- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Protocol for Isolation, Stimulation and Functional Profiling of Primary and iPSC-derived Human NK Cells
Published: Vol 10, Iss 23, Dec 5, 2020 DOI: 10.21769/BioProtoc.3845 Views: 7720
Reviewed by: Giusy Tornillorozan romel attiliJia Wang

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
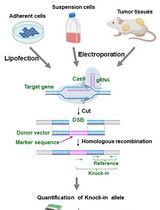
Assay for Site-Specific Homologous Recombination Activity in Adherent Cells, Suspension Cells, and Tumor Tissues
Yuki Yoshino [...] Natsuko Chiba
Apr 5, 2025 2502 Views
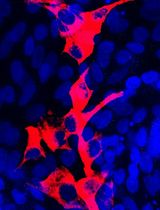
Isolation and Culture of Ferret Airway Stem Cells
Ziying Yan [...] Feng Yuan
Jul 20, 2025 2440 Views
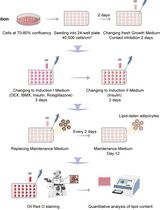
Optimization of Adipogenic Differentiation Protocol for Murine and Human Cell Culture Models
Junwan Fan [...] Wenyan He
Jan 20, 2026 292 Views
Abstract
Natural killer (NK) cells are innate immune cells, characterized by their cytotoxic capacity, and chemokine and cytokine secretion upon activation. Human NK cells are identified by CD56 expression. Circulating NK cells can be further subdivided into the CD56bright (~10%) and CD56dim NK cell subsets (~90%). NK cell-like cells can also be derived from human induced pluripotent stem cells (iPSC). To study the chemokine and cytokine secretion profile of the distinct heterogenous NK cell subsets, intracellular flow cytometry staining can be performed. However, this assay is challenging when the starting material is limited. Alternatively, NK cell subsets can be enriched, sorted, stimulated, and functionally profiled by measuring secreted effector molecules in the supernatant by Luminex. Here, we provide a rapid and straightforward protocol for the isolation and stimulation of primary NK cells or iPSC-derived NK cell-like cells, and subsequent detection of secreted cytokines and chemokines, which is also applicable for a low number of cells.
Keywords: CD56Background
Natural killer (NK) cells are part of the innate immune system and provide the first line defense against viral infections and malformations. In the human blood, two distinct NK cell populations can be identified based on CD56 and CD16 expression: CD56brightCD16+/- and CD56dimCD16+ NK cells (Melsen et al., 2016). CD56bright NK cells represent the minor subset (~10% of NK cells) and are known for their cytokine and chemokine secretion, but low cytotoxicity. In contrast, the CD56dim NK cells have high cytotoxic capacity. To test functional responses of NK cells in the absence of other cells, NK cells first need be enriched from peripheral blood mononuclear cells (PBMC) by negative enrichment. To study the production of effector molecules by the distinct NK cell populations upon stimulation, intra- and extracellular flow cytometry can be performed (Eberlein et al., 2010). However, this technique is limited to the number of effector molecule-specific antibodies available. Moreover, multiple samples are required to analyze the major effector molecules making this technique less suitable for a low number of cells. As an alternative, the distinct NK cell subsets can be sorted and stimulated. Cytokines and chemokines can be subsequently measured in the supernatant by Luminex (Lugthart et al., 2016). The advantages are: 1) distinct NK cell subsets cannot influence each other, 2) the supernatant can be harvested at multiple timepoints, which allows studying kinetics of the same cells, 3) > 25 effector molecules can be studied at once. Moreover, since no cell harvesting and fixation is required, cells could be stored or used for further experiments.
As an alternative for primary NK cells, NK cells can be derived from human iPSC. The different protocols used are based on the stepwise differentiation of human iPSC into mesoderm, hemogenic or hematopoietic progenitor cells, and subsequently into CD56+ NK cells. CD34+CD45+ hematopoietic progenitors or CD34+CD31+ hemogenic progenitors are generated using either stromal cells like OP9 or embryoid bodies in the presence of hematopoietic and vascular growth factors (Knorr et al., 2013). The CD34+ progenitors are further differentiated towards NK cells using a cytokine cocktail in either the presence or absence of OP9-DL1 stroma cells (Knorr et al., 2013; Zeng et al., 2017). Cytokine cocktails used contain SCF, FLT3L, IL-3, IL-15 and IL-7 in the absence of OP9-DL1 (Knorr et al., 2013), or SCF, FLT3L, and IL-7 when co-culturing on OP9-DL1 cells (Zeng et al., 2017). The latter conditions simultaneously generate T cells (Themeli et al., 2013 and 2020), however, substitution of IL-7 by IL-15, or the addition of IL-15 results in much purer (> 99%) CD56+ NK cell populations (Zeng et al., 2017). In our hands, CD56+ NK cells generated from iPSC best resemble primary CD56bright NK cells (Themeli et al., 2020). If the generation of T cells is hampered, by example in the case of RAG2 deficiency, previously undescribed small populations with NK cell-specific cytokine secretion profiles can be found (Themeli et al., 2020). To functionally profile rare NK cell-like populations a sensitive and easy-to-use protocol is paramount. Here we describe such a protocol that allows a rapid assessment of NK cell-specific secretion profiles using as few as 10,000 cells.
Materials and Reagents
NK cell isolation
General
Laboratory disposables:
Pipettes
15 ml tubes
50 ml tubes (Greiner, Cellstar, catalog number: 227261 )
Pasteur pipettes
Eppendorf tubes (Eppendorf, catalog number: 0030121023 )
50 ml Syringe (Becton Dickinson Medical, catalog number: BD 300865 ), 10 ml Syringe (Becton Dickinson Medical, catalog number: BD 307736 )
0.22 µm syringe filter (Whatman FP30 CA-S, catalog number: 10462300 )
Syringe needle (Becton Dickinson MICROLANCE 3, 19 G x 40 MM, catalog number: 301500 )
96-wells round bottom plate non-sterile (Corning, catalog number: 3799 )
Micronic tubes (NBS scientific, catalog number: MP32022 )
5 ml round-bottom polystyrene tubes (Corning, Falcon, catalog number: 352052 )
PBS (Fresenius Kabi, catalog number: 8717973380153 ), 4°C
Bürker-Türk counting chamber (VWR, catalog number: HECH40444702 )
Türk′s solution (Merck, Sigma-Aldrich, catalog number: 1092770100 )
Bovine serum albumin (BSA) (Merck,Sigma-Aldrich, catalog number: A9576 ), 4 °C
EDTA (Merck, Calbiochem, catalog number: 324503, Molecular weight 372.24 ), RT
Distilled water (Aqua BBraun, B Braun, catalog number: 0082479E )
NaOH (Merck, catalog number: 106498 )
Fetal Calf Serum (FCS, heat inactivated for 30 min at 56 °C to inactivate complement) (Merck, Sigma-Aldrich, catalog number: F7524 ), -20°C
EDTA solution 0.5 M (see Recipes), RT
AIMV medium (Thermo Fisher Scientific, Gibco, catalog number: 31035025 ), 4 °C
Collection medium (see Recipes), 4 °C
Antibody for cell sorting (see Recipes), 4 °C
CD56 clone N901 (ECD) (Beckman Coulter, for instance, catalog number: A82943 ), 4 °C or clone 5.1H11 (Biolegend, for instance, catalog number: 362550 ), 4°C or clone B159 (Becton Dickinson, for instance, catalog number: 560361 ), 4 °C
Only required for primary NK cell isolation
Pre-separation filters 30 µm (Miltenyi, catalog number: 130-041-407 ), RT
MACS columns MS or LS (Miltenyi, catalog number: 130-042-201 or 130-042-401 ), RT
Ficoll Paque Plus (Merck, Sigma, catalog number: GE17-1440-02 ), RT, dark
RPMI 1640 Medium (Thermo Fisher Scientific, Gibco, catalog number: 72400054 ), 4 °C
Human serum albumin 200 g/L (Sanquin, Albuman, catalog number: 8717185830897 ) 4 °C
Penicillin-Streptomycin 100x (Merck, Sigma-Aldrich, catalog number: P0781 ), -20 °C
NK cell isolation kit human (Miltenyi, catalog number: 130-092-657 ), 4 °C
CD33 clone P67.6 (PE) (Becton Dickinson, for instance, catalog number: 345799 ), 4 °C
CD14 clone M5E2 (PE-Cy7) (Becton Dickinson for instance, catalog number: 557742 ), 4 °C
CD3 clone UCHT1 (BV421) (Becton Dickinson, for instance, catalog number: 562426 ), 4 °C
CD19 clone SJ25C1 (BV510) (Becton Dickinson, for instance, catalog number: 562947 ), 4 °C
MACS buffer (see Recipes), 4 °C
Dilution medium (see Recipes), 4 °C
Wash medium (see Recipes), 4 °C
Only required for iPSC-derived NK cell isolation
30 µm CellTrics filter (Sysmex, catalog number: 04-004-2326 )
CD7 clone 124-1D1 (eBioscience, for instance, catalog number: 25-0079-41 ), 4 °C or clone 4H9 (Becton Dickinson, for instance, catalog number: 347483 ), 4 °C or clone M-T701 (Becton Dickinson, catalog number: 561934 ), 4 °C
Laboratory disposables:
Pipettes
15 ml tubes
50 ml tubes
Eppendorf tubes
96-well round bottom plate (Greiner, catalog number: 650185 )
AIMV medium (Thermo Fisher Scientific, Gibco, catalog number: 31035025 )
Fetal Calf Serum (FCS, heat inactivated for 30 min at 56 °C to inactivate complement) (Merck, Sigma-Aldrich, catalog number: F7524 ), -20 °C
Recombinant human IL-12 (Peprotech, catalog number: 200-12 ), -20 °C
Recombinant human IL-15 (Peprotech, catalog number: 200-15 ), -20 °C
Recombinant human IL-18 (MBL International, catalog number: B001-5 ), -20 °C
Interleukin mix (see Recipes)
Stimulation
Laboratory disposables:
Pipettes
15 ml polypropylene tubes
0.5 ml polypropylene tube
Reagent reservoirs
Aluminium foil
Sealing tape (for instance, Merck, Greiner, catalog number: A5596-100EA )
Ice
Paper towels
Bio-Plex Pro Human Cytokine 27-plex Immunoassay (Bio-Rad, catalog number: M50-0KCAF0Y ), 4 °C
Functional profiling
Equipment
Laminar flow cabinet for sterile work (biosafety level II) (Euroflow EF4, CleanAir by Baker)
Tube rack to hold 15, 50 ml tubes (for instance, VWR, catalog numbers: 89215-778 and 89215-778 )
Micropipettes (P10, P100, P1000) (for instance, Gilson, catalog number: F167380 )
12 channel multichannel pipette (for instance, Eppendorf, catalog number: 3125000060 )
Table top centrifuge with adapters for plates and for 15 ml and 1.5 ml tubes (Eppendorf, model: 5810 R )
CO2 Incubator (at 5% CO2 and 37 °C) (for instance, Panasonic, model: MCO 170-AIC )
Magnetic stirrer (for instance, VWR, catalog number: 89215-778 )
FACS Aria cell sorter (Becton Dickinson, Aria I, II, III) but any equivalent fluorescence-activated cell sorter should work
MiniMACS or MidiMACS separator (Miltenyi, catalog number: 130-042-102 or 130-042-302 , respectively)
MACS MultiStand (Miltenyi, catalog number: 130-042-303 , RT)
Autoclave (for instance, VWR, Ward’s, catalog number: 470230-598 )
Vortex (for instance Scientific Industries, model: Vortex Genie 2, catalog number: 200-SI-0236 )
Bio-Plex 200 system (Bio-Rad, for instance, catalog number: 171000201 )
Bio-Plex handheld magnetic washer (Bio-Rad, catalog number: 171020100 )
Plate shaker (for instance, Biosan, model: PSU-2T )
Software
Diva software (Becton Dickinson, v6.0 or later, 2007 or later)
Bio-Plex Manager software (Bio-Rad, v6.2, 2018)
Procedure
Primary NK cell isolation
Peripheral blood mononuclear cell (PBMC) isolation
Note: At room temperature (RT) unless stated otherwise.
Dilute at least 15 ml blood 2x in dilution medium and mix.
Note: From 15 ml blood typically ~1.5 x106 NK cells can be isolated. This number will yield ~7.5 x 104 CD56bright NK cells.
Add 4 ml Ficoll to a 15 ml tube.
Carefully layer the diluted blood sample (10 ml) onto the Ficoll (do not mix).
Repeat Steps 2 and 3 for the remaining diluted blood.
Centrifuge 15 min at 1,000 x g without brake.
Harvest the PBMC by carefully pipetting (using a Pasteur pipette) the white layer of cells between plasma and ficoll.
Transfer the PBMC to a new 50 ml tube (pool per sample max 25 ml).
Add wash medium up to 50 ml.
Centrifuge 10 min at 800 x g.
Remove the supernatant by aspiration and resuspend the cells.
If applicable, pool the cells in 150 ml tube.
Add wash medium up to 10 ml.
Centrifuge 10 min at 540 x g.
Remove supernatant and resuspend cells in the recommended volume of MACS buffer (~1 ml MACS buffer for every 10 ml of undiluted blood).
Count the cells with Türk′s solution (dilute 10 µl cell suspension with 90 µl Türk′s solution). Keep the remaining cells at 4 °C.
Keep 2 x 106 PBMC aside for single stain controls in sorting procedure.
NK cell isolation from PBMC using MACS
Note: Keep cells and buffers at 4 °C during the NK cell isolation. Check the manufacturer’s instructions for any updates in the protocol.
Transfer cells to 15 ml tube and add up to 14 ml MACS buffer.
Centrifuge 5 min 540 x g.
Note: Volumes for magnetic labeling are minimum volumes, when working with less than 1 x 107 cells, use this volume, when working with more than 1 x 107 cells, scale up the volumes accordingly.
Remove supernatant by aspiration. Resuspend cells in 40 µl of MACS buffer per 1 x 107 cells.
Example: For 1.5 x 107 cells, add 60 µl of MACS buffer
Add 10 µl of NK cell Biotin-Antibody Cocktail per 1 x 107 cells.
Mix well and incubate for 5 min at 4 °C.
Add 30 µl of MACS buffer per 1 x 107 cells.
Add 20 µl of NK cell MicroBead Cocktail per 1 x 107 cells.
Mix well and incubate for 10 min at 4 °C.
Place the column (MS for ≤ 1 x 107 cells, LS for > 1 x 107 PBMC) in the magnetic field of the MACS separator, which is placed on the MACS MultiStand. A maximum of 1 x 108 PBMC can be loaded per LS column.
Put the pre-separation filter on top of the column.
Position the column into a 15 ml tube.
Rinse the column by pipetting MACS buffer in the filter: 500 µl (MS column), 3 ml (LS column)
Replace the 15 ml tube filled with buffer with a new 15 ml tube (the flow-through with NK cells will be collected in this tube).
Important: Only pipet new volumes when the column reservoir is empty.
Pipette the labeled PBMC in the filter. A minimum volume of 500 µl is required for magnetic separation. If necessary, add MACS buffer to the cell suspension.
Wash column by pipetting MACS buffer in the filter: 500 µl (MS column), 3 ml (LS column)
Repeat Step 15 twice.
Centrifuge the 15 ml tube containing the unlabeled NK cells and centrifuge 5 min at 540 x g.
Resuspend in MACS buffer (~500-1,000 µl) and count.
FACS of NK cell subsets
Note: Keep cells and buffers at 4 °C.
Prepare antibody mixes (30 µl per 1 x 106 NK cells). Prevent bleaching by keeping the antibody mixes in the dark! See Recipes for example calculation. Important: use titrated antibodies.
Note: Titration is performed on 106 PBMC using the recommended dilution as the median of 5 concentrations. A 2-fold dilution factor is typically applied. Recommended dilutions should be saturating but at the same not alter the mean fluorescence intensity of the antigen negative cells.
Prepare single stain control mixes (1 antibody + MACS buffer), 25 µl per single stain. Use same dilution as used for complete antibody mix. Do not forget to include 1 unstained control.
Transfer 2 x 105 PBMC into a well of a 96 wells plate (1 well/single stain).
Pellet cells by centrifugation for 5 min, 540 x g at 4 °C (for tube), or 2 min, 540 x g at 4 °C (for plate).
For tube: remove supernatant from the 15 ml tube by aspiration and resuspend in the correct amount of antibody mix. Incubate in the dark for 30 min at 4 °C.
For plate: remove supernatant from the wells by flicking the plate once above the sink. Remove any access liquid by gently tapping the plate once onto a paper towel. Add 25 µl of single stain mix. Incubate in the dark for 30 min at 4 °C.
Wash cells by adding 10 ml (for tube) and 200 µl (for plate) MACS buffer to the cells.
Pellet cells by centrifugation as before: 5 min, 540 x g at 4 °C (for tube), 2 min, 540 x g at 4 °C (for plate).
Remove supernatant as described before (Step 5).
For tube: resuspend NK cells in MACS buffer, to a concentration of 1 x 107 NK cells per ml.
For plate: resuspend the PBMC in 70 µl MACS buffer, and collect them in micronic tubes.
Prepare round bottom polystyrene tubes with 3 ml collection medium (max. 6 x 105 cells can be collected in 1 tube).
Sort the cells at low pressure (100 µm nozzle, 20 PSI, this is important for the functioning of the NK cells after the sorting). Use the gating strategy as depicted in Figure 1. If more subsets are desired, the CD56bright can be further subdivided in a CD16- and CD16+ subset. Sorting of 1.5 x 106 NK cells will yield (when taking into account cell loss) ~7.5 x 104 CD56bright NK cells and 1 x 106 CD56dim NK cells.
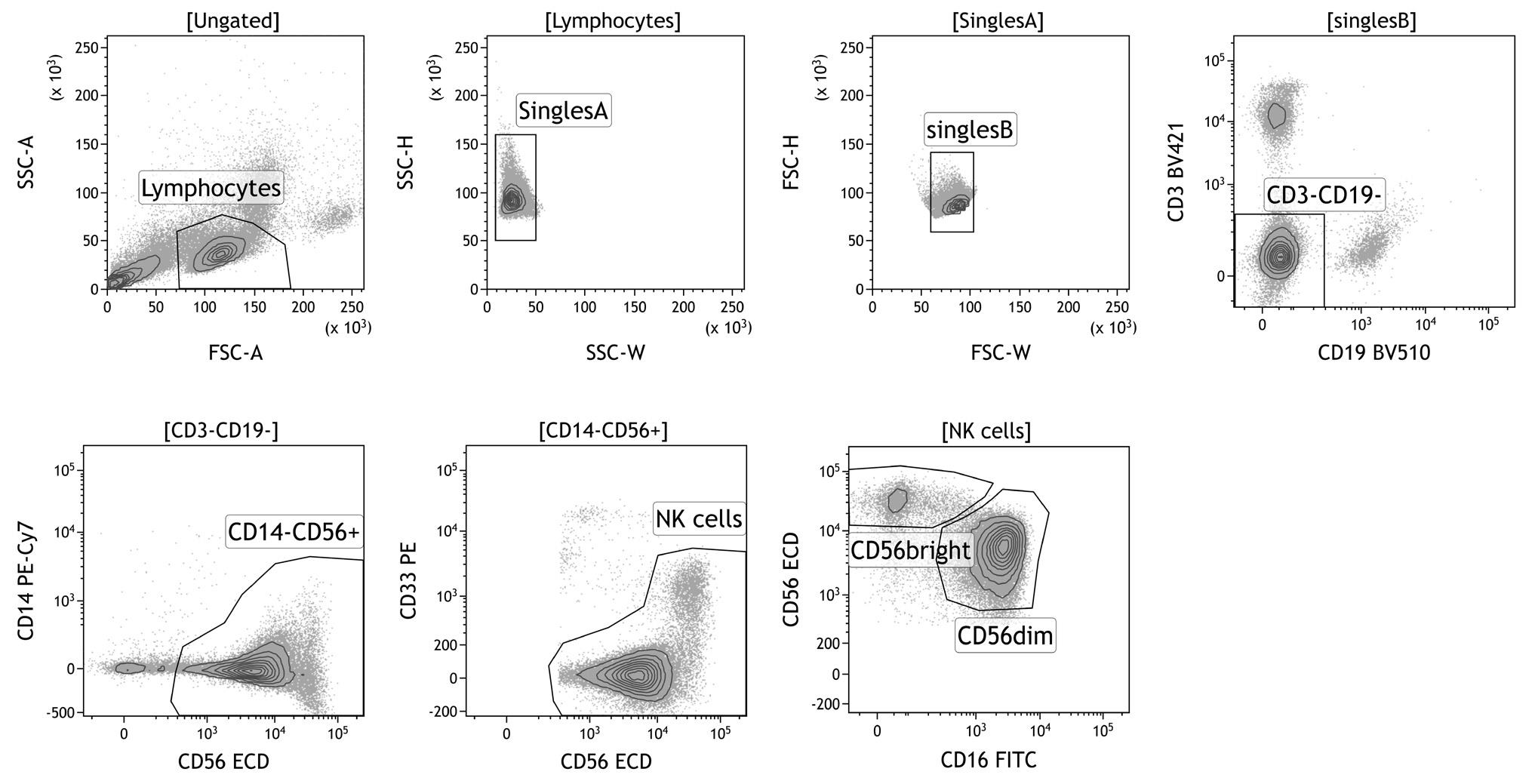
Figure 1. Gating strategy for sorting of primary NK cell subsets. First, lymphocytes are gated based on forward- and side scatter. Next, doublets, CD3+ T cells, CD19+ B cells, CD14+ monocytes, and CD33+CD56- myeloid cells are excluded. NK cells can be further subdivided based on intensity expression of CD56 and CD16: CD56brightCD16+/- and CD56dimCD16+ NK cells.
Note: Keep cells and buffers at 4 °C during the NK cell isolation.
Collect the floating cells in the iPSC differentiation culture (Figure 2A) into a 15 ml or 50 ml conically shaped tube. The tube size depends on the scale of differentiation (typical format: a few wells from a 6-well plate).
Note: ~1 x 106 floating cells are typically harvested from one 6 well plate. Depending on the success of differentiation 20-70% of the floating cells are CD7+CD56+ NK cells.
Pass the cell suspension through a 30 µm CellTrics filter (Sysmex) and collect the cells in a 15 ml or 50 ml conically shaped tube.
Centrifuge the collected cell suspension at 400 x g for 5 min at 4 °C.
Remove the supernatant by aspiration and resuspend the pelleted cells in 500 µl ice-cold PBS (wash Step 1).
Centrifuge the cells at 400 x g for 5 min at 4 °C.
Remove the supernatant by aspiration and resuspend the pelleted cells in 500 µl ice-cold MACS buffer (wash Step 2).
Centrifuge the cells at 400 x g for 5 min at 4 °C.
Remove the supernatant by aspiration and resuspend the pelleted cells in 500 µl ice-cold MACS buffer (wash Step 3).
Take a small aliquot (10 µl) and count the cells.
Transfer the cells into a 0.5 ml Eppendorf tube or 96 wells pate (round-bottom or V-shaped bottom well). Required number of tubes/wells: unstained control, single stain controls, complete stain. Use 2 x 104-5 x 104 cells for the single stain or unstained controls. The rest is used for the combi-stain of the cells that will be sorted. Never go beyond 1 x 106 cells/ 25 µl staining solution.
Centrifuge the cells at 400 x g for 5 min at 4 °C (for tube) or 2 min, 540 x g at 4 °C (for plate).
Prepare antibody mixes: single stain control mixes (1 antibody + MACS buffer), 25 µl per single stain, and a complete antibody mix containing at least anti-CD7 and anti-CD56 to identify NK cell-like cells.
Note: Use identical dilutions for the same antibodies and shield the antibodies from light, particularly, when tandem dyes are used as conjugates. Do not forget to include an unstained control and preferentially also a positive control consisting of mononuclear blood cells, by example from peripheral blood.
Remove the supernatant by aspiration ( fortube) or by flicking the plate once above the sink (for plate). Remove any access liquid by gently tapping the plate once onto a paper towel (for plate).
Resuspend the cells in the antibody mix.
Incubate in the dark for 30 min at 4 °C (typically in the fridge).
Centrifuge the cells at 400 x g for 5 min at 4 °C (for tube) or 2 min, 540 x g at 4 °C (for plate).
Remove the supernatant as described before (Step 13) and resuspend the cells in 200 µl ice-cold MACS buffer.
Centrifuge the cells at 400 x g for 5 min at 4 °C (for tube) or 2 min, 540 x g at 4 °C (for plate).
Remove the supernatant as described before (Step 13) and resuspend the single stain, unstained cells, and positive control cells in 70 µl MACS buffer and transfer them into Micronic or 5 ml round-bottom polystyrene tubes. The cells to be sorted are resuspended at a concentration of 10 x 106 cells per ml.
Prepare 5 ml round-bottom polystyrene tubes with 3 ml collection medium (max 6 x105 cells can be collected in 1 tube).
Sort the CD7+CD56+ cells at low pressure (100 µm nozzle, 20 PSI). Use the gating strategy as depicted in Figure 2B.
Note: Dependent on the question to be addressed and required purity antibodies against additional NK cell markers such as CD16 may be used.
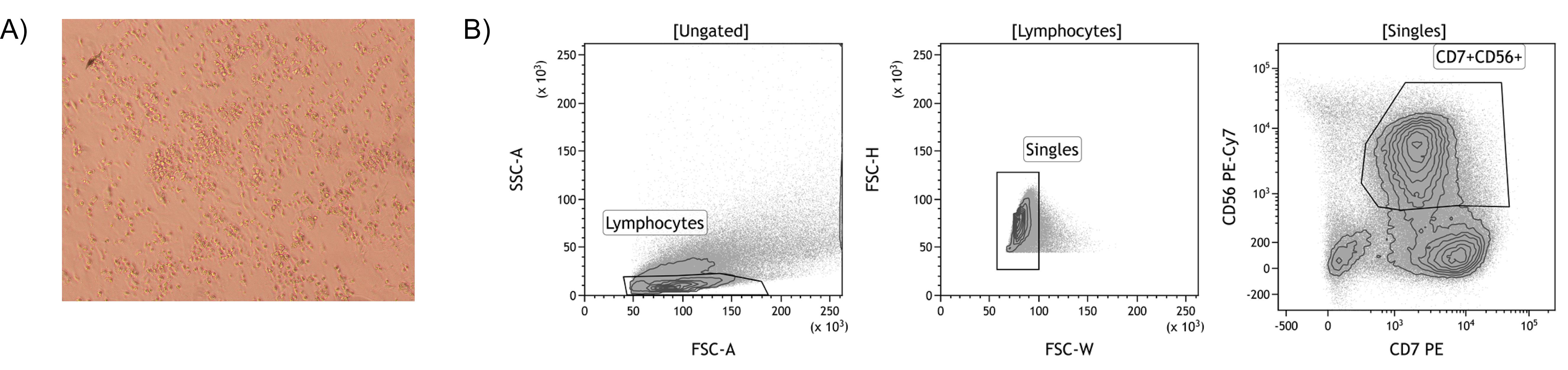
Figure 2. Isolation of iPSC-derived NK cells by FACS. A. An example of the differentiation of iPSC towards NK cells after co-culturing with OP9-DL1 cells for 3 weeks (100x magnification). Non-adherent cells are harvested and CD56+ NK cells are isolated using FACS. B. Gating strategy to isolate iPSC-derived CD7+CD56+ NK cells.NK cell stimulation
Spin down the sorted NK cell subsets and resuspend at a concentration of 5.6 x 104 cells/ml in AIMV medium with 1% Penicillin-Streptomycin (the final FCS concentration will be ~5%).
Add 180 µl /well NK cells in a sterile 96-wells round-bottom plate.
Add 20 µl of IL-12, IL-15 and IL-18 mix and medium without cytokines (AIMV + 1% Penicillin-Streptomcyin) to the different wells.
Note: Cells from healthy donors that are cultured in IL-12, IL-15 and IL-18 are the positive control for IFN-γ production. Cells cultured in medium without cytokines act as negative control. Preferentially, stimulations should be performed in triplicate. If more cells were sorted than needed stimulations with single or double cytokine mixtures should be included.
Include 1 well with 200 µl medium (AIMV + 1% Penicillin-Streptomcyin + 5% FCS) and 1 well with 200 µl medium supplemented with interleukins (as background for the Luminex).
Wrap plastic foil around plate to prevent evaporation.
Culture cells for 20 h in incubator at 37 °C, 5% CO2.
Note: When studying kinetics, take a sample of 10 µl at multiple time points.
Spin down plate, 540 x g for 2 min.
Harvest supernatant and store in sealed 96 wells plate or Micronic tubes at -20 °C.
Luminex
Note: The protocol below has been optimized and therefore differs from the manufacture’s instruction.
Spin down the standard vial and reconstitute a single vial of standard in 500 µl of diluent (= AIMV medium + 5% FCS), gently vortex for 5 s and incubate on ice for 30 min.
Label 9 0.5 ml polypropylene tubes Std1-Std8 and Blank.
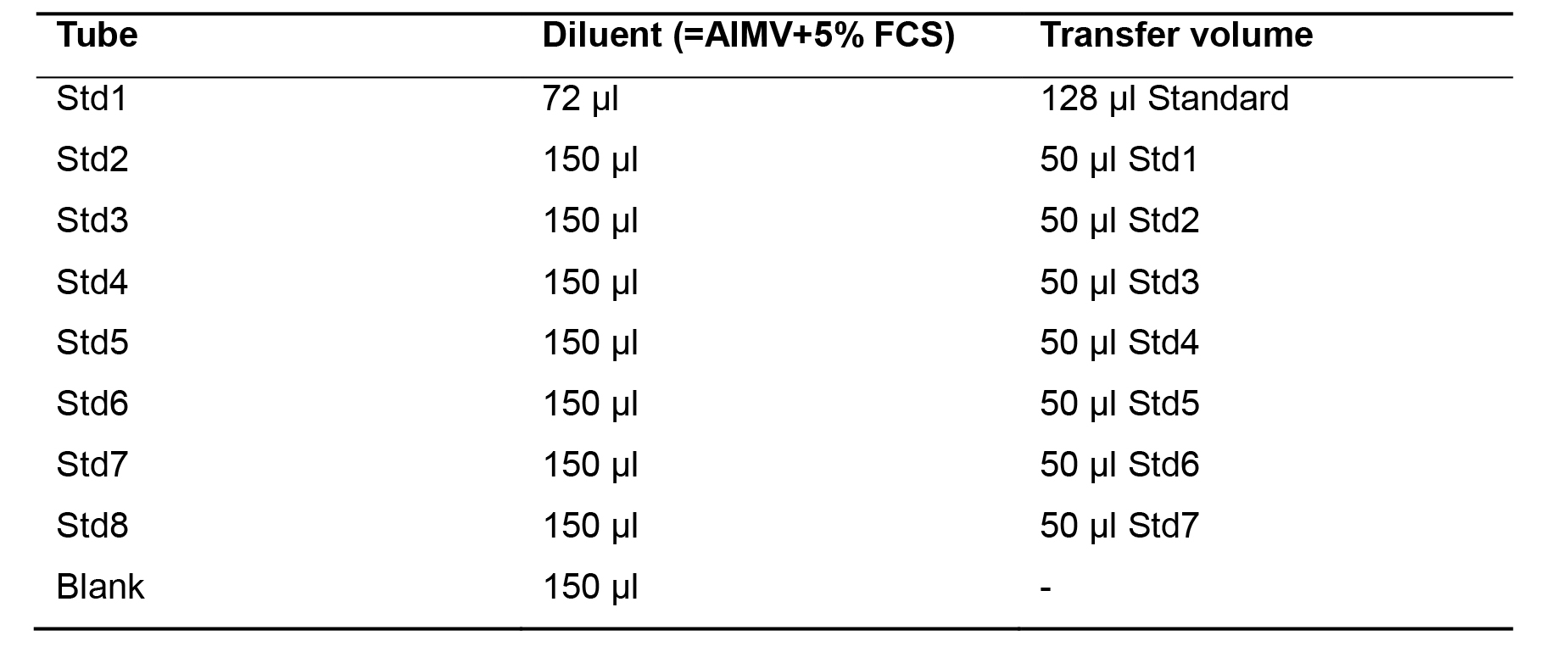
Add the specified volume of diluent to each tube (Table 1).
Vortex the reconstituted standard gently for 5 sec, take 128 µl and add to tube Std1.
Vortex Std1 and transfer 50 µl from tube Std1 to tube Std2 and vortex. Important: use new pipet tips for every volume transfer.
Continue with serial dilutions form tube S2 to S8 by transferring 50 µl each time.
Table 1. Serial dilutions to generate a standard curve
Thaw supernatant, and centrifuge at 1,000 x g for 4 min at RT.
Transfer supernatant to clean polypropylene tube.
Dilute samples 1:6 in diluent (once thawed keep samples on ice) and equilibrate to RT before use.
Make a plate lay-out (for example Figure 3) to check the number of wells required. Use duplicates for the standards and blanks. Fill the plate vertically.
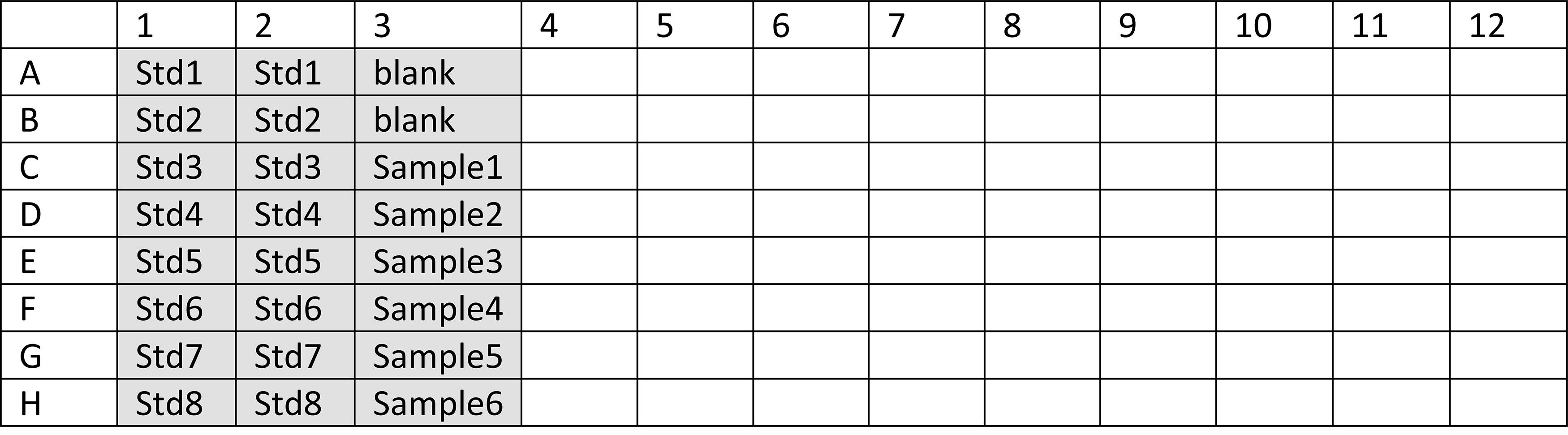
Figure 3. Plate lay-out. An example of a plate lay-out with 8 standards and 1 blank in duplicate, and 6 samples.Add the required volume of assay buffer to a 15 ml tube (Table 2).
Vortex the stock beads for 30 s at medium speed. Carefully open the cap and pipet any liquid trapped in the cap back into the vial. Important: do not centrifuge the stock beads, since the beads will be spun down.
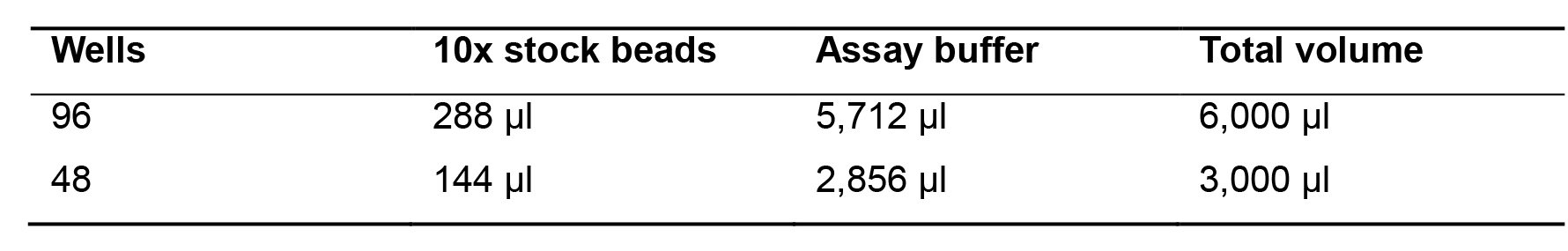
Dilute the beads by pipetting the required volume of stock beads into the 15 ml tube (Table 2). Important: protect the beads from light with aluminium foil.
Table 2. Bead dilutions
Bring standards and samples to RT before use.
Vortex the diluted beads at medium speed for 30 s and pour the beads into a reagent reservoir. Transfer 50 µg of beads to each well of the flat bottom plate.
Add 100 µl wash buffer per well. Position the plate for at least 60 s on the handheld magnet and quickly decant the waste solution. Remove any access liquid by tapping the plate onto a paper towel. Remove plate from the magnet, carefully resuspend the beads by tapping the plate, and repeat this washing step.
Gently vortex the diluted standards, blanks and samples for 5 s. Transfer 50 µl to each well. Important: use a new pipette tip for each volume transfer.
Cover the plate with a new sheet of sealing tape and protect from light with aluminum foil. Incubate on shaker at 450 rpm for 45 min at RT.
Prepare the dilution of detection antibodies 10 min before use. Add the required volume of detection antibody diluent to a new 15 ml tube (Table 3).
Vortex the stock detection antibodies at medium speed for 15-20 s. Spin down the stock for 30 s.
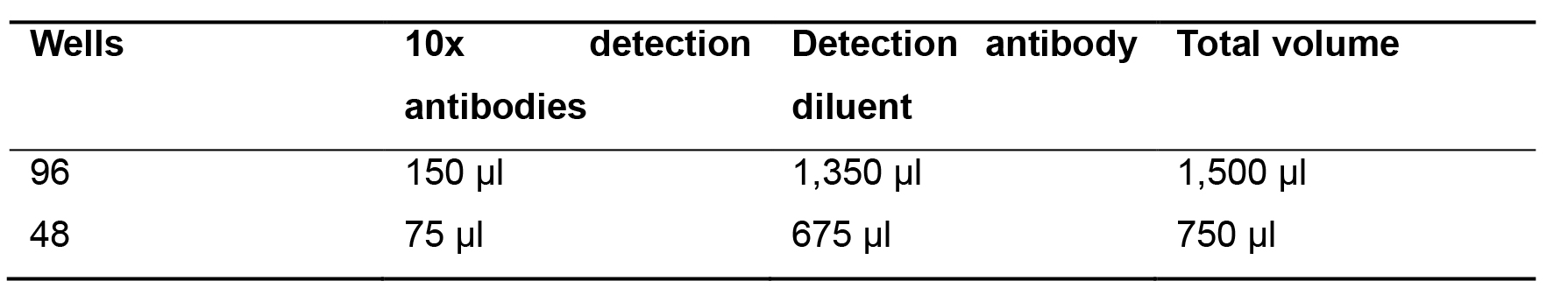
Dilute the 10x detection antibodies by pipetting the required volume into the 15 ml tube (Table 3).
After the 45 min incubation, remove the sealing tape and wash 3x with 100 µl washing buffer.
Vortex the diluted detection antibodies for 5 s, pour into a reagent reservoir and transfer 12.5 µl to each well.
Cover plate with a new sealing tape and protect from light with aluminum foil. Incubate on shaker at 450 rpm for 30 min at RT.
Table 3. Dilution of detection antibodies
Prepare the streptavidin-PE dilution 10 min before use. Add the required volume of assay buffer to a new 15 ml tube (Table 4).
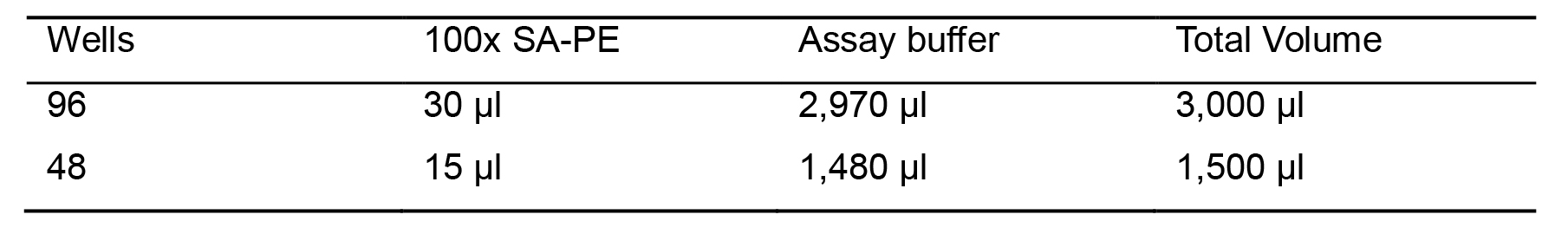
Table 4. Dilution streptavidin-PE
Vortex the 100x SA-PE for 5 s at medium speed and spin down for 30 s.
Pipette the required volume of SA-PE to the 15 ml tube (Table 4). Vortex and protect from light.
Power-up the Bio-plex system (30 min in advance).
After the 30 min incubation of the detection antibody, remove the sealing tape and wash the plate 3x with 100 µl washing buffer.
Vortex the diluted SA-PE for 5 s, pour into a reagent reservoir, and transfer 25 µl to each well.
Cover the plate with sealing tape and aluminum foil. Incubate on shaker at 450 rpm for 10 min at RT.
Wash the plate 3x with 100 µl wash buffer.
To resuspend beads for plate reading, add 80 µl of assay buffer to each well and cover the plate with sealing tape. Incubate on shaker at 850 ± 50 rpm for 30 s at RT.
Remove the tape and read the plate (using low PMT settings). In case you expect low concentrations, use high PMT settings.
Data analysis: For a detailed handbook we refer to online tutorials by Bio-Rad (Reference 1 for instance).
Note: It is important to subtract the background values (the wells with only medium or medium supplemented with interleukins) from the other values, which are automatically generated by the software based on the standard curve. In addition, check whether the experiment was successful by verifying that the negative control (cells without stimulation) is truly negative, and that the positive control (cells stimulated with IL-12, IL-15 and IL-18) shows IFN-γ production. An example of cytokine production by a variety of NK cells is shown in Figure 4.
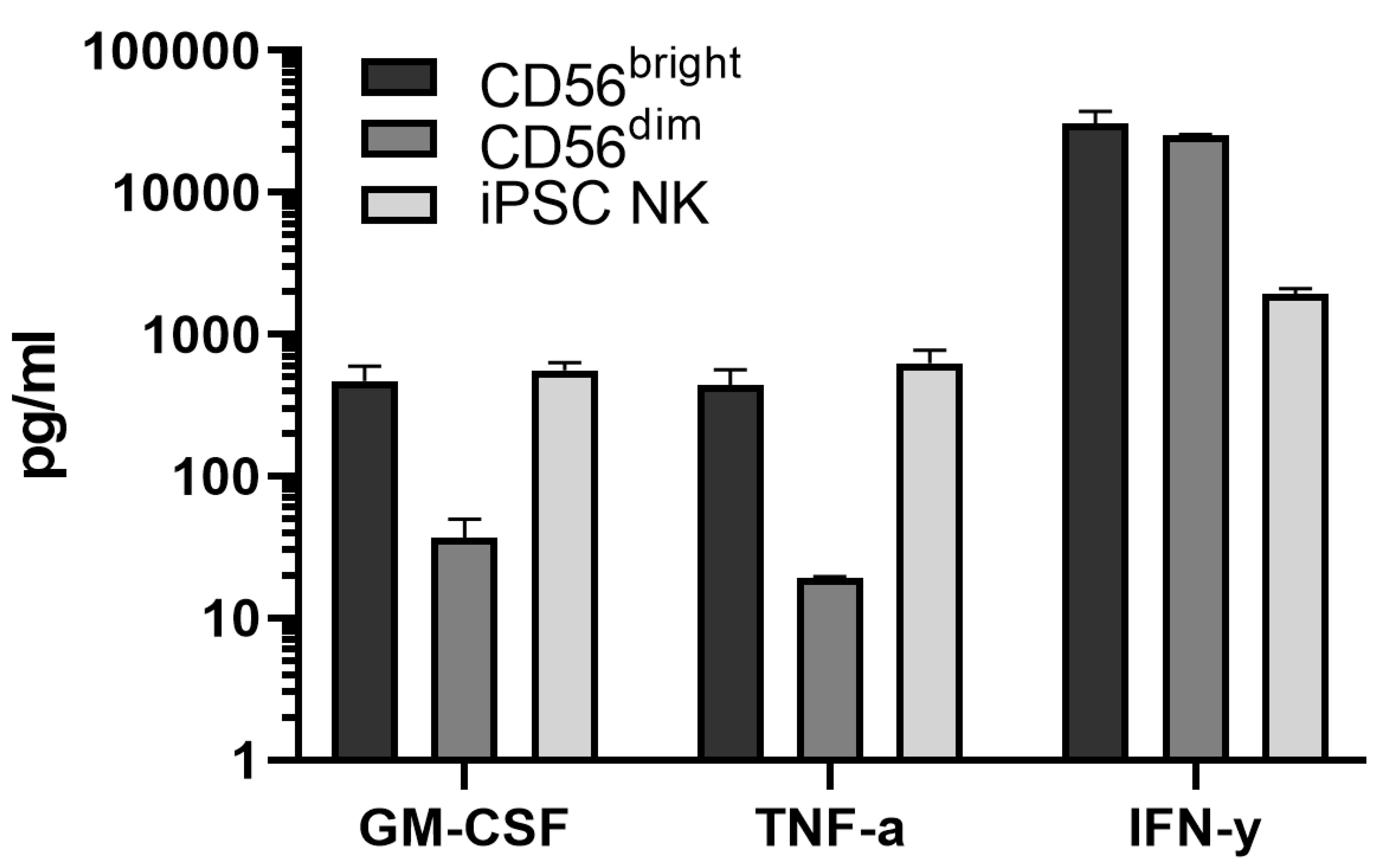
Figure 4. Example of Luminex measurements of stimulated primary and iPSC-derived NK cells. The Bio-Plex Pro Human Cytokine 27-plex Immunoassay allows detection of 27 cytokines/chemokines. NK cells are producers of predominantly GM-CSF, TNF-α and IFN-γ and various chemokines. GM-CSF, TNF-α and IFN-γ production is shown upon the culture of primary NK cell subsets (CD56bright and CD56dim) and iPSC-derived NK cells (CD7+CD56+) in the presence of IL-12, IL-15 and IL-18 for 20 h. The data is a representative of at least 2 independent experiments. The means and standard deviations are shown.
Recipes
Dilution medium
500 ml RPMI 1640 medium
5 ml penicillin-streptomycin 100x
Add Penicillin-Streptomycin to RPMI 1640 medium and shake
Wash medium
500 ml RPMI 1640 medium
5 ml Penicillin-Streptomycin 100x
2 ml Human serum albumin 200 g/L (final = 0.8 g/L)
Add Penicillin-Streptomycin and human serum albumin to RPMI 1640 medium and shake
EDTA solution 0.5 M
1,000 ml distilled water
186.12 g EDTA
Add EDTA to a bottle with distilled water, add magnetic stirring bar, and put the bottle on a magnetic stirrer. Add pellets NaOH until EDTA dissolves (around pH 7.5-8). Autoclave and store de bottle at RT. Use a syringe filter 0.22 µm to filter the solution before use
MACS buffer
500 ml PBS
8.3 ml 30% BSA (final = 0.5%)
2 ml 0.5 M EDTA solution (final = 2 mM)
Add BSA and filtered EDTA solution to the PBS and mix
Antibody mix for primary NK cells
MACS buffer
1:20 CD33 PE
1:200 CD16 FITC
1:100 CD56 ECD
1:100 CD14 PE-Cy7
1:200 CD3 BV421
1:100 CD19 BV510
Add the antibodies to the MACS buffer (in an Eppendorf tube) and vortex. Keep the mix in the dark at 4 °C
Collection medium
AIMV medium
30% FCS
1% penicillin-streptomycin
Add the FCS and Penicillin-Streptomycin to the AIMV medium and mix
Interleukin mix
AIMV medium
1% Penicillin-Streptomycin
10 ng/ml IL-12
10 ng/ml IL-15
20 ng/ml IL-18
Add the Penicillin-Streptomycin and interleukins to the AIMV medium and vortex
Acknowledgments
This study was supported by the graduate program of NWO (J.M.), Leiden University Medical Center fellowship (J.M.), LSBR foundation (H.M.) (LSBR09-11), the European Commission (Marie Curie Individual Fellowship to M.T.), the Dutch Cancer Society (KWF) (M.T.), and Stichting VUmc CCA (M.T.).
Competing interests
All authors have declared to have no competing interests.
Ethics
Primary NK cells were isolated from buffy coats of healthy adult donors (Sanquin Blood bank, Region Southwest, Rotterdam, The Netherlands). Human materials for iPSC were collected according to the approval by the “Medical Ethics Committees” of the Erasmus MC (MEC-2016-606) or the LUMC (P13-080). The experiments involving human materials were done in accordance with the principles outlined in the “Declaration of Helsinki”.
References
- Bio-Plex ManagerTM and Data ProTM Software: Streamline Multiplex Data Analysis (2016). YouTube, 1 Feb.https://www.youtube.com/watch?v=v2i3Vp6MwyE.
- Eberlein, J., Nguyen, T. T., Victorino, F., Golden-Mason, L., Rosen, H. R. and Homann, D. (2010). Comprehensive assessment of chemokine expression profiles by flow cytometry. J Clin Invest 120(3): 907-923.
- Knorr, D. A., Ni, Z., Hermanson, D., Hexum, M. K., Bendzick, L., Cooper, L. J., Lee, D. A. and Kaufman, D. S. (2013). Clinical-scale derivation of natural killer cells from human pluripotent stem cells for cancer therapy. Stem Cells Transl Med 2(4): 274-283.
- Lugthart, G., Melsen, J. E., Vervat, C., van Ostaijen-Ten Dam, M. M., Corver, W. E., Roelen, D. L., van Bergen, J., van Tol, M. J., Lankester, A. C. and Schilham, M. W. (2016). Human Lymphoid Tissues Harbor a Distinct CD69+CXCR6+ NK Cell Population. J Immunol 197(1): 78-84.
- Melsen, J. E., Lugthart, G., Lankester, A. C. and Schilham, M. W. (2016). Human Circulating and Tissue-Resident CD56bright Natural Killer Cell Populations. Front Immunol 7: 262.
- Themeli, M., Chhatta, A., Boersma, H., Prins, H. J., Cordes, M., de Wilt, E., Farahani, A. S., Vandekerckhove, B., van der Burg, M., Hoeben, R. C., Staal, F. J. T. and Mikkers, H. M. M. (2020). iPSC-Based Modeling of RAG2 Severe Combined Immunodeficiency Reveals Multiple T Cell Developmental Arrests. Stem Cell Reports 14(2): 300-311.
- Themeli, M., Kloss, C. C., Ciriello, G., Fedorov, V. D., Perna, F., Gonen, M. and Sadelain, M. (2013). Generation of tumor-targeted human T lymphocytes from induced pluripotent stem cells for cancer therapy. Nat Biotechnol 31(10): 928-933.
- Zeng, J., Tang, S. Y., Toh, L. L. and Wang, S. (2017). Generation of“Off-the-Shelf” Natural Killer Cells from Peripheral Blood Cell-Derived Induced Pluripotent Stem Cells. Stem Cell Reports 9: 1796-1812.
Article Information
Copyright
© 2020 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Melsen, J., Themeli, M., van Ostaijen-ten Dam, M., van Beelen, E., Lugthart, G., Hoeben, R., Schilham, M. and Mikkers, H. (2020). Protocol for Isolation, Stimulation and Functional Profiling of Primary and iPSC-derived Human NK Cells. Bio-protocol 10(23): e3845. DOI: 10.21769/BioProtoc.3845.
Category
Immunology > Immune cell function
Stem Cell > Pluripotent stem cell > Cell induction
Cell Biology > Cell isolation and culture > Cell differentiation
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link