- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Identification of Buffer Conditions for Optimal Thermostability and Solubility of Herpesviral Protein UL37 Using the Thermofluor Assay
Published: Vol 10, Iss 12, Jun 20, 2020 DOI: 10.21769/BioProtoc.3662 Views: 3899
Reviewed by: Anca Flavia SavulescuCristina SuárezVikas Duhan

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
Production and Purification of Cell Culture–generated Hepatitis B Virus by Transient Transfection and Density Gradient
Asako Murayama [...] Takanobu Kato
Jul 20, 2023 1940 Views
Flotation Assay With Fluorescence Readout to Study Membrane Association of the Enteroviral Peripheral Membrane Protein 2C
Kasturika Shankar [...] Lars-Anders Carlson
Apr 5, 2025 1659 Views

Detection and Analysis of S-Acylated Proteins via Acyl Resin–Assisted Capture (Acyl-RAC)
Dina A. Abdulrahman and Michael Veit
Apr 5, 2025 1936 Views
Abstract
Structural and biochemical studies of proteins require high amounts of stable, purified proteins. Protein stability often depends on the buffer composition, which includes pH and concentration of salts or other solutes such as glycerol, hence an efficient method for identifying optimal buffer conditions for stability would minimize time and resources used for protein purification and further studies. This protocol describes the use of the Thermofluor assay, in combination with a custom 24-condition screen, to identify buffer conditions that increase protein thermostability, using the conserved herpesviral protein UL37 as an example. Detailed instructions on screen conditions, running the Thermofluor MATLAB script, and analyzing the data are provided. In comparison to circular dichroism (CD), the buffer screen in combination with Thermofluor assay provides a faster and more informative method to analyze protein thermostability.
Keywords: ThermofluorBackground
Conserved across herpesviruses, the protein UL37 has multiple functions in the viral lifecycle and is required for efficient viral replication. As a component of viral tegument–a layer sandwiched between the genome-containing capsid and the lipid envelope–UL37 is not only necessary for viral assembly (Desai et al., 2001; Jambunathan et al., 2014), but is also required for efficient capsid trafficking (Leege et al., 2009), neuroinvasion (Richards et al., 2017), and counteraction of the host innate immune response (Liu et al., 2008; Zhao et al., 2016). A better understanding of the nature of its multifunctionality requires detailed knowledge of its structure and biochemical properties, which, in turn requires well-behaved purified protein.
Initial preparations of different UL37 constructs varied in solubility, monodispersity, and yield of purified protein, making protein preparations inconsistent and further characterizations irreproducible. Since buffer optimization is a straightforward approach to addressing these concerns, we developed a 24-condition screen that varies buffering agents, pH, and concentrations of NaCl and glycerol and used it in combination with a Thermofluor assay (Phillips et al., 2011) to identify buffer conditions that maximize thermostability. The N-terminal and C-terminal halves of UL37 had variable average thermal stabilities and optimal buffer conditions (Figure 1, Table 1). Yet in all cases, conditions that increased protein stability also improved protein solubility, ultimately increasing the yields of purified protein, which facilitated downstream biochemical characterization.
The Thermofluor assay has many advantages over circular dichroism (CD), which is also used to monitor protein stability (Greenfield, 2006). First, the CD can only test one condition at a time whereas the Thermofluor assay is high throughout, screening up to 384 conditions simultaneously. Second, CD only estimates the secondary structure, whereas the Thermofluor assay provides an assessment of the tertiary structure. Third, CD signal is sensitive to changes in buffer components whereas essentially any buffer can be used in the Thermofluor assay. The buffer screen described here is a good starting point for characterizing a protein of unknown biochemical properties (stability, solubility) but can be customized or expanded for individual proteins based on available biochemical information. Finally, the Thermofluor assay can also be used to assess the domain organization of larger proteins by determining whether multiple domains of a large protein represent independently folded units, thereby increasing understanding of protein structure and function.
Materials and Reagents
- 1.7 ml microcentrifuge tubes (Thomas Scientific, catalog number: 1159M35 )
- 96-well PCR microplate (Thermo Scientific, catalog number: AB0800 )
- TBS (Tris-buffered saline) (150 mM NaCl, 20 mM Tris pH 7.5)
- SYPRO Orange Protein Gel Stain (Thermo Fisher Scientific, catalog number: S6650 )
- Transparent microplate sealing film (Hampton Research, ClearSeal, catalog number: HR4-521 )
- Sodium Acetate (Fisher Scientific, catalog number: BP334 )
- Sodium Citrate (Fisher Scientific, catalog number: S279 )
- MES (Sigma-Aldrich, catalog number: M2933 )
- Sodium Cacodylate (Sigma-Aldrich, catalog number: C4945 )
- Bis-Tris (Fisher Scientific, catalog number: BP301 )
- Imidazole (Fisher Scientific, catalog number: AC301872500 )
- MOPS (American Bioanalytical, catalog number: AB01270 )
- HEPES (Fisher Scientific, catalog number: BP310500 )
- Tris (Fisher Scientific, catalog number: BP152 )
- Bicine (Sigma-Aldrich, catalog number: B3876 )
- Glycine (Fisher Scientific, catalog number: BP381 )
- CHES (Sigma-Aldrich, catalog number: C2885 )
- Ethanolamine (ACROS organics, catalog number: AC149582500 )
- CAPS (Sigma-Aldrich, catalog number: C2632 )
- Glycerol (Chem-Impex, catalog number: 30144 )
- Sodium Chloride (Fisher Scientific, catalog number: S271 )
- 45 µg protein of interest (for 24 conditions).
Notes:- Protein should be homogenous (single band on an SDS-PAGE gel) and monodisperse (a single peak on size-exclusion chromatography).
- Proteins used for this protocol included the N and C terminal domains of UL37 homolog from pseudorabies virus (PRV UL37N, PRV UL37C) and the full-length protein (PRV UL37FL). All proteins used here were purified using two rounds of affinity chromatography and one round of size-exclusion chromatography. Detailed instructions on purification of PRV UL37 can be found in Pitts et al. (2014) and Koenigsberg et al. (2018). Multiple protein batches resulted in comparable thermostability profiles for each protein.
Equipment
- Benchtop centrifuge (Beckman Coulter, model: Allegra X-15R )
- qPCR machine (Roche, LightCycler 480, catalog number: 0 5015278001 )
Software
- ThermoQ (MATLAB, MathWorks, http://www.hotproteins.com)
Procedure
- Buffer and Protein Preparation
- Make 1 ml of the following buffers:
100 mM Sodium Acetate pH 5.0
100 mM Sodium Citrate pH 5.5
100 mM MES pH 6.0
100 mM Sodium Cacodylate pH 6.5
100 mM Bis-Tris pH 6.5
100 mM Imidazole pH 7.0
100 mM PIPES pH 7.0
100 mM Sodium Malonate pH 7.0
100 mM MOPS pH 7.5
100 mM HEPES pH 7.5
100 mM Tris pH 8.0
100 mM Bicine pH 9.0
100 mM Glycine pH 9.0
100 mM CHES pH 9.5
100 mM CAPS pH 9.5
200 mM HEPES pH 7.5
200 mM HEPES pH 7.5, 50 mM NaCl
200 mM HEPES pH 7.5, 150 mM NaCl
200 mM HEPES pH 7.5, 300 mM NaCl
200 mM HEPES pH 7.5, 500 mM NaCl
200 mM HEPES pH 7.5, 150 mM NaCl, 5% glycerol
200 mM HEPES pH 7.5, 150 mM NaCl, 10% glycerol
200 mM HEPES pH 7.5, 500 mM NaCl, 5% glycerol
Note: Buffers should be made 2x the concentration intended to be screened since addition of the protein solution in a 1:1 ratio will dilute the buffer by half. Any buffers (i.e., pH, salt, detergents, etc.) can be used. Here a screen of a pH range from 5.0-9.5 is used as well as targeted optimization of the buffer used for protein purification. Optimization includes increasing salt and glycerol concentrations, both of which are known to improve protein stability (Yamasaki et al., 1990 and Vagenende et al., 2009). - Make a 300 µl solution of protein to a final concentration of 0.15 mg/ml in TBS.
- Add SYPRO orange dye to protein solution at a 1:1,000 dilution.
- Pipet 10 µl of screen buffers (from Step A1) per well of 96-well microplate.
- Add 10 µl protein-dye solution to each well containing buffer (1:1).
Note: Final protein concentration is 0.075 mg/ml and final buffer concentrations are 0.5x what is listed in Step A1. - Set-up control well with buffer and dye (no protein).
- Seal plate with transparent film.
- Spin 96-well microplate at 500 x g for 1 min at room temperature.
- Make 1 ml of the following buffers:
- Create Thermofluor method on Roche LightCycler 480
- Set filters to 465 nm for excitation and 610 nm for emission
- Click the New Method icon, select Melting Curves as the Analysis Mode in the Programs window
- Create a two-step method:
- Step 1: Set a target of 25 °C held for 10 s (without data acquisition)
- Step 2: Set a target of 95 °C with continuous acquisition at a rate of 3 fluorescent measurements per °C
- Save method as a template
- Insert 96-well microplate containing protein-dye-buffer solution
- New Method Template > Thermofluor > Apply Template
- Click Start Run, enter name for data file
- When run is finished, export data as a .txt file, save to drive
Data analysis
- Import .txt file onto computer.
- Open the MATLAB ThermoQ.m script (downloaded from www.hotproteins.com).
- Click Run > Roche > 96 wells > Select File > choose .txt file > Next > Analyze.
- Script will generate four figures:
- ThermoQ figure 1: raw fluorescence data from all wells.
- ThermoQ figure 2: smoothed, polynomial fit of the curves in ThermoQ figure 1.
- ThermoQ figure 3: first derivative of melting curves in ThermoQ figure 1, where peak corresponds to the unfolding transition.
- ThermoQ figure 4: overview of melting temperatures for all wells.
- To get exact melting temperature from ThermoQ figure 4 select Data Cursor from top menu, then click the lower left corner of a square, the Z coordinate represents the melting temperature (Figure 2).
- ThermoQ figure 3 can be used to determine presence of multiple melting events (i.e., multiple peaks).
Thermofluor was performed with UL37N, two versions of UL37C, and the full-length UL37 (Figure 1). The UL37N protein had the highest thermal stability in PIPES (pH 7.0) or Na-Malonate (pH 7.0) buffers, very low concentrations of NaCl, and glycerol (Figure 1C, Table 1). Thus, all subsequent purification batches and biochemical and structural assays, including crystallization, for UL37N protein was conducted in its optimal buffer of 50 mM PIPES pH 7.0, 50 mM NaCl, and 0.5 mM TCEP. Glycerol was added to cell lysis buffer but was left out during the rest of purification and storage because it can hinder crystallization.
The longer UL37C construct was most stable in HEPES buffer (pH 7.5), with the highest melting temperature (TM) in HEPES pH 7.5, 150 mM NaCl and 10% glycerol (B11) (Figure 1A, Table 1), which was used in subsequent purification, characterization and storage of all UL37C constructs. The melting curve of the full-length UL37 resulted in two distinct melting events, referred to as TM low and TM high, which are nearly identical to the respective TM of UL37C and UL37N (Figure 1D, Table 1). This suggests that UL37N and UL37C are two separately folded domains of UL37. In the case of UL37N and UL37C, increased thermostability correlated with increased solubility. No specific assay was used to determine protein solubility. A protein construct was considered to have an increased solubility if it could be concentrated to a higher concentration and remained in solution without precipitating at that concentration.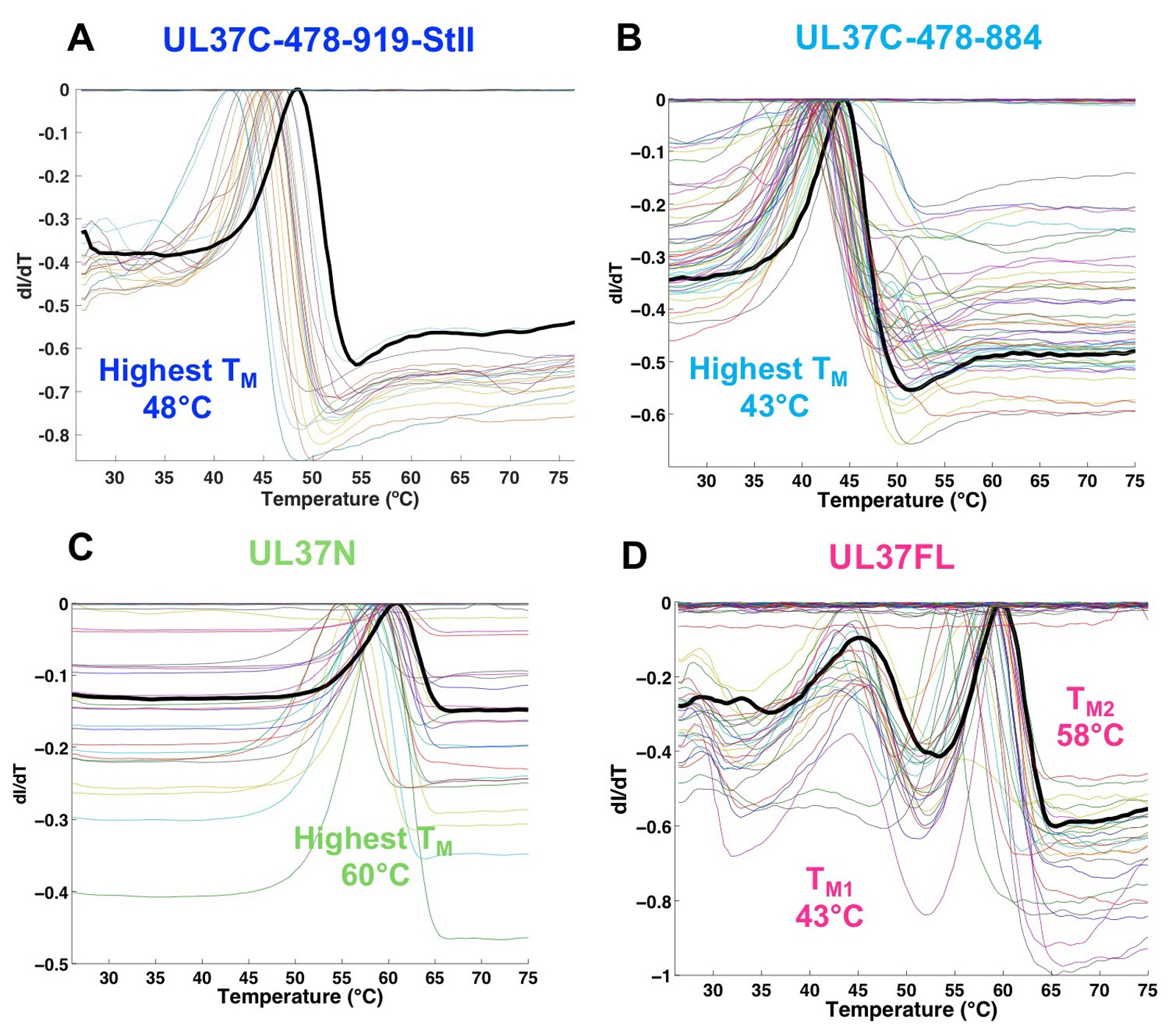
Figure 1. Results of Thermofluor assay of PRV UL37. ThermoQ script generated images (ThermoQ figure 3) that represent the first derivative of the melting curves (ThermoQ figure 1). Maxiumum values represent the unfolding transition or melting temperature (TM). Each colored line represents a different buffer condition from the screen. Black lines represent the buffer condition resulting in the highest TM. Results are shown for PRV UL37 constructs containing (A) the C terminal half (UL37C), residues 478-919; (B) UL37C, residues 478-884; (C) the N-terminal half (UL37N), residues 1-496; and (D) the full-length UL37, residues 1-919. This research was originally published in the Journal of Biological Chemistry (Koenigsberg and Heldwein, 2018).
Table 1. UL37 melting temperatures determined by Thermofluor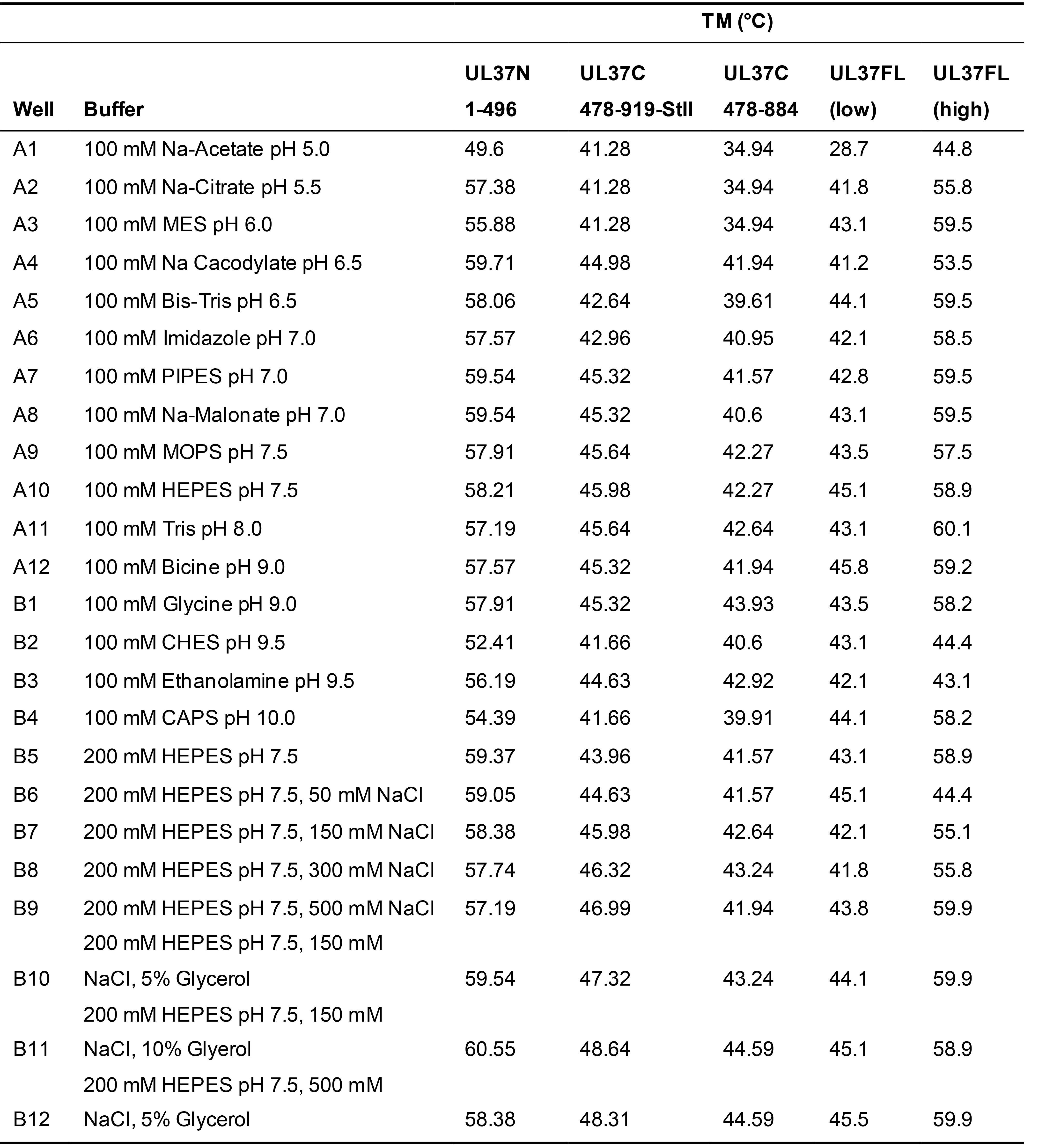
Exact buffer conditions screened for all UL37 constructs with the corresponding melting temperature, as determined by using the method described in Data analysis step 5.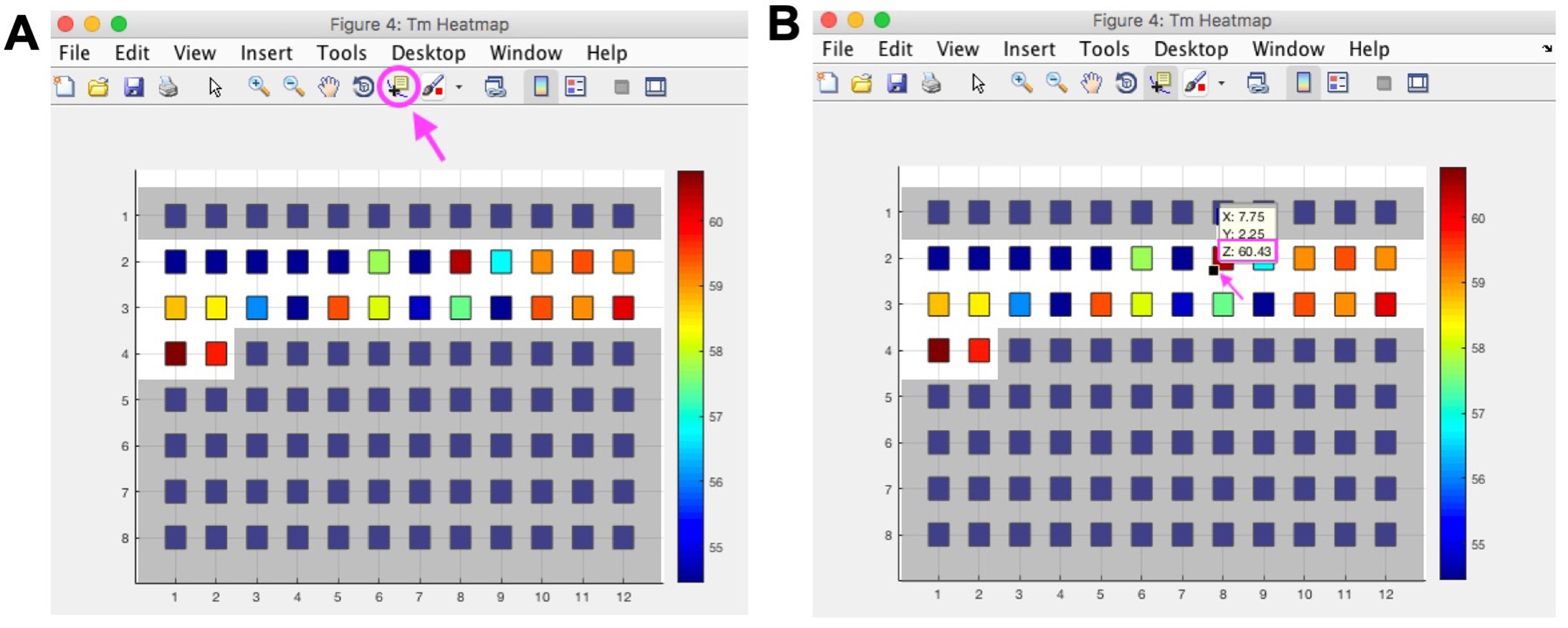
Figure 2. Determine exact melting temperature from ThermoQ figure 4. The ThermoQ script displays melting temperatures as a heatmap (ThermoQ figure 4) to visualize relative melting temperatures between different buffer conditions. To determine the exact melting temperature, click the “Data Cursor” option circled in pink in Figure 2A. Once “Data Cursor” is selected the mouse is displayed as a crossbar. Identify the square on the heatmap that corresponds to the well of interest on the plate and click the bottom left corner of the square indicated by the pink arrow in Figure 2B. A small yellow box with XYZ coordinates will appear. The Z coordinate represents the melting temperature for that square in °C. Blue squares on gray background did not contain buffer and were not part of the screen. Note: this is example data that does not correspond to data shown in Figure 1 or Table 1.
Acknowledgments
This work was supported by National Institutes of Health Grant R01 AI056346 (to E. E. H.), the Investigators in the Pathogenesis of Infectious Disease Award from the Burroughs Wellcome Fund (to E. E. H.), and a Faculty Scholar grant from the Howard Hughes Medical Institute (to E. E. H.). A.L.K. was supported by NIH training grant T32 AI007422. This protocol was alluded to in the original research papers which also contain some of the data reported here (Pitts et al., 2014 and Koenigsberg et al., 2018).
Competing interests
E.E.H. is the scientific co-founder of Thyreos, LLC. J.D.P. is currently an employee of Gilead Sciences. The authors declare that they have no conflicts of interest with the contents of this article.
References
- Desai, P., Sexton, G. L., McCaffery, J. M. and Person, S. (2001). A null mutation in the gene encoding the herpes simplex virus type 1 UL37 polypeptide abrogates virus maturation. J Virol 75(21): 10259-10271.
- Greenfield, N. (2006). Using circular dichroism as a function of temperature to determine the thermodynamics of protein unfolding and binding interactions. Nat Protoc 1(6): 2527-2535.
- Jambunathan, N., Chouljenko, D., Desai, P., Charles, A. S., Subramanian, R., Chouljenko, V. N. and Kousoulas, K. G. (2014). Herpes simplex virus 1 protein UL37 interacts with viral glycoprotein gK and membrane protein UL20 and functions in cytoplasmic virion envelopment. J Virol 88(11): 5927-5935.
- Koenigsberg, A. L. and Heldwein, E. E. (2018). The dynamic nature of the conserved tegument protein UL37 of herpesviruses. J Biol Chem 293(41): 15827-15839.
- Leege, T., Granzow, H., Fuchs, W., Klupp, B. G. and Mettenleiter, T. C. (2009). Phenotypic similarities and differences between UL37-deleted pseudorabies virus and herpes simplex virus type 1. J Gen Virol 90(Pt 7): 1560-1568.
- Liu, X., Fitzgerald, K., Kurt-Jones, E., Finberg, R. and Knipe, D. M. (2008). Herpesvirus tegument protein activates NF-kappaB signaling through the TRAF6 adaptor protein. Proc Natl Acad Sci U S A 105(32): 11335-11339.
- Pitts, J. D., Klabis, J., Richards, A. L., Smith, G. A. and Heldwein, E. E. (2014). Crystal structure of the herpesvirus inner tegument protein UL37 supports its essential role in control of viral trafficking. J Virol 88(10): 5462-5473.
- Phillips, K. and de la Pena, A. H. (2011). The combined use of the Thermofluor assay and ThermoQ analytical software for the determination of protein stability and buffer optimization as an aid in protein crystallization. Curr Protoc Mol Biol Chapter 10: Unit10 28.
- Richards, A. L., Sollars, P. J., Pitts, J. D., Stults, A. M., Heldwein, E. E., Pickard, G. E. and Smith, G. A. (2017). The pUL37 tegument protein guides alpha-herpesvirus retrograde axonal transport to promote neuroinvasion. PLoS Pathog 13(12): e1006741.
- Vagenende, V., Yap, M. G. and Trout, B. L. (2009). Mechanisms of protein stabilization and prevention of protein aggregation by glycerol. Biochemistry 48(46): 11084-11096.
- Yamasaki, M., Yano, H. and Aoki, K. (1990). Differential scanning calorimetric studies on bovine serum albumin: I. Effects of pH and ionic strength. Int J Biol Macromol 12(4): 263-268.
- Zhao, J., Zeng, Y., Xu, S., Chen, J., Shen, G., Yu, C., Knipe, D., Yuan, W., Peng, J., Xu, W., Zhang, C., Xia, Z. and Feng, P. (2016). A viral deamidase targets the helicase domain of RIG-I to block RNA-induced activation. Cell Host Microbe 20(6): 770-784.
Article Information
Copyright
© 2020 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Readers should cite both the Bio-protocol article and the original research article where this protocol was used:
- Koenigsberg, A. L., Pitts, J. D. and Heldwein, E. E. (2020). Identification of Buffer Conditions for Optimal Thermostability and Solubility of Herpesviral Protein UL37 Using the Thermofluor Assay. Bio-protocol 10(12): e3662. DOI: 10.21769/BioProtoc.3662.
- Koenigsberg, A. L. and Heldwein, E. E. (2018). The dynamic nature of the conserved tegument protein UL37 of herpesviruses. J Biol Chem 293(41): 15827-15839.
Category
Microbiology > Microbial biochemistry > Protein
Biochemistry > Protein > Fluorescence
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link











