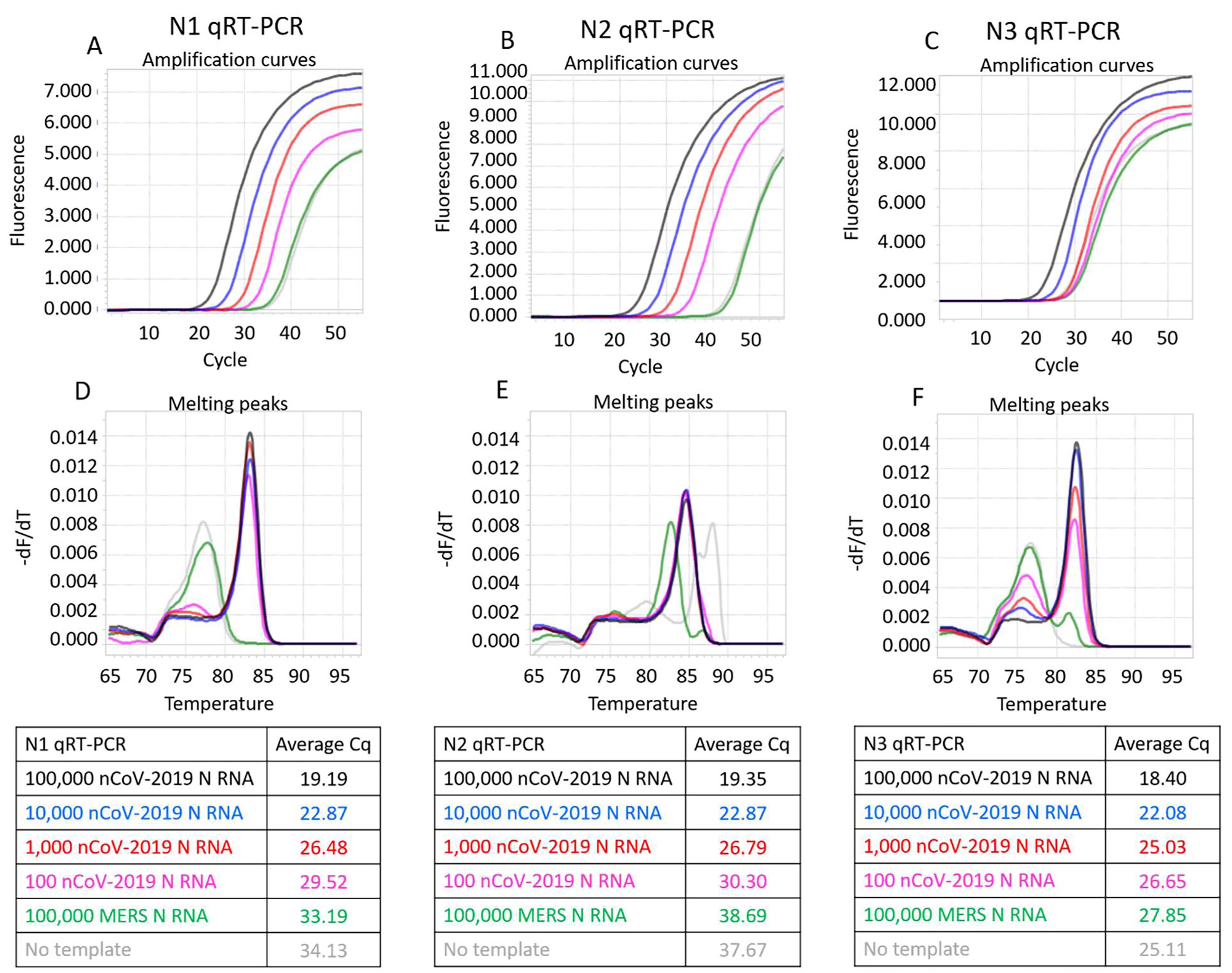
Figure 1. CDC SARS-CoV-2 N1, N2, and N3 RT-qPCR assays performed using indicated copies of synthetic RNA and RTX DNA polymerase. Panels A, B, and C depict N1, N2, and N3 assays measured in real-time using EvaGreen dye. Amplification curves from reactions containing 100,000 (black traces), 10,000 (blue traces), 1,000 (red traces), and 100 (pink traces) copies of SARS-CoV-2 synthetic N RNA are depicted. Negative control reactions either contained no templates (gray traces) or contained 100,000 copies of synthetic N RNA from MERS-CoV (green traces). Panels D, E, and F depict melting peaks of amplicons determined using the ‘Tm calling’ analysis in the LightCycler96 software. Note that the amplicons observed in negative controls were spurious, as indicated by their distinct melting temperatures compared to target-derived amplicons. Average Cq values of all assays are tabulated.
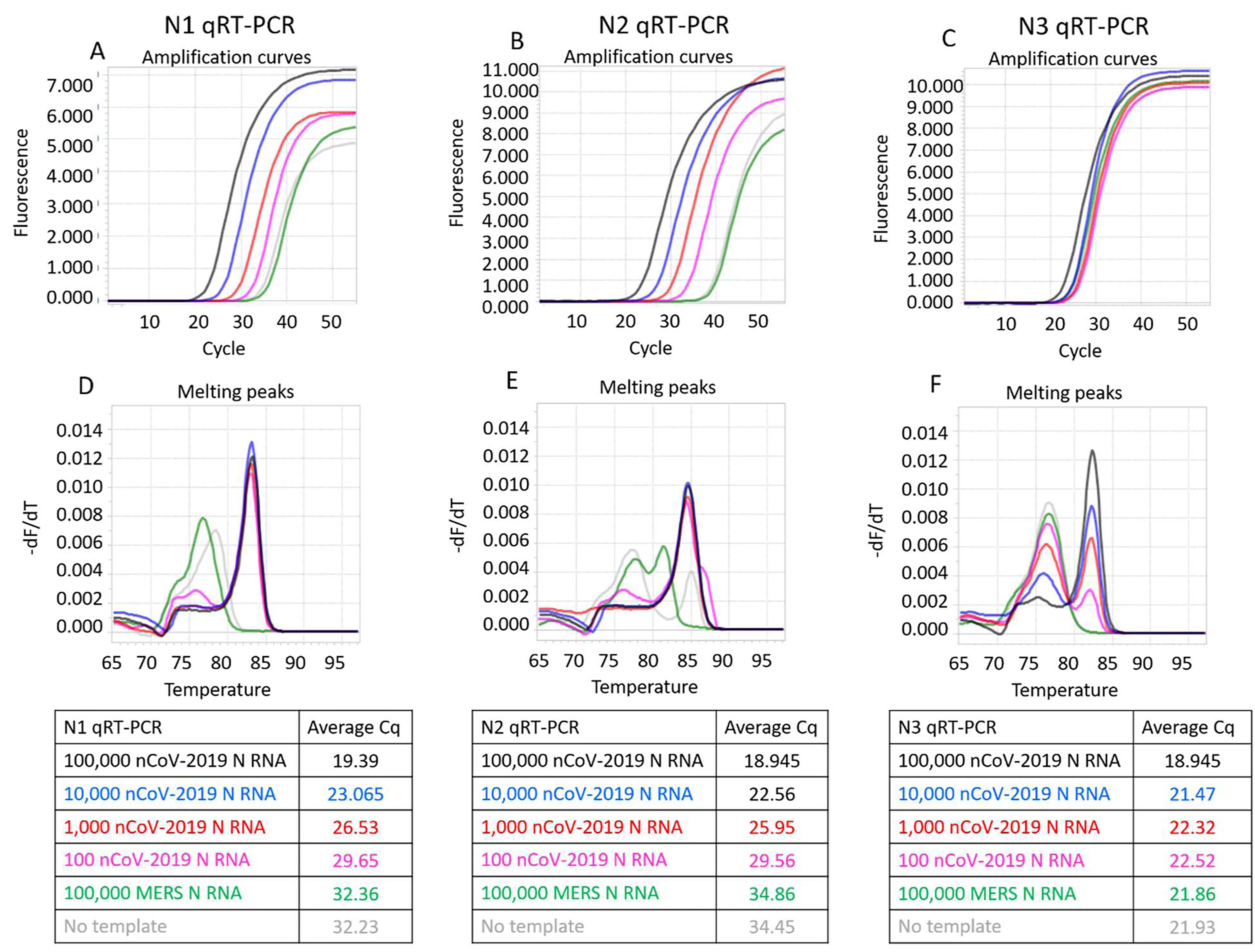
Figure 2. CDC SARS-CoV-2 N1, N2, and N3 RT-qPCR assays performed using indicated copies of synthetic RNA and RTX Exo- DNA polymerase. Panels A, B, and C depict N1, N2, and N3 assays measured in real-time using EvaGreen dye. Amplification curves from reactions containing 100,000 (black traces), 10,000 (blue traces), 1,000 (red traces), and 100 (pink traces) copies of SARS-CoV-2 synthetic N RNA are depicted. Negative control reactions either contained no templates (gray traces) or contained 100,000 copies of synthetic N RNA from MERS-CoV (green traces). Panels D, E, and F depict melting peaks of amplicons determined using the ‘Tm calling’ analysis in the LightCycler96 software. Average Cq values of all assays are tabulated.
Example data for RTX polymerase-based TaqMan RT-qPCR
Representative results of RTX polymerase-based SARS-CoV-2 TaqMan RT-qPCR tests performed using synthetic RNA templates are depicted in Figures 3 and 4. Both RTX and RTX Exo- when combined with Taq DNA polymerase are able to support one-pot TaqMan RT-qPCR assays for all three CDC SARS-CoV-2 assays. Under both buffer conditions tested, 1x ThermoPol versus 1x RTX buffer, RTX polymerase yielded more consistent amplification curves for all three CDC assays compared to RTX Exo- polymerase.
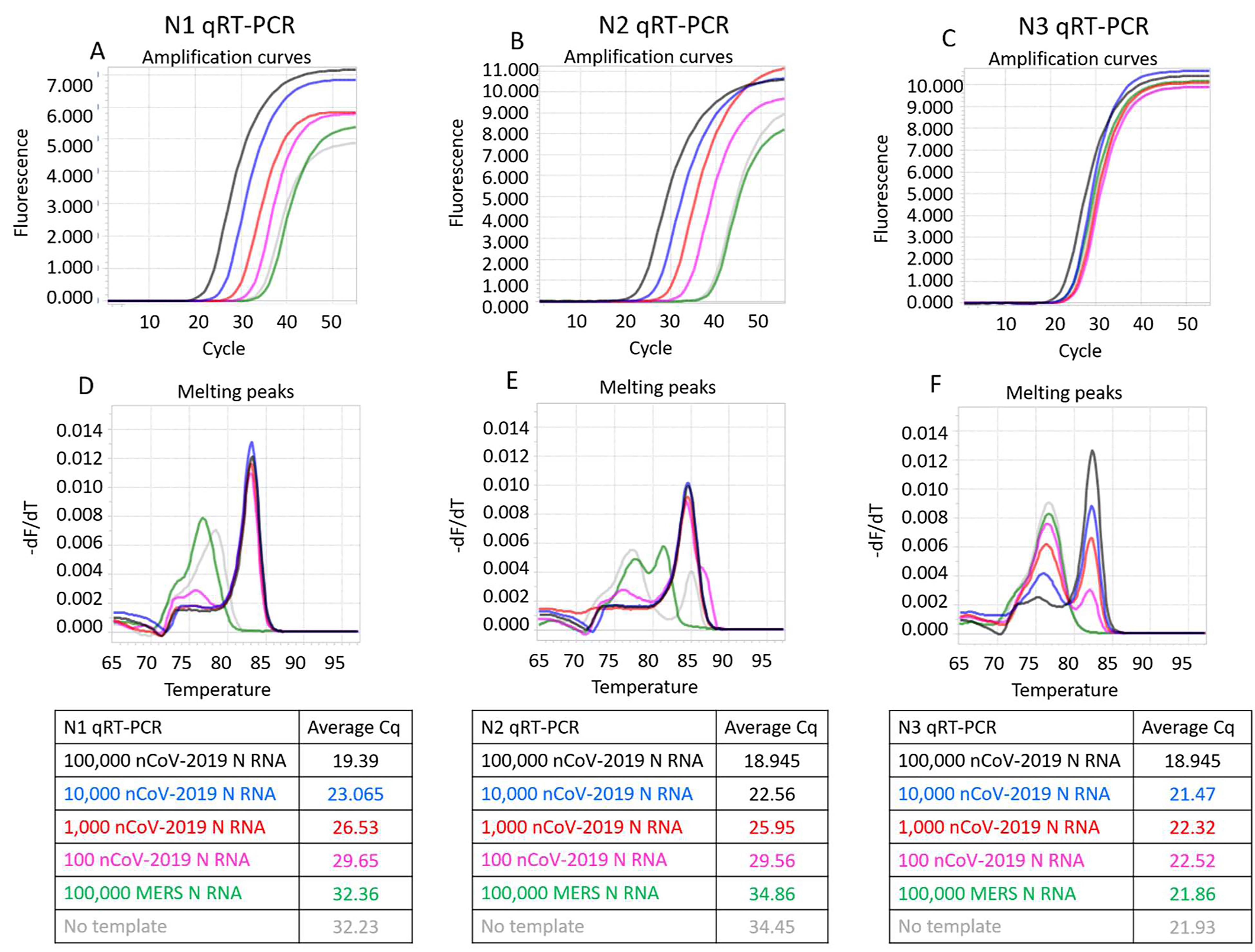
Figure 3. CDC SARS-CoV-2 N1, N2, and N3 TaqMan RT-qPCR assays performed in 1X ThermoPol buffer using indicated copies of synthetic RNA and RTX or RTX Exo- and Taq DNA polymerases.Panels A, B, and C depict TaqMan assays containing both RTX and Taq DNA polymerases. Panels D, E, and F depict TaqMan assays containing only Taq DNA polymerase. Panels G, H, and I depict TaqMan assays containing RTX Exo- and Taq DNA polymerases. Amplification curves from reactions containing 100,000 (black traces), 10,000 (blue traces), 1,000 (red traces), and 100 (pink traces) copies of SARS-CoV-2 synthetic N RNA are depicted. Negative control reactions either contained no templates (gray traces) or contained 100,000 copies of synthetic N RNA from MERS-CoV (green traces). Cq values of all assays are tabulated.
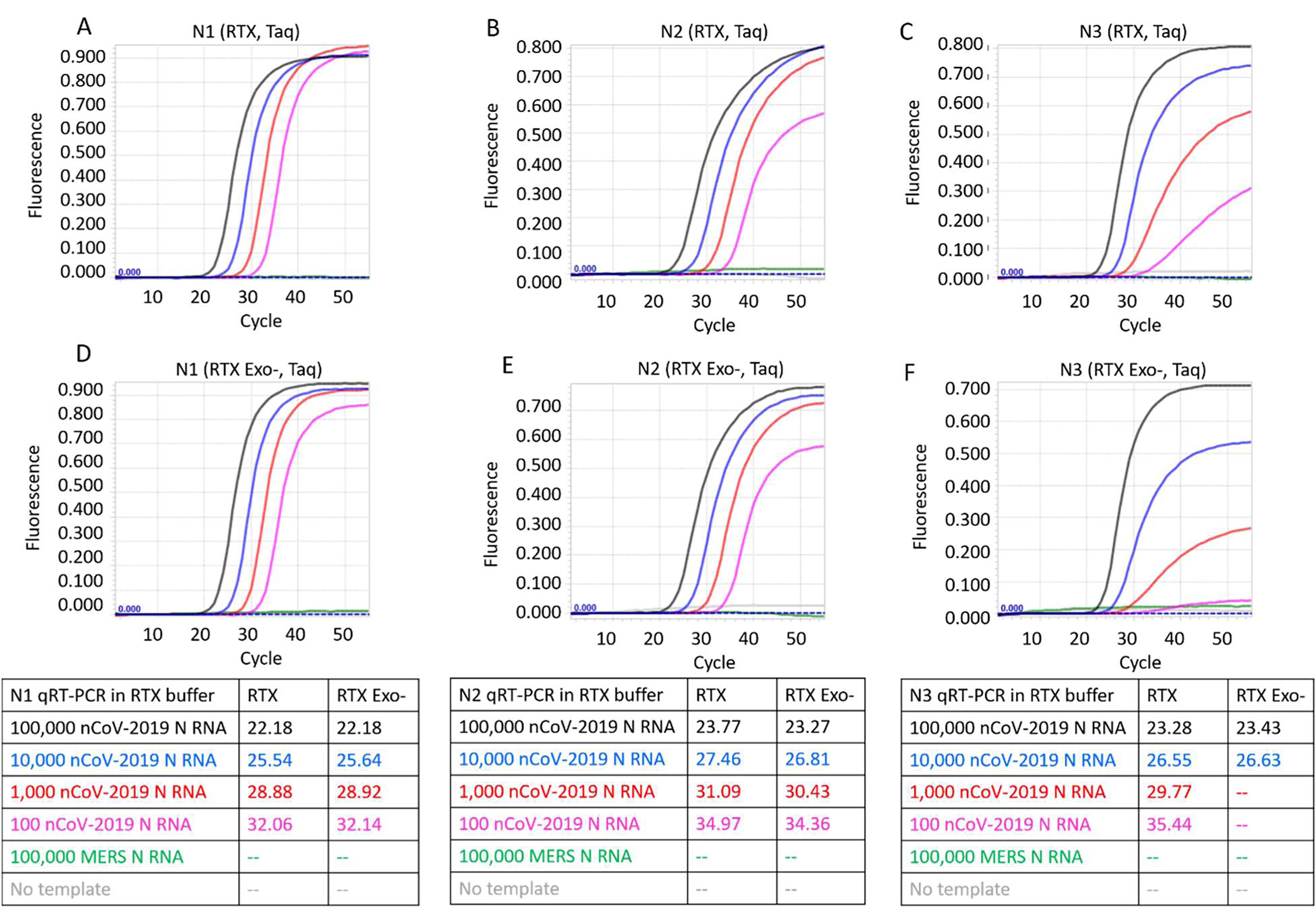
Figure 4. CDC SARS-CoV-2 N1, N2, and N3 TaqMan RT-qPCR assays performed in 1X RTX buffer using indicated copies of synthetic RNA and RTX or RTX Exo- and Taq DNA polymerases. Panels A, B, and C depict TaqMan assays containing RTX and Taq DNA polymerases. Panels D, E, and F depict TaqMan assays containing RTX Exo- and Taq DNA polymerases. Amplification curves from reactions containing 100,000 (black traces), 10,000 (blue traces), 1,000 (red traces), and 100 (pink traces) copies of SARS-CoV-2 synthetic N RNA are depicted. Negative control reactions either contained no templates (gray traces) or contained 100,000 copies of synthetic N RNA from MERS-CoV (green traces). Cq values of all assays are tabulated.
Example data for TaqPathTM TaqMan RT-qPCR
Figure 5 depicts typical results from CDC N1, N2, and N3 TaqMan qRT-PCR reactions performed using synthetic RNA and TaqPathTM commercial RT-qPCR mastermix.
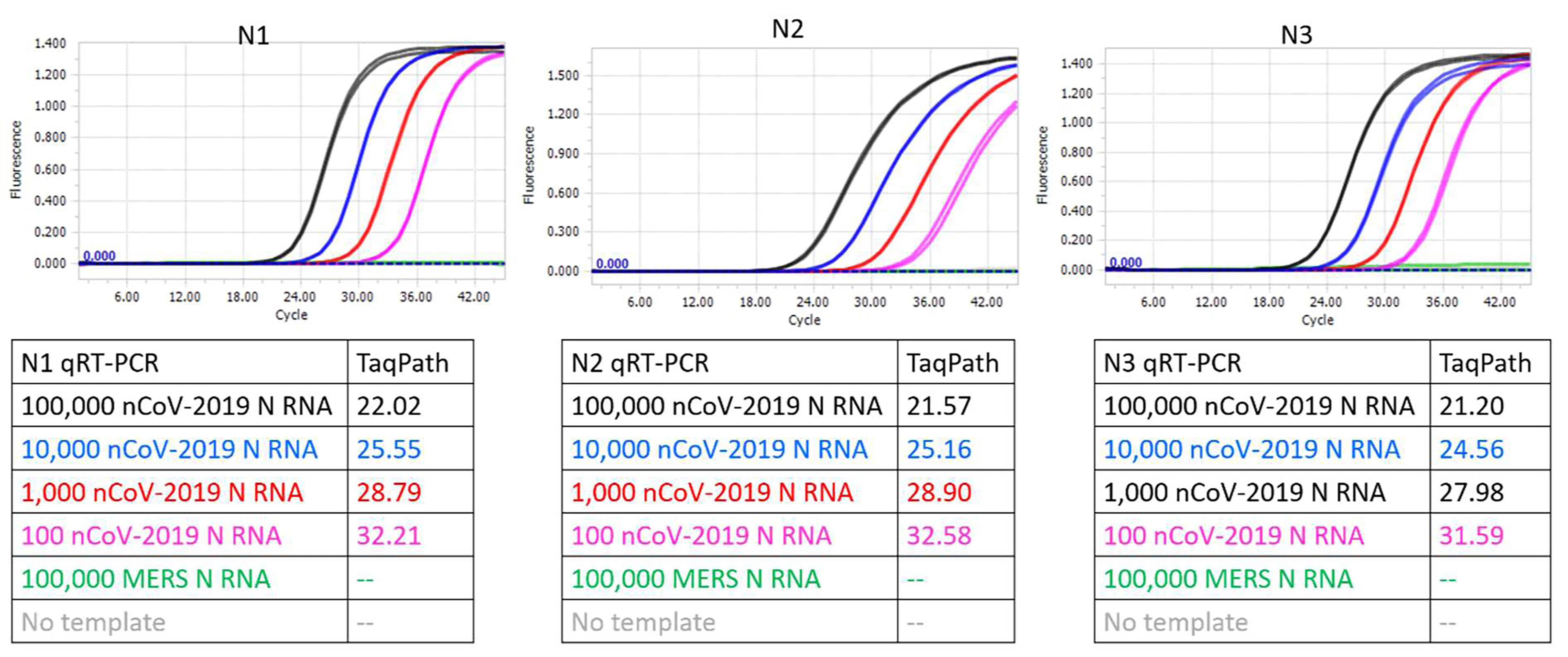
Figure 5. CDC SARS-CoV-2 N1, N2, and N3 TaqMan RT-qPCR assays performed using indicated copies of synthetic RNA and TaqPathTM commercial RT-qPCR mastermix. Amplification curves from reactions containing 100,000 (black traces), 10,000 (blue traces), 1,000 (red traces), and 100 (pink traces) copies of SARS-CoV-2 synthetic N RNA are depicted. Negative control reactions either contained no templates (gray traces) or contained 100,000 copies of synthetic N RNA from MERS-CoV (green traces). Cq values of all assays are tabulated.
The results with either a viral RT (in TaqPathTM commercial mastermix) or RTX are summarized in Figure 6. As can be seen, RTX-based TaqMan assays are of comparable sensitivity and specificity to the gold standard TaqPath assay. While all three RTX-based TaqMan assays performed in ThermoPol buffer consistently yielded Cq values, these were somewhat delayed compared to Cq values obtained with the commercial TaqPathTM Master Mix. In contrast, RTX-based TaqMan assays when executed in RTX buffer yielded Cq values that were closer to Cq values obtained with the commercial mastermix.
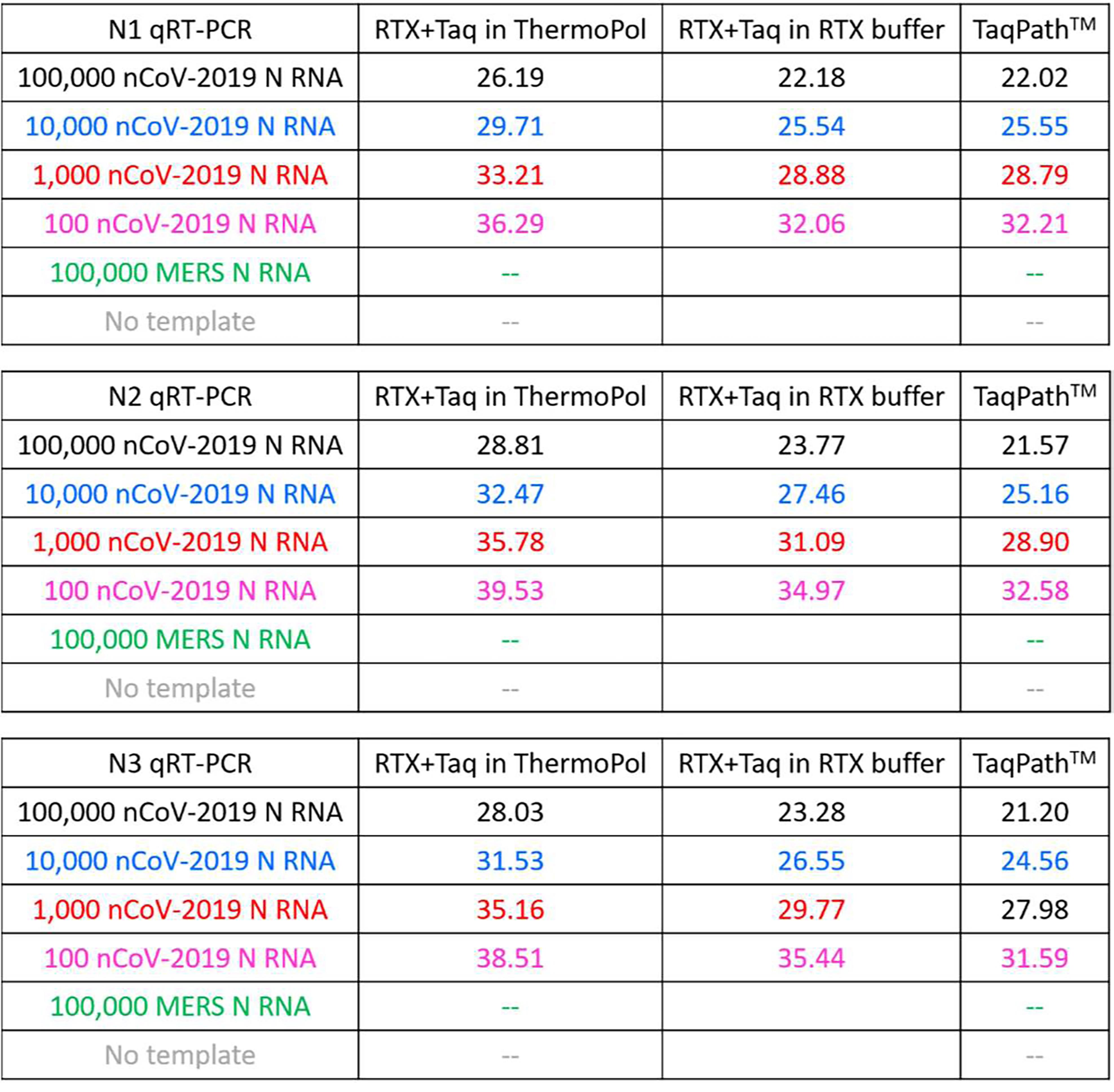
Figure 6. Comparison of Cq values of RTX-based and TaqPath-based SARS-CoV-2 TaqMan RT-qPCR assays
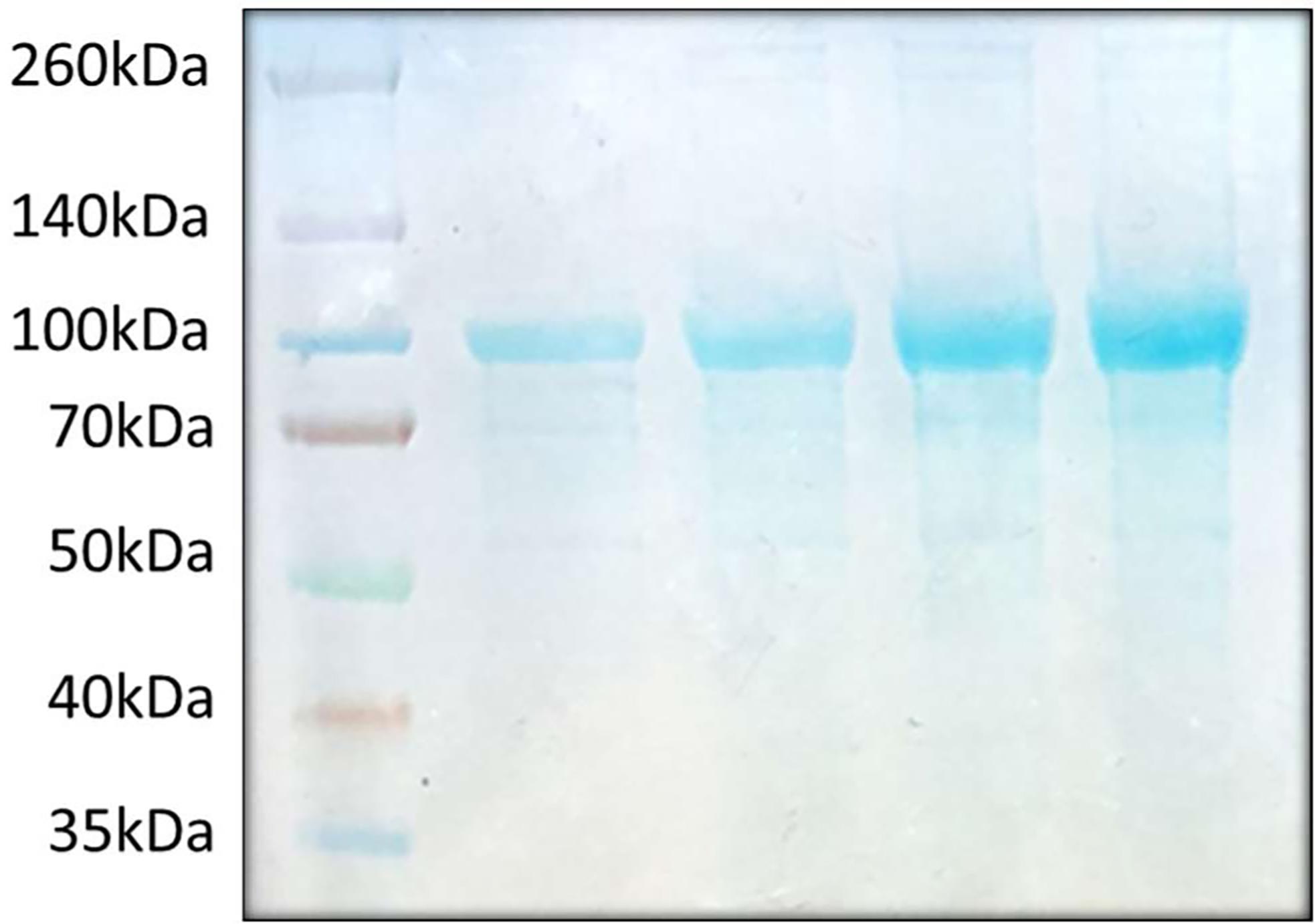
Figure 7. An example of RTX purification analyzed by SDS-PAGE. A total of ten to forty micrograms of purified RTX was developed on a NuPAGETM 4 to 12%, Bis-Tris mini protein gel. RTX bands appear near the 100kDa size marker. 1st lane: 10 µg, 2nd lane: 20 µg, 3rd lane: 30 µg, 4th lane: 40 µg.
Notes
Overall, RTX performs well either as a single enzyme for RT-qPCR with intercalating dyes, such as EvaGreen, or as the RT component of a TaqMan based assay. At the lowest RNA concentrations examined (100 copies), the gold standard TaqPath assay generally gave a signal at a Cq value of ca. 32. The more robust full-length RTX on its own actually performed slightly better, with Cq values typically from 29-30 (although the N3 primer set gives a higher signal and higher background). When used as a substitute for a viral RT, RTX is slightly less sensitive overall, with Cq values typically from 32-35.
By obtaining a RTX expression vector and purifying the thermostable reverse transcriptase it should prove possible to carry out RT-qPCR reactions with a sensitivity similar to that already observed for approved kits, with few or no false positive results. Plasmids and sequence information for 6xHis tagged RTX and RTX Exo- protein expression are available from Addgene (pET_RTX: https://www.addgene.org/102787/ and pET_RTX_(exo-): https://www.addgene.org/102786/), or can be obtained via https://reclone.org/.
Recipes
10x RTX buffer
600 mM Tris-HCl
250 mM (NH4)2SO4
100 mM KCl
20 mM MgSO4
pH 8.4, 25 °C
10x ThermoPol buffer
200 mM Tris-HCl
100 mM (NH4)2SO4
100 mM KCl
20 mM MgSO4
1% Triton® X-100
pH 8.8, 25 °C
Buffers for RTX DNA polymerase purification
Resuspension Buffer
50 mM Phosphate Buffer, pH 7.5
300 mM NaCl
20 mM imidazole
0.1% Igepal CO-630 (non-toxic Non-idet P40 equivalent)
5 mM MgSO4
Equilibration Buffer
50 mM Phosphate Buffer, pH 7.5
300 mM NaCl 20 mM imidazole
Lysis Buffer
30 ml Resuspension Buffer
1x EDTA-free protease inhibitor tablet
30-60 mg HEW Lysozyme
Wash Buffer
50 mM Phosphate Buffer, pH 7.5
300 mM NaCl
50 mM imidazole
Elution Buffer
50 mM Phosphate Buffer, pH 7.5
300 mM NaCl
250 mM imidazole
Heparin Buffer A
40 mM Tris-HCl, pH 7.5
100 mM NaCl
0.1% Igepal CO-630 (non-toxic Non-idet P40 equivalent)
Heparin Buffer B
40 mM Tris-HCl, pH 7.5
2 M NaCl
0.1% Igepal CO-630 (non-toxic Non-idet P40 equivalent)
Buffers for dialysis of RTX DNA polymerase
Ni-NTA Dialysis Buffer (2 L x2)
40 mM Tris-HCl, pH 7.5
100 mM NaCl
10 mM beta-mercapto ethanol (BME) or 1 mM 1,4-Dithiothreitol (DTT)
0.1% Igepal CO-630 (non-toxic Non-idet P40 equivalent)
Heparin Dialysis Buffer (2 L)
50 mM Tris-HCl, pH 8.0
50 mM KCl
0.1% Tween-20
0.1% Igepal CO-630 (non-toxic Non-idet P40 equivalent)
Final Dialysis Buffer (1 L)
50% Glycerol
50 mM Tris-HCl, pH 8.0
50 mM KCl
0.1% Tween-20
0.1% Igepal CO-630 (non-toxic Non-idet P40 equivalent)
1 mM DTT
Acknowledgments
We acknowledge the extraordinary contributions of the Schoggins lab at the University of Texas Southwestern Medical Center in making synthetic RNA available on short notice for these assays. We would like to acknowledge funding from the Promega Corporation (UTA18-000656), the National Institutes of Health (1R01EB027202-01A1), and the Welch Foundation (F-1654).
Competing interests
The Board of Regents of The University of Texas has licensed IP covering RTX to Promega Corporation.
References
- Basu, A., Zinger, T., Inglima, K., Woo, K. M., Atie, O., Yurasits, L., See, B. and Aguero-Rosenfeld, M. E. (2020). Performance of Abbott ID Now COVID-19 Rapid Nucleic Acid Amplification Test Using Nasopharyngeal Swabs Transported in Viral Transport Media and Dry Nasal Swabs in a New York City Academic Institution. J Clin Microbiol 58(8): e01136-20.
- Bhadra, S., Pothukuchy, A., Shroff, R., Cole, A. W., Byrom, M., Ellefson, J. W., Gollihar, J. D. and Ellington, A. D. (2018). Cellular reagents for diagnostics and synthetic biology. PLoS One 13(8): e0201681.
- D'Cruz, R. J., Currier, A. W. and Sampson, V. B. (2020). Laboratory Testing Methods for Novel Severe Acute Respiratory Syndrome-Coronavirus-2(SARS-CoV). Front Cell Dev Biol 8: 468.
- Ellefson, J. W., Gollihar, J., Shroff, R., Shivram, H., Iyer, V. R. and Ellington, A. D. (2016). Synthetic evolutionary origin of a proofreading reverse transcriptase. Science 352(6293): 1590-1593.
- Kavanagh, M. M., Erondu, N. A., Tomori, O., Dzau, V. J., Okiro, E. A., Maleche, A., Aniebo, I. C., Rugege, U., Holmes, C. B. and Gostin, L. O. (2020). Access to lifesaving medical resources for African countries: COVID-19 testing and response, ethics, and politics. Lancet 395(10238): 1735-1738.
- Peck, K. R. (2020). Early diagnosis and rapid isolation: response to COVID-19 outbreak in Korea. Clin Microbiol Infect 26(7): 805-807.
- Pettit, S. D., Jerome, K. R., Rouquie, D., Mari, B., Barbry, P., Kanda, Y., Matsumoto, M., Hester, S., Wehmas, L., Botten, J. W. and Bruce, E. A. (2020). All In': a pragmatic framework for COVID-19 testing and action on a global scale. EMBO Mol Med 12(6): e12634.
- Tang, Y. W., Schmitz, J. E., Persing, D. H. and Stratton, C. W. (2020). Laboratory Diagnosis of COVID-19: Current Issues and Challenges. J Clin Microbiol 58(6): e00512-20.
- To, K. K., Tsang, O. T., Yip, C. C., Chan, K. H., Wu, T. C., Chan, J. M., Leung, W. S., Chik, T. S., Choi, C. Y., Kandamby, D. H., Lung, D. C., Tam, A. R., Poon, R. W., Fung, A. Y., Hung, I. F., Cheng, V. C., Chan, J. F. and Yuen, K. Y. (2020). Consistent Detection of 2019 Novel Coronavirus in Saliva. Clin Infect Dis 71(15): 841-843.
Article Information
Copyright
© 2021 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Bhadra, S., Maranhao, A. C., Paik, I. and Ellington, A. D. (2021). A One-enzyme RT-qPCR Assay for SARS-CoV-2, and Procedures for Reagent Production. Bio-protocol 11(2): e3898. DOI: 10.21769/BioProtoc.3898.
Category
Microbiology > Pathogen detection
Molecular Biology > RNA > qRT-PCR
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










