- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Expression and Purification of Recombinant Skd3 (Human ClpB) Protein and Tobacco Etch Virus (TEV) Protease from Escherichia coli
Published: Vol 10, Iss 23, Dec 5, 2020 DOI: 10.21769/BioProtoc.3858 Views: 5109
Reviewed by: Toshitsugu FujitaMasato YanoAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
Monitoring Protein Stability In Vivo Using an Intein-Based Biosensor
John S. Smetana [...] Christopher W. Lennon
Apr 20, 2025 1619 Views
Endo-1,4-β-D-xylanase Assay Using Azo-Xylan and Variants Thereof
Luca Bombardi [...] Salvatore Fusco
Apr 20, 2025 1963 Views
Thermus thermophilus CRISPR Cas6 Heterologous Expression and Purification
Junwei Wei [...] Yingjun Li
Jul 20, 2025 2203 Views
Abstract
Skd3 (encoded by human CLPB) is a mitochondrial AAA+ protein comprised of an N-terminal ankyrin-repeat domain and a C-terminal HCLR-clade nucleotide-binding domain. The function of Skd3 has long remained unknown due to challenges in purifying the protein to high quality and near homogeneity. Recently we described Skd3 as a human mitochondrial protein disaggregase that solubilizes proteins in the mitochondrial intermembrane space. This protocol overcomes the challenges associated with purifying Skd3 and allows for in depth in vitro study of Skd3 activity. Tobacco etch virus (TEV) protease is required in the purification of Skd3. Thus, we also describe how to purify high quality TEV protease for use in the purification of Skd3, other purification protocols, and in vitro assays requiring TEV protease.
Keywords: Skd3Background
Skd3 is a mitochondrial AAA+ protein that has been implicated in the multisystem mitochondrial disorder, 3-methylglutaconic aciduria type VII (MGCA7) (Capo-Chichi et al., 2015; Kanabus et al., 2015; Saunders et al., 2015; Wortmann et al., 2015; Kiykim et al., 2016). The study of the biological function and the effect of these mutations on the activity of Skd3 had remained elusive due to limited ability to study Skd3 in vitro (Cupo and Shorter, 2020; Mroz et al., 2020). The challenges of studying Skd3 in vitro are threefold: (1) we have found that Skd3 is largely insoluble when expressed recombinantly in Escherichia coli. (2) Soluble fractions of Skd3 from E. coli were found to be highly proteolyzed. (3) Purification products were limited in quantity, with low microgram yields. With the protein purification scheme described herein, we are able to reproducibly purify Skd3 (and Skd3 variants) in milligram quantities to crystallography-grade purity (> 95%), thus overcoming the previous limitations of studying Skd3 in vitro (Cupo and Shorter, 2020). By harnessing biochemical assays and this purification scheme, we have shown that Skd3 is a potent mitochondrial protein disaggregase that chaperones the mitochondrial intermembrane space and maintains the solubility of proteins such as HAX1 and MICU2 (Cupo and Shorter, 2020). Furthermore, we have found that MGCA7-linked mutations disrupt Skd3 disaggregase activity (but not necessarily ATPase activity) in a manner that correlates with the clinical severity of the mutation (Cupo and Shorter, 2020).
The purification scheme for Skd3 requires large (milligram) quantities of high-purity tobacco etch virus (TEV) protease. Commercial purchase of TEV protease for in depth study of Skd3 activity is expensive. Thus, we also describe an in-house purification scheme for TEV protease. Similar to the Skd3 purification scheme, this protocol yields milligram quantities of highly active and > 95% pure TEV protease that can be used for Skd3 purification, other purification schemes, or in vitro assays that require TEV protease ( Mann et al., 2019; Cupo and Shorter, 2020).
Materials and Reagents
Pipette tips (USA Scientific TipOne)
Amicon Ultra-15 Centrifugal Filter Unit (15 ml) (Millipore Sigma, catalog number: UFC905024 )
Poly-Prep® Chromatography Columns (Bio-Rad, catalog number: 7311550 )
Greiner Bio-OneTM Polypropylene Conical Bottom Test Tubes (15 ml) (Fisher Scientific, catalog number: 10384601 )
Greiner Bio-OneTM Polypropylene Conical Bottom Test Tubes (50 ml) (Fisher Scientific, catalog number: 10711212 )
Protein LoBind Tubes (Eppendorf, catalog number: 022431081 )
Spectra/Por 6 Dialysis Tubing (8kD MWCO, 40 mm width) (Repligen, catalog number: 132584 )
Steriflip-GP Sterile Centrifuge Tube Top Filter Unit (50 ml) (Millipore Sigma, catalog number: SCGP00525 )
Sterile 100 mm x 15 mm Polystyrene Petri Dish (Fisher Scientific, catalog number: FB0875712 )
FisherbrandTM Disposable Borosilicate Glass Tubes with Plain End (6 ml) (Fisher Scientific, catalog number: 14-961-26 )
Escherichia coli BL21-CodonPlus (DE3)-RIL Competent Cells (Aligent, catalog number: 230245 )
pHis-TEV plasmid (available upon request)
pMALC2_MBP-PARLSkd3 or pMALC2_MBP-MPPSkd3 plasmid (available upon request)
SpectrumTM Labs Spectra/PorTM 6 8000 D MWCO Standard RC Pre-wetted Dialysis Kits 5.1 ml/cm vol./Length (Fisher Scientific, catalog number: 08-700-187 )
2-Mercaptoethanol (BME) (Bio-Rad, catalog number: 1610710 , CAS number: 60-24-2)
Adenosine 5′-triphosphate (ATP) disodium salt hydrate, Grade II, ≥98.5% (HPLC), crystalline, from microbial (Millipore Sigma, catalog number: A3377-100G , CAS number: 34369-07-8)
Coomassie Brilliant Blue G-250 (Bio-Rad, catalog number: 1610406 )
Coomassie Brilliant Blue R-250 (Bio-Rad, catalog number: 1610400 )
Agar (Fisher Scientific, catalog number: BP1423-500 , CAS number: 9002-18-0)
Ampicillin Sodium Salt (Fisher Scientific, catalog number: BP1760-25 , CAS number: 69-52-3)
Amylose Resin (New England Biolabs, catalog number: E8021L )
BL21-CodonPlus (DE3)-RIL competent cells (Aligent, catalog number: 230245 )
Chloramphenicol (Fisher Scientific, catalog number: BP904-100 , CAS number: 56-75-7)
cOmpleteTM, Mini, EDTA-free Protease Inhibitor Cocktail Tablets (Millipore Sigma, catalog number: 11836170001 )
cOmpleteTM Protease Inhibitor Cocktail Tablets (Millipore Sigma, catalog number: 11697498001 )
Dextrose Anhydrous (glucose) (Fisher Scientific, catalog number: BP350-1, CAS number: 50-99-7 )
DL-Dithiothreitol (DTT) (MP Biomedicals, catalog number: 100597 , CAS number: 3483-12-3)
Glycerol (Fisher Scientific, catalog number: BP229-4 , CAS number: 56-81-5)
HEPES (Fisher Scientific, catalog number: BP310-1 , CAS number: 7365-45-9 )
Hydrochloric acid, ACS reagent, 37% (Millipore Sigma, catalog number: 320331-500ML , CAS number: 7647-01-0)
Imidazole (Fisher Scientific, catalog number: O3196-500 , CAS number: 288-32-4)
Isopropyl β-D-1-thiogalactopyranoside (IPTG), Dioxane-Free, High Purity (Millipore, catalog number: 420322-25GM, CAS number: 367-93-1 )
Luria Broth (LB) [Miller's LB Broth] 1 L Buffered Capsules (RPI Corporation, catalog number: L24045-5000.0 )
Lysozyme from chicken egg white, lyophilized powder, protein ≥ 90%, ≥ 40,000 units/mg protein (Millipore Sigma, catalog number: L6876-25G , CAS number: 12650-88-3)
Magnesium chloride hexahydrate, ReagentPlus®, ≥99.0% (Millipore Sigma, catalog number: M0250-1KG , CAS number: 7791-18-6)
Ni-NTA Agarose Resin (Qiagen, catalog number: 30250 )
Pepstatin from Streptomyces species (Millipore Sigma, catalog number: 11359053001 , CAS number: 26305-03-3)
Potassium Chloride (Fisher Scientific, catalog number: BP366-1 , CAS number: 7447-40-7)
Potassium hydroxide, ACS reagent, ≥ 85%, pellets (Millipore Sigma, catalog number: 221473-1KG , CAS number: 1310-58-3)
Sodium Chloride (Fisher Scientific, catalog number: BP358-212 , CAS number: 7647-14-5)
Sodium hydroxide, ACS reagent, ≥ 97.0%, pellets (Millipore Sigma, catalog number: 221465-2.5KG , CAS number: 1310-73-2)
Stericup Quick Release-GP Sterile Vacuum Filtration System (Millipore Sigma, catalog number: S2GPU11RE )
Tris (Bio-Rad, catalog number: 1610719 , CAS number: 77-86-1)
TEV protease (purified in house, described below)
Agar Plates (see Recipes)
Growth Media (see Recipes)
Lysozyme Stock Solution (see Recipes)
TEV Lysis Buffer (see Recipes)
TEV Wash Buffer (see Recipes)
TEV Elution Buffer (see Recipes)
TEV Dialysis Buffer (see Recipes)
TEV Low-Salt IEX Buffer (see Recipes)
TEV High-Salt IEX Buffer (see Recipes)
Skd3 Lysis Buffer (see Recipes)
Skd3 Wash Buffer (see Recipes)
Skd3 ATP Wash Buffer (see Recipes)
Skd3 Elution Buffer (see Recipes)
Skd3 Sizing Buffer (see Recipes)
ATP Stock (see Recipes)
Equipment
Pipettes (Eppendorf)
PYREX Baffled Shaker Erlenmeyer Flask (250 ml) (Corning, catalog number: 4444-250 )
PYREX® 2,800 ml Fernbach-Style Culture Flask with Baffles (Corning, catalog number: 4423-2XL )
New BrunswickTM Innova® 44/44R Shaker (or equivalent 250 RPM shaker)
Polycarbonate Baffled Erlenmeyer Flasks (1 L) (Corning, catalog number: 431402 )
SorvallTM RC3BP Plus Low-Speed Centrifuge (ThermoFisher, catalog number: 75007530 ) (or equivalent low-speed centrifuge for 1 L cultures)
NalgeneTM PPCO 1 L Centrifuge Bottles (ThermoFisher, catalog number: 3120-1010 )
Misonix Sonicator® 3000 Ultrasonic Cell Disruptor (Misonix, catalog number: K-04711-79 ) (or equivalent sonicator)
NalgeneTM Oak Ridge High-Speed PPCO Centrifuge Tubes (30 ml) (ThermoFisher, catalog number: 3119-0030 )
SorvallTM RC 6 Plus Superspeed Centrifuge (ThermoFisher, catalog number: 36-101-0816) fitted with SS-34 Fixed Angle Rotor (ThermoFisher, catalog number: 28020TS ) (or equivalent high-speed centrifuge)
Centrifuge 5810 R (Eppendorf, catalog number: 022625101 ) (or equivalent low-speed centrifuge for 50 ml conical tubes)
LabnetTM LabRollerTM II w/Carousel for 50 ml Tubes (Labnet, catalog number: H5000 )
Econo-Column® Chromatography Columns (Bio-Rad, catalog number: 7372512 )
SnakeSkinTM Dialysis Clips (ThermoFisher, catalog number: 68011 )
GE Healthcare ÄKTA PurifierTM 10 FPLC with ÄKTA Sample Pump P960 (Millipore Sigma, catalog number: GE18672700 ) (or equivalent Fast Protein Liquid Chromatography [FPLC] system with UV-Vis detection and fraction collector)
GE Healthcare HiPrepTM 26/60 Sephacryl S-300 HR Size-Exclusion Column (Millipore Sigma, catalog number: GE17-1196-01 )
GE Healthcare HiTrap® SP XL Column (Cytiva Life Sciences, catalog number: 17516001 )
Software
FIJI ImageJ software
Procedure
Purification of TEV protease
Note: The following protocol is for purifying TEV from a 6 L culture volume of bacteria which yields approximately 100 mg of > 95% pure protein.
Bacterial cell transformation, induction, and harvest
Transform ~100 ng of pHis-TEV plasmid (ampicillin resistant; available from the authors) into Escherichia coli BL21 (DE3) RIL cells (chloramphenicol resistant) and plate on Agar Plates supplemented with ampicillin and chloramphenicol. Grow transformants at 37 °C overnight.
The next day, harvest the transformants by scraping the entire plate into a starter culture of 90 ml Growth Media supplemented with ampicillin and chloramphenicol.
Grow starter culture at 37 °C with 250 RPM shaking for 2 h.
Add 1 L Growth Media to each of 6 PYREX® 2,800 ml Fernbach-Style Culture Flask with Baffles. Add 10 ml starter culture to each 1 L baffled shaker flask.
Grow cultures at 37 °C with 250 RPM shaking until cultures reach OD600 = 0.7 (approximately 1-3 h).
Once cultures reach OD600 = 0.7, rapidly cool shakers and media to 15 °C (approximately 30-60 min).
Once temperature is equilibrated to 15 °C, supplement 1 mM IPTG and continue shaking at 15 °C and 250 RPM for 16 h.
Spin cells in 1 L centrifuge bottles at 4,658 x g for 25 min to pellet the cells. Discard the supernatant.
Bacterial cell lysis
Note: All steps should be performed on ice/at 4 °C with pre-chilled tubes unless otherwise indicated.
Resuspend the bacterial cell pellet in 72 ml ice-cold TEV Lysis Buffer and transfer to 50 ml conical tubes.
Add 6 ml of 20 mg/ml lysozyme to the TEV Lysis Buffer. Incubate cells on ice with lysozyme for 30 min.
Sonicate sample with one 30 s cycle and three 15 s cycles at power level 9. Keep conical tubes on ice while sonicating and let lysate cool on ice for 5 min between sonication cycles.
Note: It is extremely important to keep the lysate cool at this step as the sonication can heat the lysates and denature the protein.
Transfer lysate into NalgeneTM Oak Ridge High-Speed PPCO Centrifuge Tubes and centrifuge lysate at 30,597 x g and 4 °C for 20 min to clear the lysates.
Purification against N-terminal His tag
Equilibrate 21 ml Ni-NTA resin with two bed volumes (BV) of ice-cold TEV Lysis Buffer in 50 ml conical tubes. Centrifuge at 796 x g and 4 °C for 5 min. Discard supernatant. Repeat 3-4 times.
Note: Higher centrifuge speeds at this step will damage the amylose resin.
Add cleared bacterial cell lysates to equilibrated Ni-NTA resin. Nutate slowly at 4 °C for 2 h.
Note: If lysates are nutated too rapidly, bubbles will form in the buffer which can drive protein denaturation.
Centrifuge at 796 x g and 4 °C for 5 min. Discard supernatant.
Add two BV of ice-cold TEV Wash Buffer to the resin and gently but fully resuspend the resin in the TEV Wash Buffer. Centrifuge at 796 x g and 4 °C for 5 min. Discard supernatant. Repeat until the resin has been washed with ~25 BV of TEV Wash Buffer.
Using ~2 BV of ice-cold TEV Wash Buffer, load Ni-NTA resin onto an Econo-Column® Chromatography Column and let TEV Wash Buffer drain through resin until level with top of resin.
Elute protein from an Econo-Column® Chromatography Column using 5-10 BV of TEV Elution Buffer, one BV at a time.
Note: It is useful to run a Bradford assay for each column volume and continue eluting until < 0.1 mg of protein elutes per column volume.
Pool elution and spin concentrate to 10 ml by centrifuging at 3,184 x g and 4 °C in an Amicon Ultra-15 Centrifugal Filter Unit.
Transfer protein solution to a 15 ml conical tube and spin at 3,184 x g and 4 °C for 3 min to remove aggregates.
Dialysis and purification using ion-exchange chromatography
Wash dialysis tubing (SpectrumTM Labs Spectra/PorTM 6 8000 D MWCO Standard RC Pre-wetted Dialysis Kits 5.1 ml/cm vol./Length) in TEV Dialysis Buffer.
Seal one end of dialysis tubing with a dialysis tube clip (SnakeSkinTM Dialysis Clips).
Transfer protein solution into dialysis tubing and seal the other end of the tubing with another dialysis clip.
Place dialysis bag gently into 5 L of TEV Dialysis Buffer and dialyze overnight (~12 h) at 4 °C.
Transfer protein solution to a 15 ml conical tube and spin at 3,184 x g and 4 °C for 10 min to remove any protein that may have precipitated during dialysis.
Remove supernatant and filter protein through Steriflip® 0.22 µm filter prior to loading on FPLC.
Pre-equilibrate the HiTrap® SP XL Column in two column volumes (CV) of water at a flow rate of 0.5 ml/min and then two CV of TEV Low-Salt IEX Buffer at a flow rate of 0.5 ml/min.
Load filtered protein onto the HiTrap® SP XL Column at a flow rate of 0.5 ml/min.
Wash with two CV of TEV Low-Salt IEX Buffer at a flow rate of 0.5 ml/min.
Elute using a gradient from 0% TEV High-Salt IEX Buffer to 80% TEV High-Salt IEX Buffer at a flow rate of 0.5 ml/min over the course of 40 min. Collect 0.5 ml fractions in FisherbrandTM Disposable Borosilicate Glass Tubes with Plain End. Make sure the entire FPLC system is kept at 4 °C.
Using the UV chromatogram at 280 nm, asses the location of the main peak of the TEV protease.
Assess the purity of the putative TEV protease-containing fractions using Coomassie Brilliant Blue by loading 20 µl of each fraction onto an SDS-PAGE gel (Figure 1). Only pool fractions with >95% purity. See data analysis section for further details.
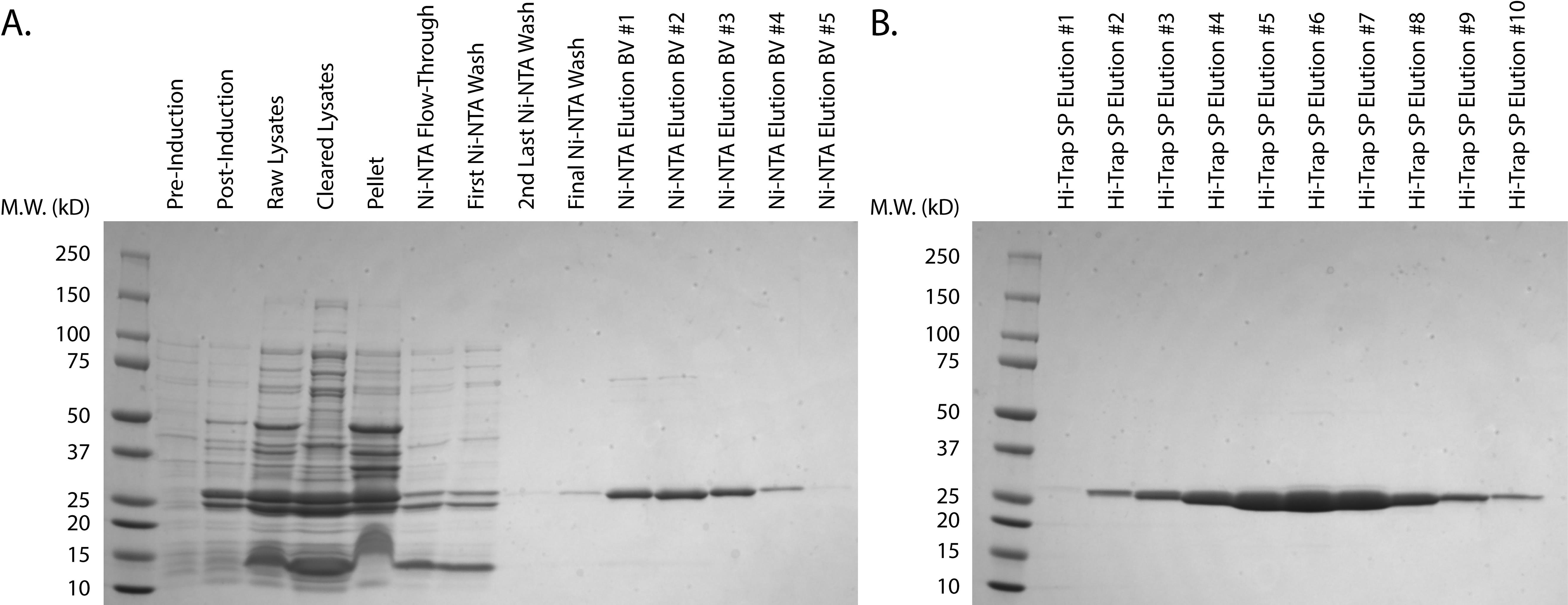
Figure 1. Purification of His-TEV protease. A. Purification of His-TEV protease from transformed, uninduced cells through elution from Ni-NTA resin. B. Elution of His-TEV protease from HiTrap® SP XL Column. Protein elution fractions were determined via chromatogram. Hi-Trap SP elution fractions #4-8 were pooled based on purity, spin concentrated, and snap-frozen.Storage
Transfer pooled elution to Amicon Ultra-15 Centrifugal Filter Unit.
Spin concentrate to ~100 µM by centrifuging at 3,184 x g and 4 °C.
Note: Protein concentration can be measured using a Bradford assay, BCA assay, or a Nanodrop.
Supplement protein to 50% (v/v) glycerol by adding filtered 100% glycerol.
Aliquot protein in protein LoBind tubes and snap freeze using liquid nitrogen. Aliquot size can vary but generally 100 µl-1,000 µl aliquots are most useful.
Store samples at -80 °C. Protein can be stored indefinitely.
Purification of Skd3
Note: The following protocol is for purifying Skd3 from a 6 L culture volume of bacteria which yields approximately 20 mg of >95% pure protein.
Bacterial cell transformation, induction, and harvest
Transform ~100 ng of pMALC2_MBP-PARLSkd3 or pMALC2_MBP-MPPSkd3 plasmid (ampicillin resistant; available from the authors) into Escherichia coli BL21 (DE3) RIL cells (chloramphenicol resistant) and plate on Agar Plates supplemented with ampicillin and chloramphenicol (Cupo and Shorter, 2020). Grow transformants at 37 °C overnight.
The next day, harvest the transformants by scraping the entire plate into a starter culture of 90 ml Growth Media supplemented with ampicillin and chloramphenicol.
Grow starter culture at 37 °C with 250 RPM shaking for 2 h.
Add 1 L Growth Media to each of 6 PYREX® 2,800 ml Fernbach-Style Culture Flask with Baffles. Add 10 ml starter culture to each 1 L baffled shaker flask.
Growth cultures at 37 °C with 250 RPM shaking until cultures reach OD600 = 0.3 to 0.6 (approximately 1-3 h).
Once cultures reach OD600 = 0.3 to 0.6, rapidly cool shakers and media to 15 °C (approximately 30-60 min).
Once temperature is equilibrated to 15 °C, supplement 1 mM IPTG and continue shaking at 15 °C and 250 RPM for 16 h.
Spin cells in 1 L centrifuge bottles at 4,658 x g for 25 min to pellet the cells. Discard the supernatant.
Bacterial cell lysis
Note: All steps should be performed on ice/at 4 °C with pre-chilled tubes unless otherwise indicated.
Resuspend the bacterial cell pellet in 72 ml ice-cold Skd3 Lysis Buffer and transfer to 50 ml conical tubes.
Add 6 ml of 20 mg/ml lysozyme to the Skd3 Lysis Buffer. Incubate cells on ice with lysozyme for 30 min.
Sonicate sample with one 30 s cycle and three 15 s cycles at power level 9. Keep conical tubes on ice while sonicating and let lysate cool on ice for 5 min between sonication cycles.
Note: It is extremely important to keep the lysate cool at this step as the sonication can heat the lysates and denature the protein.
Transfer lysate into NalgeneTM Oak Ridge High-Speed PPCO Centrifuge Tubes and centrifuge lysate at 30,597 x g and 4 °C for 20 min to clear the lysates.
Purification against N-terminal maltose binding protein (MBP) tag and ATP wash
Equilibrate 30 ml amylose resin with two BV of ice-cold Skd3 Lysis Buffer in 50 ml conical tubes. Centrifuge at 796 x g and 4 °C for 5 min. Discard supernatant. Repeat 3-4 times.
Note: Higher centrifuge speeds at this step will damage the amylose resin.
Add cleared bacterial cell lysates to equilibrated amylose resin. Nutate slowly at 4 °C for 4 h.
Note: If lysates are nutated too rapidly, bubbles will form in the buffer which can drive protein denaturation.
Centrifuge at 796 x g and 4 °C for 5 min. Discard supernatant.
Add two BV of ice-cold Skd3 Wash Buffer to the resin and gently but fully resuspend the resin in the Skd3 Wash Buffer. Centrifuge at 796 x g and 4 °C for 5 min. Discard supernatant. Repeat until the resin has been washed with ~15 BV of Skd3 Wash Buffer.
Add one BV of room temperature Skd3 ATP Wash Buffer. Centrifuge at 796 x g and 25 °C for 5 min. Discard supernatant.
Add one BV of room temperature Skd3 ATP Wash Buffer. Nutate at 25 °C for 30 min. Centrifuge at 796 x g and 25 °C for 5 min. Discard supernatant.
Add one BV of room temperature Skd3 ATP Wash Buffer. Centrifuge at 796 x g and 25 °C for 5 min. Discard supernatant and put the resin back on ice or at 4 °C.
Add two BV of ice-cold Skd3 Wash Buffer to the resin and gently but fully resuspend the resin in the Wash Buffer. Centrifuge at 796 x g and 4 °C for 5 min. Discard supernatant. Repeat until the resin has been washed with another ~15 BV of Skd3 Wash Buffer.
Elution from amylose resin and TEV cleavage
Add two BV of ice-cold Skd3 Elution Buffer to the resin and gently but fully resuspend the resin in the Skd3 Elution Buffer. Centrifuge at 796 x g and 4 °C for 5 min. Discard supernatant. Repeat four times to exchange the resin into Skd3 Elution Buffer.
Add one BV of Skd3 Elution Buffer to resin.
Add 5 mg of TEV. Nutate for 4 h at 34 °C.
Transfer resin and Skd3 Elution Buffer slurry to the chromatography columns set inside 15 ml conical tubes and centrifuge at 4,658 x g and 4 °C for 10 min.
Pool flow-through of chromatography columns and keep on ice.
Purification using size-exclusion chromatography
Pre-equilibrate the HiPrepTM 26/60 Sephacryl S-300 HR Column first in one CV of filtered, sterile water at a flow rate of 0.5 ml/min. Then equilibrate with one CV Skd3 Sizing Buffer at a flow rate of 0.5 ml/min.
Note: It is important to start equilibration into water the day before lysing the cells and start the equilibration into Sizing Buffer first thing the morning the cells are lysed as this process takes 6 h for each equilibration step.
Transfer pooled flow-through to spin concentration columns.
Spin concentrate to < 13 ml by centrifuging at 3,184 x g and 4 °C in an Amicon Ultra-15 Centrifugal Filter Unit.
Note: This is the maximum volume that can be loaded onto a HiPrepTM 26/60 Sephacryl S-300 HR Column with good column resolution. If another size-exclusion chromatography column is used, this value will differ.
Filter protein through Steriflip® 0.22 µm filter prior to loading on FLPC.
Note: This step is important to prevent the loading of aggregates or debris onto the FPLC and size-exclusion column which can be severely damaging.
Load filtered protein onto the HiPrepTM 26/60 Sephacryl S-300 HR Column at a flow rate of 0.5 ml/min.
Flow 1.5 CV of Skd3 Sizing Buffer through the HiPrepTM 26/60 Sephacryl S-300 HR Column at a flow rate of 0.5 ml/min and collect 1 ml fractions in FisherbrandTM Disposable Borosilicate Glass Tubes with Plain End. Make sure the entire FPLC system is kept at 4 °C.
Note: Once the sizing step has started, it can run overnight, and purification can resume the next morning.
The next morning, using the UV chromatogram at 280 nm, asses the location of the main peak of Skd3. Note that Skd3 generally elutes at a broad peak from 57 ml to 69 ml. Smaller peaks will also be observed for some residual cleaved MBP and TEV protease.
Assess the purity of the putative Skd3-containing fractions using Coomassie Brilliant Blue by loading 20 µl of each fraction onto an SDS-PAGE gel (Figure 2). Only pool fractions with > 95% purity. See data analysis section for further details.
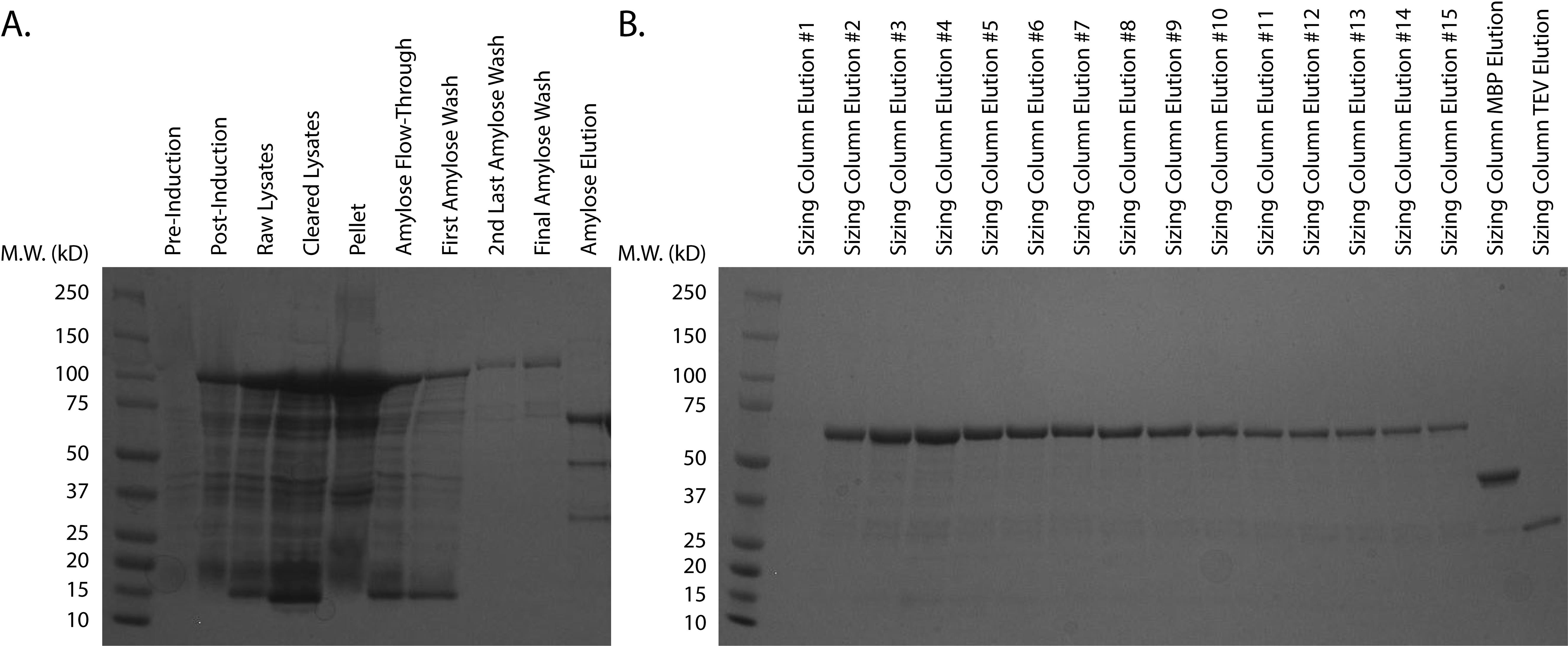
Figure 2. Purification of MPPSkd3. A. Purification of MPPSkd3 from transformed, uninduced cells through elution from amylose resin. B. Elution of MPPSkd3 from HiPrepTM 26/60 Sephacryl S-300 HR Column. Protein elution fractions were determined via chromatogram. Fractions of separated cleaved-MBP tag and TEV protease are shown for reference. HiPrepTM 26/60 Sephacryl S-300 HR Column #2-15 were pooled based on purity, spin concentrated, and snap-frozen.Storage
Transfer pooled elution to spin concentration columns.
Spin concentrate to ~100 µM by centrifuging at 3,184 x g and 4 °C in an Amicon Ultra-15 Centrifugal Filter Unit.
Note: Protein concentration can be measured using a Bradford assay, BCA assay, or a Nanodrop.
Supplement protein to 5 mM ATP using 250 mM ATP Stock (pH = 8.0).
Aliquot protein in protein LoBind tubes and snap freeze using liquid nitrogen. Aliquot size can vary but generally 10 µl-100 µl aliquots are most useful.
Store samples at -80 °C.
Data analysis
To evaluate purity of the proteins a purity gel can be run by loading 2 µg, 4 µg, 6 µg, 8 µg, and 10 µg of the final product onto an SDS-PAGE gel and stain with Coomassie Brilliant Blue. Percent purity can be assessed by measuring protein band intensity using FIJI ImageJ software.
To evaluate the identity of the purified proteins, a Western blot can be run using an anti-Skd3 antibody for Skd3 and anti-His antibody for TEV protease. An assortment of anti-Skd3 antibodies can be found in Cupo and Shorter (2020).
To evaluate the activity of Skd3, ATPase and luciferase disaggregation assays can be performed as described previously (Cupo and Shorter, 2020). To evaluate the activity of TEV protease, a substrate containing a TEV-cleavage site can be incubated with TEV protease for a period of 2 h with samples taken at 15 min, 30 min, 1 h, and 2 h time points. The samples can then be analyzed by running on an SDS-PAGE gel and staining with Coomassie Brilliant Blue. The cleavage activity of TEV protease can be compared to a commercial (or previously purified) stock to calculate relative activity.
Recipes
Note: Store all buffers at 4 °C.
Agar Plates
1 capsule LB per 1 L
20g agar per 1 L
100 µg/ml ampicillin
34 µg/ml chloramphenicol
Notes:
Autoclave and mix well.
Add the ampicillin and chloramphenicol after the agar has cooled but before it has solidified.
Growth Media
1 capsule LB per 1 L
0.2% (w/v) glucose
100 µg/ml ampicillin
34 µg/ml chloramphenicol
Notes:
Autoclave and mix well.
Add the ampicillin and chloramphenicol fresh before use.
The glucose is important to prevent degradation of the amylose resin by bacterial amylase enzymes.
Lysozyme Stock Solution
20 mg/ml Lysozyme from chicken egg white
Note: Dissolve in water.
TEV Lysis Buffer
25 mM Tris-HCl pH = 8.0
500 mM NaCl
10 mM BME
1 cOmpleteTM, Mini, EDTA-free Protease Inhibitor Cocktail tablet per 50 ml
Note: Add BME and cOmplete Protease Inhibitor fresh before use and steri-filter all buffers using Stericup Quick Release-GP Sterile Vacuum Filtration System.
TEV Wash Buffer
25 mM Tris-HCl pH = 8.0
500 mM NaCl
25 mM imidazole
10 mM BME
Note: Add BME fresh before use and steri-filter all buffers using Stericup Quick Release-GP Sterile Vacuum Filtration System.
TEV Elution Buffer
25 mM Tris-HCl pH = 8.0
500 mM NaCl
300 mM imidazole
10 mM BME
Note: Add BME fresh before use and steri-filter all buffers using Stericup Quick Release-GP Sterile Vacuum Filtration System.
TEV Dialysis Buffer
25 mM HEPES-NaOH pH = 7.0
5 mM BME
5% (v/v) glycerol
Note: Add BME fresh before use.
TEV Low-Salt IEX Buffer
25 mM HEPES-NaOH pH = 7.0
5 mM DTT
5% (v/v) glycerol
Note: Add DTT fresh before use and steri-filter all buffers using Stericup Quick Release-GP Sterile Vacuum Filtration System.
TEV High-Salt IEX Buffer
25 mM HEPES-NaOH pH = 7.0
750 mM NaCl
5 mM DTT
5% (v/v) glycerol
Note: Add DTT fresh before use and steri-filter all buffers using Stericup Quick Release-GP Sterile Vacuum Filtration System.
Skd3 Lysis Buffer
40 mM HEPES-KOH pH = 7.4
500 mM KCl
20% (w/v) glycerol
5 mM ATP
10 mM MgCl2
2 mM BME
1 cOmpleteTM, Protease Inhibitor Cocktail tablet per 50 ml
5 µM Pepstatin
Note: Add BME, cOmplete Protease Inhibitor, and Pepstatin fresh before use and steri-filter all buffers using Stericup Quick Release-GP Sterile Vacuum Filtration System.
Skd3 Wash Buffer
40 mM HEPES-KOH pH = 7.4
500 mM KCl
20% (w/v) glycerol
5 mM ATP
10 mM MgCl2
2 mM BME
1 cOmpleteTM, Protease Inhibitor Cocktail tablet per 250 ml
2.5 µM Pepstatin
Note: Add BME, cOmplete Protease Inhibitor, and Pepstatin fresh before use and steri-filter all buffers using Stericup Quick Release-GP Sterile Vacuum Filtration System.
Skd3 ATP Wash Buffer
40 mM HEPES-KOH pH = 7.4
500 mM KCl
20% (w/v) glycerol
20 mM ATP
10 mM MgCl2
2 mM BME
1 cOmpleteTM, Protease Inhibitor Cocktail tablet per 250 ml
2.5 µM Pepstatin
Note: Add BME, cOmplete Protease Inhibitor, and Pepstatin fresh before use and steri-filter all buffers using Stericup Quick Release-GP Sterile Vacuum Filtration System.
Skd3 Elution Buffer
50 mM Tris-HCl pH=8.0
300 mM KCl
10% (w/v) glycerol
5 mM ATP
10 mM MgCl2
2 mM BME
Note: Add BME fresh before use and steri-filter all buffers using Stericup Quick Release-GP Sterile Vacuum Filtration System.
Skd3 Sizing Buffer
50 mM Tris-HCl pH = 8.0
500 mM KCl
10% (w/v) glycerol
1 mM ATP
10 mM MgCl2
1 mM DTT
Note: Add DTT fresh before use and steri-filter all buffers using Stericup Quick Release-GP Sterile Vacuum Filtration System.
ATP Stock
250 mM ATP
Adjust pH to 8.0 using KOH
Acknowledgments
We thank JiaBei Lin, Bede Portz, Edward Chuang, and Zachary March for providing key feedback to develop this protein purification protocol. Our work was funded by the Blavatnik Family Foundation Fellowship (R.R.C.), The G. Harold and Leila Y. Mathers Foundation (J.S.), and NIH grants T32GM008275 (R.R.C.), F31AG060672 (R.R.C.), R01GM099836 (J.S.), and R21AG061784 (J.S.).
Competing interests
J.S. is a consultant for Dewpoint Therapeutics and Maze Therapeutics.
References
- Capo-Chichi, J. M., Boissel, S., Brustein, E., Pickles, S., Fallet-Bianco, C., Nassif, C., Patry, L., Dobrzeniecka, S., Liao, M., Labuda, D., Samuels, M. E., Hamdan, F. F., Vande Velde, C., Rouleau, G. A., Drapeau, P. and Michaud, J. L. (2015). Disruption of CLPB is associated with congenital microcephaly, severe encephalopathy and 3-methylglutaconic aciduria. J Med Genet 52(5): 303-311.
- Cupo, R. R. and Shorter, J. (2020). Skd3(human ClpB) is a potent mitochondrial protein disaggregase that is inactivated by 3-methylglutaconic aciduria-linked mutations. Elife 9: 55279.
- Kanabus, M., Shahni, R., Saldanha, J. W., Murphy, E., Plagnol, V., Hoff, W. V., Heales, S. and Rahman, S. (2015). Bi-allelic CLPB mutations cause cataract, renal cysts, nephrocalcinosis and 3-methylglutaconic aciduria, a novel disorder of mitochondrial protein disaggregation. J Inherit Metab Dis 38(2): 211-219.
- Kiykim, A., Garncarz, W., Karakoc-Aydiner, E., Ozen, A., Kiykim, E., Yesil, G., Boztug, K. and Baris, S. (2016). Novel CLPB mutation in a patient with 3-methylglutaconic aciduria causing severe neurological involvement and congenital neutropenia. Clin Immunol 165: 1-3.
- Mann, J. R., Gleixner, A. M., Mauna, J. C., Gomes, E., DeChellis-Marks, M. R., Needham, P. G., Copley, K. E., Hurtle, B., Portz, B., Pyles, N. J., Guo, L., Calder, C. B., Wills, Z. P., Pandey, U. B., Kofler, J. K., Brodsky, J. L., Thathiah, A., Shorter, J. and Donnelly, C. J. (2019). RNA Binding Antagonizes Neurotoxic Phase Transitions of TDP-43. Neuron 102(2): 321-338e328.
- Mroz, D., Wyszkowski, H., Szablewski, T., Zawieracz, K., Dutkiewicz, R., Bury, K., Wortmann, S. B., Wevers, R. A. and Zietkiewicz, S. (2020). CLPB (caseinolytic peptidase B homolog), the first mitochondrial protein refoldase associated with human disease. Biochim Biophys Acta Gen Subj 1864(4): 129512.
- Saunders, C., Smith, L., Wibrand, F., Ravn, K., Bross, P., Thiffault, I., Christensen, M., Atherton, A., Farrow, E., Miller, N., Kingsmore, S. F. and Ostergaard, E. (2015). CLPB variants associated with autosomal-recessive mitochondrial disorder with cataract, neutropenia, epilepsy, and methylglutaconic aciduria. Am J Hum Genet 96(2): 258-265.
- Wortmann, S. B., Zietkiewicz, S., Kousi, M., Szklarczyk, R., Haack, T. B., Gersting, S. W., Muntau, A. C., Rakovic, A., Renkema, G. H., Rodenburg, R. J., Strom, T. M., Meitinger, T., Rubio-Gozalbo, M. E., Chrusciel, E., Distelmaier, F., Golzio, C., Jansen, J. H., van Karnebeek, C., Lillquist, Y., Lucke, T., Ounap, K., Zordania, R., Yaplito-Lee, J., van Bokhoven, H., Spelbrink, J. N., Vaz, F. M., Pras-Raves, M., Ploski, R., Pronicka, E., Klein, C., Willemsen, M. A., de Brouwer, A. P., Prokisch, H., Katsanis, N. and Wevers, R. A. (2015). CLPB mutations cause 3-methylglutaconic aciduria, progressive brain atrophy, intellectual disability, congenital neutropenia, cataracts, movement disorder. Am J Hum Genet 96(2): 245-257.
Article Information
Copyright
Cupo and Shorter. This article is distributed under the terms of the Creative Commons Attribution License (CC BY 4.0).
How to cite
Readers should cite both the Bio-protocol article and the original research article where this protocol was used:
- Cupo, R. R. and Shorter, J. (2020). Expression and Purification of Recombinant Skd3 (Human ClpB) Protein and Tobacco Etch Virus (TEV) Protease from Escherichia coli. Bio-protocol 10(23): e3858. DOI: 10.21769/BioProtoc.3858.
- Cupo, R. R. and Shorter, J. (2020). Skd3(human ClpB) is a potent mitochondrial protein disaggregase that is inactivated by 3-methylglutaconic aciduria-linked mutations. Elife 9: e55279.
Category
Microbiology > Microbial biochemistry > Protein
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










