- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
An Optimized Method to Isolate Human Fibroblasts from Tissue for Ex Vivo Analysis
(*contributed equally to this work) Published: Vol 9, Iss 23, Dec 5, 2019 DOI: 10.21769/BioProtoc.3440 Views: 6337
Reviewed by: Gal HaimovichMarzia Di DonatoAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
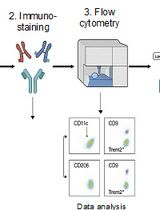
Identification and Sorting of Adipose Inflammatory and Metabolically Activated Macrophages in Diet-Induced Obesity
Dan Wu [...] Weidong Wang
Oct 20, 2025 2233 Views
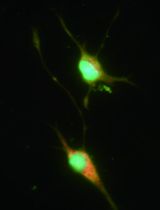
Selective Enrichment and Identification of Cerebrospinal Fluid-Contacting Neurons In Vitro via PKD2L1 Promoter-Driven Lentiviral System
Wei Tan [...] Qing Li
Nov 20, 2025 1335 Views
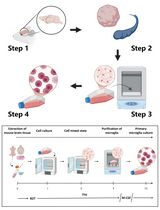
Revisiting Primary Microglia Isolation Protocol: An Improved Method for Microglia Extraction
Jianwei Li [...] Guohui Lu
Dec 5, 2025 1460 Views
Abstract
Despite their involvement in many physiological and pathological processes, fibroblasts remain a poorly-characterized cell type. Analysis of primary fibroblasts while maintaining their in vivo phenotype is challenging: standard methods for fibroblast isolation require cell culture in vitro, which is known to alter phenotypes. Previously-described protocols for the dissociation of primary tissues fail to extract sufficient numbers of fibroblasts, instead largely yielding immune cells. Here, we describe an optimized method for generating a fibroblast-enriched single-cell suspension from human tissues using combined mechanical and enzymatic dissociation. This allows analysis of ex vivo fibroblasts without the need for culture in vitro.
Keywords: Tissue disaggregationBackground
Fibroblasts are almost ubiquitous in human tissues, and the most common cell type in the stroma of a number of solid tumours, where they are referred to as cancer-associated fibroblasts (CAFs) (Ishii et al., 2016; Kalluri and Zeisberg, 2006; Rupp et al., 2014; Servais and Erez, 2013). CAFs are associated with multiple hallmarks of malignancy (Ishii et al., 2016; Tao et al., 2017) and correlate with poor prognosis in multiple solid tumours (Hanley et al., 2018).
Given these tumour-promoting effects, and their genetic stability relative to cancer cells (Ishii et al., 2016), it is unsurprising that fibroblasts are an attractive therapeutic target. However, clinical trials targeting CAFs have so far yielded disappointing results (Hofheinz et al., 2003; Narra et al., 2007). This may, in part, be due to variation within the fibroblast population: these cells are known to be heterogeneous in both normal and disease states (Desmoulière et al., 2004; Sugimoto et al., 2006; Anderberg and Pietras, 2009; Servais and Erez, 2013; Witowski et al., 2015; Kalluri, 2016; Mellone et al., 2017). However, this heterogeneity remains poorly-characterised and it is not yet clear how many subtypes are present within a given tissue or tumour type, or the nature of functional differences between groups (Herrera et al., 2013; Servais and Erez, 2013; Ishii et al., 2016).
Characterising heterogeneity of cell populations within human tissues often requires analysis at a single-cell level. Single-cell RNA sequencing is a valuable platform for characterising multicellular ecosystems. However, fibroblasts are embedded within extracellular matrix and are particularly difficult to isolate: this has led to under-representation in, for example, single-cell RNA sequencing datasets (Lambrechts et al., 2018). Unlike many murine models, there is no standardized disaggregation protocol for human solid tissues. A number of different immune cell populations have been successfully isolated and analyzed directly from tissues (Holt et al., 1986; Perrot et al., 2007; Grange et al., 2011; Quatromoni et al., 2015; Ganesan et al., 2017). However, epithelial and non-immune stromal cells are usually cultured in vitro prior to analysis (Lurton et al., 1999; Koumas et al., 2003; Comhair et al., 2012; Barkauskas et al., 2013; Mackay et al., 2013). Culture in vitro has been shown to change fibroblast phenotypes (Öhlund et al., 2017; Waise et al., 2019); thus, how and whether the functional differences described in vitro are maintained in vivo is not yet known (Lurton et al., 1999).
Here, we describe an optimized protocol for the isolation of fibroblasts from primary human tissues, allowing immediate analysis without the need for culture in vitro. In brief, primary samples undergo mechanical and enzymatic dissociation, followed by incubation with TrypLE and red cell lysis buffer (to disrupt intercellular adhesions and remove red blood cells, respectively). Use of this approach yields a single-cell suspension consisting of multiple cell types, with approximately a 4-fold greater proportion of fibroblasts compared to other disaggregation strategies (Waise et al., 2019). We describe use of this protocol for the ex vivo analysis of fibroblasts in both normal and disease states using single-cell RNA sequencing, and highlight alternative downstream applications. In addition, this approach may have applications in the analysis of other cell types (e.g., epithelial cells).
Materials and Reagents
- 5 ml syringes (BD Plastipak, catalog number: 307731)
- 10 ml syringes (BD Plastipak, catalog number: 307736)
- Pasteur pipette (Scientific Laboratory Supplies, catalog number: PIP4210)
- 50 ml Falcon tube (Sarstedt, Brand, model/catalog number: 114 x 28 mm, 62.547.004)
- 15 ml Falcon tube (Sarstedt, Brand, model/catalog number: 120 x 17 mm, 62.554.002)
- Sterile scalpel #21 blade (Swann-Morton, catalog number: 0507)
- Polystyrene cell culture dish (Sarstedt, catalog number: 83.3902)
- Syringe filtration unit Filtropur S 0.2 (Sarstedt, catalog number: 83.1826.001)
- Scissors
- Collagenase P from Clostridium histolyticum (Merck, Roche, catalog number: 11213857001). Reconstitute in PBS to 150 U/ml, store 100 μl aliquots at -20 °C
- TrypLE Express Enzyme (no phenol red; Thermo Fisher, catalog number: 12604013). Store at room temperature protected from light for up to 2 years
- Fetal bovine serum (Biosera, catalog number: FB-1001/500). Store at -20 °C for up to 60 months
- Deoxyribonuclease I from bovine pancreas (Merck, Sigma-Aldrich, catalog number: D4263). Reconstitute in 1 ml PBS (2000 U/ml), store 40 μl aliquots at -20 °C
- Dulbecco’s Modified Eagle Medium (Merck, Sigma-Aldrich, catalog number: D5671-500ML). Store at 4 °C
- L-glutamine (Merck, Sigma-Aldrich, catalog number: G7513-100ML). Store at -20 °C for up to 2 years
- Penicillin-streptomycin (Merck, Sigma-Aldrich, catalog number: P4333-100ML). Store at -20 °C for up to 2 years
- Phosphate-buffered saline (PBS)
- Amphotericin B (250 μg/ml; Gibco, catalog number: 15290-018). Store at -20 °C for 1 year
- Sterile double-distilled H2O
- Red blood cell lysis buffer (10x; BioLegend, catalog number: 420301). Store at 4 °C
- DNase stock solutions
- “Complete” DMEM (see Recipes)
- “Empty” DMEM (see Recipes)
Equipment
- Orbital shaker-incubator (e.g., Grant-bio Orbital Shaker-Incubator ES-20)
- EASYStrainer 40 μm (Grener Bio-One, catalog number: 542040)
- Centrifuge
Procedure
- Tissue dissociation
- Sample collection
Transport the tissue sample in “empty” DMEM (Recipe 2) on ice - Prepare working solutions
- In a 50 ml Falcon tube, add 100 μl Collagenase P and 40 μl DNase stock solutions to 5 ml “complete” DMEM (Recipe 1)
- Make PBS-A: add 1 μl Amphotericin to 10 ml PBS in a 15 ml Falcon tube
- Mechanical dissociation
In a polystyrene Petri dish, incise the tissue 10-12 times to relax the tissue - Wash sample in PBS-A
- Add 5 ml PBS-A to incised tissue, leave at room temperature for 5 min
- This process may be repeated if the sample is particularly congested
- Enzymatic dissociation
- Using scissors, cut the Pasteur pipette bulb at a 45-degree angle to create a scoop. Use this to remove the tissue from the PBS-A
- Transfer to the 50 ml Falcon tube containing the Collagenase P/DNase solution
- Transfer the 50 ml Falcon tube to the orbital shaker
- Incubate at 37 °C with agitation (200 rpm) for 15 min
- After 15 min, remove from the orbital shaker and sequentially pipette with 50 ml, 10 ml and 5 ml pipette tips to promote dissociation
- Return to the orbital shaker for a further 15 min, then repeat sequential pipetting
- Return to the orbital shaker for a further 30 min, then repeat sequential pipetting
- TrypLE treatment
- Centrifuge the sample at 450 x g for 5 min
- Remove the supernatant and re-suspend the resulting pellet in 1 ml of undiluted TrypLE
- Incubate at 37 °C for 10 min
- Removing non-digested tissue fragments
- Add tissue strainer to a new 50 ml Falcon tube on ice
- Strain tissue/enzyme suspension, pressing through with plunger of 5 ml syringe, simultaneously washing with “empty” DMEM
- Keep the sample on ice from this point
- Red blood cell lysis
- Centrifuge the sample at 450 x g for 5 min at 4 °C
- Aspirate the medium
- Re-suspend the pellet in 1 ml red blood cell lysis solution
- Incubate at 4 °C for 10 min
- Centrifuge at 450 x g for 5 min at 4 °C
- Sample collection
- Aspirate the red cell lysis buffer
- Re-suspend the pellet in 1 ml of “complete” DMEM
- The cells are now ready for quantification (if necessary) and use
- Sample collection
Notes
- The single-cell suspension may be used for a number of downstream applications, including single-cell RNA sequencing, establishing primary fibroblast cultures, and isolating fibroblast subpopulations. For single-cell RNA sequencing with the Drop-seq platform, re-suspend the cells in double-distilled H2O supplemented with 9% Optiprep (Sigma-Aldrich), 1% PBS and 0.1% BSA, and perform as per Macosko et al. (2015) with the following modifications: 500 ng cDNA for PCR, 15 PCR cycles. The number of cells it is feasible to use will depend on the yield from the microfluidic step.
- To establish primary cell cultures, plate 1 x 105 cells/cm2 to tissue culture plates in ‘Complete’ DMEM. Incubate in a humidified incubator at 37 °C and 5% CO2 for 2 h to allow cells to adhere, before washing three times with PBS to remove non-adherent cells. This typically yields a 99.1% pure fibroblast (CD45-EpCAM-CD31-CD90+) culture, as determined by flow cytometry analysis. However, users should note that these culture conditions will not maintain in vivo fibroblast phenotypes (Waise et al., 2019).
- Antibodies directed to specific fibroblast surface markers (e.g., PDGFR-α [Erez et al., 2010]), in combination with fluorescence-activated or magnetic cell sorting, can be used for isolation of fibroblast subpopulations. It is of note when employing these methods that enzymatic disaggregation can alter surface marker expression (Gray et al., 2002; Grange et al., 2011; Quatromoni et al., 2015), and that no single surface marker will reliably identify or differentiate all fibroblast populations (Sugimoto et al., 2006; Lambrechts et al., 2018).
Recipes
- “Complete” DMEM
Dulbecco’s Modified Eagle Medium (Sigma-Aldrich)
10% (v/v) fetal calf serum (Biosera)
1% (v/v) L-glutamine (Sigma-Aldrich)
1% (v/v) penicillin-streptomycin (Sigma-Aldrich) - “Empty” DMEM
Dulbecco’s Modified Eagle Medium (Sigma-Aldrich) only
Acknowledgments
This protocol was derived from previously-published data 28. This work was supported by Cancer Research UK and Medical Research Council Clinical Research Training Fellowships and a Pathological Society Trainee’s Small Grant to SW. Implementation of Drop-seq was supported by a Medical Research Council Discovery award (MC_PC_15078) and a Southampton Cancer Research UK Centre Development Fund Award to MJJRZ, CHO, JW, CJH & GJT. RP was supported by a John Goldman Fellowship for Future Science (2016/JGF/0003; Leuka Charity) awarded to MJJRZ. The authors thank Evan Macosko, Melissa Goldman and Steve McCarroll for their helpful advice, Dr. Serena Chee (University Hospital Southampton), Benjamin Johnson, Carine Fixmer and Maria Lane (TargetLung Clinical Trials Associates) for enabling access to clinical samples, and the patients involved in this study.
Competing interests
The authors declare no competing interests.
Ethics
Lung samples were received fresh from patients undergoing surgery at Southampton General Hospital (TargetLung study; approved by NRES Committee South Central: Hampshire A, REC number 14/SC/0186). All research was performed in accordance with the appropriate regulations. Informed consent was obtained from patients or their legal guardians.
References
- Anderberg, C. and Pietras, K. (2009). On the origin of cancer-associated fibroblasts. Cell Cycle 8(10): 1461-1462.
- Barkauskas, C. E., Cronce, M. J., Rackley, C. R., Bowie, E. J., Keene, D. R., Stripp, B. R., Randell, S. H., Noble, P. W. and Hogan, B. L. (2013). Type 2 alveolar cells are stem cells in adult lung. J Clin Invest 123(7): 3025-3036.
- Comhair, S. A., Xu, W., Mavrakis, L., Aldred, M. A., Asosingh, K. and Erzurum, S. C. (2012). Human primary lung endothelial cells in culture. Am J Respir Cell Mol Biol 46(6): 723-730.
- Desmouliere, A., Guyot, C. and Gabbiani, G. (2004). The stroma reaction myofibroblast: a key player in the control of tumor cell behavior. Int J Dev Biol 48(5-6): 509-517.
- Erez, N., Truitt, M., Olson, P., Arron, S. T. and Hanahan, D. (2010). Cancer-associated fibroblasts are activated in incipient neoplasia to orchestrate tumor-promoting inflammation in an NF-kappaB-Dependent manner. Cancer Cell 17(2): 135-147.
- Ganesan, A. P., Clarke, J., Wood, O., Garrido-Martin, E. M., Chee, S. J., Mellows, T., Samaniego-Castruita, D., Singh, D., Seumois, G., Alzetani, A., Woo, E., Friedmann, P. S., King, E. V., Thomas, G. J., Sanchez-Elsner, T., Vijayanand, P. and Ottensmeier, C. H. (2017). Tissue-resident memory features are linked to the magnitude of cytotoxic T cell responses in human lung cancer. Nat Immunol 18(8): 940-950.
- Gray, D. H., Chidgey, A. P. and Boyd, R. L. (2002). Analysis of thymic stromal cell populations using flow cytometry. J Immunol Methods 260(1-2): 15-28.
- Grange, C., Letourneau, J., Forget, M. A., Godin-Ethier, J., Martin, J., Liberman, M., Latour, M., Widmer, H., Lattouf, J. B., Piccirillo, C. A., Cailhier, J. F. and Lapointe, R. (2011). Phenotypic characterization and functional analysis of human tumor immune infiltration after mechanical and enzymatic disaggregation. J Immunol Methods 372(1-2): 119-126.
- Hanley, C. J., Mellone, M., Ford, K., Thirdborough, S. M., Mellows, T., Frampton, S. J., Smith, D. M., Harden, E., Szyndralewiez, C., Bullock, M., Noble, F., Moutasim, K. A., King, E. V., Vijayanand, P., Mirnezami, A. H., Underwood, T. J., Ottensmeier, C. H. and Thomas, G. J. (2018). Targeting the myofibroblastic cancer-associated fibroblast phenotype through inhibition of NOX4. J Natl Cancer Inst 110(1).
- Herrera, M., Islam, A. B., Herrera, A., Martin, P., Garcia, V., Silva, J., Garcia, J. M., Salas, C., Casal, I., de Herreros, A. G., Bonilla, F. and Pena, C. (2013). Functional heterogeneity of cancer-associated fibroblasts from human colon tumors shows specific prognostic gene expression signature. Clin Cancer Res 19: 5914-5926.
- Hofheinz, R. D., al-Batran, S. E., Hartmann, F., Hartung, G., Jager, D., Renner, C., Tanswell, P., Kunz, U., Amelsberg, A., Kuthan, H. and Stehle, G. (2003). Stromal antigen targeting by a humanised monoclonal antibody: an early phase II trial of sibrotuzumab in patients with metastatic colorectal cancer. Onkologie 26(1): 44-48.
- Holt, P. G., Robinson, B. W., Reid, M., Kees, U. R., Warton, A., Dawson, V. H., Rose, A., Schon-Hegrad, M. and Papadimitriou, J. M. (1986). Extraction of immune and inflammatory cells from human lung parenchyma: evaluation of an enzymatic digestion procedure. Clin Exp Immunol 66(1): 188-200.
- Ishii, G., Ochiai, A. and Neri, S. (2016). Phenotypic and functional heterogeneity of cancer-associated fibroblast within the tumor microenvironment. Adv Drug DelivRev 99(Pt B): 186-196.
- Kalluri, R. (2016). The biology and function of fibroblasts in cancer. Nat Rev Cancer 16(9): 582-598.
- Kalluri, R. and Zeisberg, M. (2006). Fibroblasts in cancer. Nat Rev Cancer 6(5): 392-401.
- Koumas, L., Smith, T. J., Feldon, S., Blumberg, N. and Phipps, R. P. (2003). Thy-1 expression in human fibroblast subsets defines myofibroblastic or lipofibroblastic phenotypes. Am J Pathol 163(4): 1291-1300.
- Lambrechts, D., Wauters, E., Boeckx, B., Aibar, S., Nittner, D., Burton, O., Bassez, A., Decaluwe, H., Pircher, A., Van den Eynde, K., Weynand, B., Verbeken, E., De Leyn, P., Liston, A., Vansteenkiste, J., Carmeliet, P., Aerts, S. and Thienpont, B. (2018). Phenotype molding of stromal cells in the lung tumor microenvironment. Nat Med 24(8): 1277-1289.
- Lurton, J., Rose, T. M., Raghu, G. and Narayanan, A. S. (1999). Isolation of a gene product expressed by a subpopulation of human lung fibroblasts by differential display. Am J Respir Cell Mol Biol 20(2): 327-331.
- Mackay, L. S., Dodd, S., Dougall, I. G., Tomlinson, W., Lordan, J., Fisher, A. J. and Corris, P. A. (2013). Isolation and characterisation of human pulmonary microvascular endothelial cells from patients with severe emphysema. Respir Res 14: 23.
- Macosko, E. Z., Basu, A., Satija, R., Nemesh, J., Shekhar, K., Goldman, M., Tirosh, I., Bialas, A. R., Kamitaki, N., Martersteck, E. M., Trombetta, J. J., Weitz, D. A., Sanes, J. R., Shalek, A. K., Regev, A. and McCarroll, S. A. (2015). Highly parallel genome-wide expression profiling of individual cells using nanoliter droplets. Cell 161(5): 1202-1214.
- Mellone, M., Hanley, C. J., Thirdborough, S., Mellows, T., Garcia, E., Woo, J., Tod, J., Frampton, S., Jenei, V., Moutasim, K. A., Kabir, T. D., Brennan, P. A., Venturi, G., Ford, K., Herranz, N., Lim, K. P., Clarke, J., Lambert, D. W., Prime, S. S., Underwood, T. J., Vijayanand, P., Eliceiri, K. W., Woelk, C., King, E. V., Gil, J., Ottensmeier, C. H. and Thomas, G. J. (2017). Induction of fibroblast senescence generates a non-fibrogenic myofibroblast phenotype that differentially impacts on cancer prognosis. Aging (Albany NY) 9(1): 114-132.
- Narra, K., Mullins, S. R., Lee, H. O., Strzemkowski-Brun, B., Magalong, K., Christiansen, V. J., McKee, P. A., Egleston, B., Cohen, S. J., Weiner, L. M., Meropol, N. J. and Cheng, J. D. (2007). Phase II trial of single agent Val-boroPro (Talabostat) inhibiting Fibroblast Activation Protein in patients with metastatic colorectal cancer. Cancer Biol Ther 6(11): 1691-1699.
- Ohlund, D., Handly-Santana, A., Biffi, G., Elyada, E., Almeida, A. S., Ponz-Sarvise, M., Corbo, V., Oni, T. E., Hearn, S. A., Lee, E. J., Chio, II, Hwang, C. I., Tiriac, H., Baker, L. A., Engle, D. D., Feig, C., Kultti, A., Egeblad, M., Fearon, D. T., Crawford, J. M., Clevers, H., Park, Y. and Tuveson, D. A. (2017). Distinct populations of inflammatory fibroblasts and myofibroblasts in pancreatic cancer. J Exp Med 214(3): 579-596.
- Perrot, I., Blanchard, D., Freymond, N., Isaac, S., Guibert, B., Pacheco, Y. and Lebecque, S. (2007). Dendritic cells infiltrating human non-small cell lung cancer are blocked at immature stage. J Immunol, 178(5): 2763-2769.
- Quatromoni, J. G., Singhal, S., Bhojnagarwala, P., Hancock, W. W., Albelda, S. M. and Eruslanov, E. 2015. An optimized disaggregation method for human lung tumors that preserves the phenotype and function of the immune cells. J Leukoc Biol 97(1): 201-209.
- Rupp, C., Scherzer, M., Rudisch, A., Unger, C., Haslinger, C., Schweifer, N., Artaker, M., Nivarthi, H., Moriggl, R., Hengstschläger, M., Kerjaschki, D., Sommergruber, W., Dolznig, H. and Garin-Chesa, P. (2014). IGFBP7, a novel tumor stroma marker, with growth-promoting effects in colon cancer through a paracrine tumor–stroma interaction. Oncogene, 34(7): 815-825.
- Servais, C. and Erez, N. (2013). From sentinel cells to inflammatory culprits: cancer-associated fibroblasts in tumour-related inflammation. J Pathol 229(2): 198-207.
- Sugimoto, H., Mundel, T. M., Kieran, M. W. and Kalluri, R. (2006). Identification of fibroblast heterogeneity in the tumor microenvironment. Cancer Biol Ther 5(12): 1640-1646.
- Tao, L., Huang, G., Song, H., Chen, Y. and Chen, L. (2017). Cancer associated fibroblasts: An essential role in the tumor microenvironment. Oncol Lett 14(3): 2611-2620.
- Waise, S., Parker, R., Rose-Zerilli, M. J. J., Layfield, D. M., Wood, O., West, J., Ottensmeier, C. H., Thomas, G. J. and Hanley, C. J. (2019). An optimised tissue disaggregation and data processing pipeline for characterising fibroblast phenotypes using single-cell RNA sequencing. Sci Rep 9(1): 9580.
- Witowski, J., Kawka, E., Rudolf, A. and Jorres, A. (2015). New developments in peritoneal fibroblast biology: implications for inflammation and fibrosis in peritoneal dialysis. Biomed Res Int 2015: 134708.
Article Information
Copyright
© 2019 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Waise, S., Parker, R., Rose-Zerilli, M. J. J., Layfield, D. M., Wood, O., West, J., Ottensmeier, C. H., Thomas, G. J. and Hanley, C. J. (2019). An Optimized Method to Isolate Human Fibroblasts from Tissue for Ex Vivo Analysis. Bio-protocol 9(23): e3440. DOI: 10.21769/BioProtoc.3440.
Category
Cancer Biology > General technique > Tumor microenvironment
Cell Biology > Cell isolation and culture > Cell isolation
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link








