- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Measurement of Mitotic Spindle Angle and Mitotic Cell Distance in Fixed Tissue of Drosophila Larval Brains
Published: Vol 9, Iss 22, Nov 20, 2019 DOI: 10.21769/BioProtoc.3432 Views: 3886
Reviewed by: Sunanda MarellaPradeep Kumar BhaskarAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
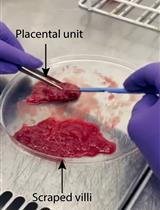
Isolation of First-Trimester and Full-term Human Placental Hofbauer Cells
Anna Appios [...] Naomi McGovern
Jun 5, 2021 4687 Views
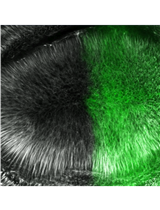
Ex-vivo Microtubule Stability Assay Using Drosophila Wing Disc
Jung-Wan Mok and Kwang-Wook Choi
Dec 5, 2021 3104 Views
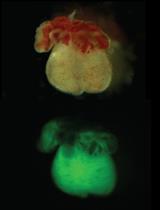
Isolation of Embryonic Cardiomyocytes and Cell Proliferation Assay Using Genetically Engineered Reporter Mouse Model
Maren Beall [...] Jihyun Jang
Sep 5, 2023 1798 Views
Abstract
The positioning and the cleavage plane orientation of mitotic cells in pseudostratified epithelia (PSE) must be tightly regulated since failures in any of these processes might have fatal consequences during development. Here we present a simple method to determine the spindle orientation as well as the positioning of neuroepithelial mitotic cells within the Outer Proliferation Center (OPC) of Drosophila larval brains.
Keywords: MitosisBackground
Pseudostratified epithelia (PSE) are epithelia in which the interphase nuclei of the epithelial cells dynamically position at different depths within the epithelial monolayer, while mitotic nuclei localize at the most apical side of the epithelium. PSE are usually highly proliferative tissues acting as organ precursors in many organisms, including the building blocks of the epiblast in the gastrulating mouse embryo, the pancreatic buds and different areas of the central nervous system in vertebrates, as well as the imaginal discs and the optic lobe anlage in Drosophila (Meyer et al., 2011; Ichikawa et al., 2013; Strzyz et al., 2016).
Almost a century ago, Sauer observed that in PSE nuclei moved to the apical surface to divide (Sauer, 1935). The apical migration of nuclei is linked to the cell cycle progression, occurring at G2 phase, and it is known as Interkinetic Nuclear Migration (INM) or Pre-mitotic Rapid Apical Migration (PRAM). Thus, INM is responsible for the spatial organization of proliferating cells in PSE and the localization of mitotic nuclei at the PSE basal side is frequently linked to failures in INM (Tsuda et al., 2009). The apical “mitotic zone” serves as a niche for signaling and spatial control of mitotic entry, and it has been recently shown that apical daughter cells reintegrate better into the PSE (reviewed in Norden, 2017).
Better known is the control of the mitotic spindle orientation, which will determine tissue architecture or cell fate specification in the context of symmetric or asymmetric cell divisions, respectively. As failures in both processes are implicated in tumorigenesis, spindle misorientation might promote or, at least, facilitate tumor development (Pease and Tirnauer 2011; Noatynska et al., 2012, Bergstralh and St Johnston 2014). Epithelial cells tend to divide with their mitotic spindle parallel to the plane of the epithelial sheet. The significance of tightly regulating this process during early development, when epithelia are in a high proliferative phase, is manifest by the existence of several neurological diseases associated with mitotic spindle orientation failures, such as microcephaly, lissencephaly and Huntington disease (Noatynska et al., 2012).
Several protocols for 3D automated measurements of mitotic spindles have been recently published (Juschkeet al., 2014; Lázaro-Diéguez et al., 2014). Although these protocols are very reliable as they eliminate any bias due to hand processing of the images, they are established to work in vitro, in cell culture. In Franco and Carmena (2019), we have developed a simple and reliable method to determine the angle of the mitotic spindle and the positioning of the mitotic nucleus of neuroepithelial cells in fixed whole brains of Drosophila third instar larvae, making possible to visualize in vivo, the morphogenetic consequences of failures in any of these processes. Since planar divisions and INM are considered hallmarks of some neural progenitors, these measurements can be an effective way of studying the architecture and organization of a developing neuroepithelium.
Materials and Reagents
- Dissection wells (Winlab, catalog number: 190029500)
- Dissection forceps (Fine Science Tools, Dumont, catalog number: 55)
- Glass slides and covers
- Adhesive tape
- Yellow and white pipette tips
- Ice bucket
- Drosophila early third instar larvae (60 h after larval hatching ALH)
- BSA (Albumin from bovine serum, Sigma-Aldrich, catalog number: A3059)
- Primaries antibodies against a spindle pole protein, for example an anti-γTubulin (Mouse monoclonal anti-γTubulin, Sigma-Aldrich, catalog number: T5326), and against Phospho-Histone 3 (PH3) to determine metaphasic nuclei (Rabbit polyclonal anti-PH3 (Ser10), Millipore, catalog number: 06-570)
- Secondary antibodies coupled to Alexa Fluor 488 (Goat anti-Mouse IgG, Alexa Fluor Plus 488, Invitrogen, catalog number: A32723) and 555 (Goat anti-Rabbit IgG, Alexa Fluor Plus 555, Invitrogen, catalog number: A32732)
- Na2HPO4
- KCl
- NaCl
- Triton X-100 (Sigma-Aldrich, catalog number: T8787)
- PFA 16% solution (EM grade, Electron Microscopy Sciences, catalog number: 15710)
- Phalloidin linked to a fluorophore (Alexa Fluor 633 phalloidin, Invitrogen, catalog number: A22284), used to determine the cellular and tissue morphology
- Vectashield Mounting medium (Linaris, catalog number: H-1000)
- Phosphate-buffered saline (PBS) (see Recipes)
- PBT (see Recipes)
- Blocking Reagent (see Recipes)
- 4% Paraformaldehyde (PFA) in PBT-0.1T (see Recipes)
Equipment
- Orbital shaker
- Confocal microscope
- Autoclave
Software
- Lite Leika
- Image J
- Excel
Procedure
- Stain larval brains according to the following protocol
Note: All the washes and incubations are done on an orbital shaker at slow motion.- Roughly dissect about 15 brains per genotype in cold PBS. Try not to prolong more than 20 min the dissection time to preserve the quality of the material. Keep the material already dissected on ice-cold PBS.
- Fix by incubation in 4% PFA in PBT-0.1T for 20 min with gentle agitation.
- Rinse with PBT by replacing the solution 3 times and wash by gentle agitation for another 3 x 15 min each. During these washes, clean the brains of other pieces of tissue sticking to it.
- Incubate brains in PBT + 0.1% BSA for at least 1 h at Room Temperature (RT).
- Incubate brains with the corresponding primary antibodies at the right dilution in blocking reagent overnight at 4 °C.
- Rinse with PBT by replacing the solution 3 times and wash for another 3 x 15 min each.
- Incubate with secondary antibodies at the right dilution in blocking reagent for 2 h at RT in the dark.
- Rinse with PBT by replacing the solution 3 times and wash for another 3 x 15 min each.
- Wash with PBS for 10 min at RT.
- After removing the PBS, add Vectashield to the well and leave at 4 °C overnight (this incubation facilitates the mounting step).
- Mount brains onto glass slides using the bridge method illustrated in Figure 1, preserving their three-dimensional configuration.
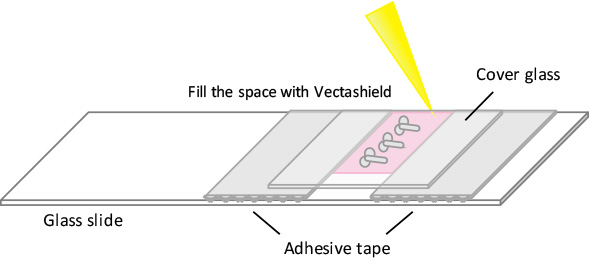
Figure 1. Bridge method for mounting Drosophila brains - Glue two pieces of adhesive tape leaving a thin space (approx. 5 mm) in between as shown in Figure 1.
- Add the brains in the smallest possible volume of Vectashield with a cut yellow pipette tip.
- Remove as much as possible the excess of Vectashield with a cut white pipette tip.
- Arrange the brains with the ventral side facing up.
- Add a cover glass and fix it with 2 pieces of adhesive tape.
- Fill the space under the bridge with Vectashield.
- Imaging acquisition
Fluorescent images were recorded using a Leica upright DMR microscope with an HCX Plan Apochromat 63x/1.32-0.6 NA oil confocal scanning objective. 2 x Zoom with a distance between focal planes in each Z-stack of 0.8 μm was used.
Data analysis
Open the Z-stack with ImageJ (shown in Figure 2)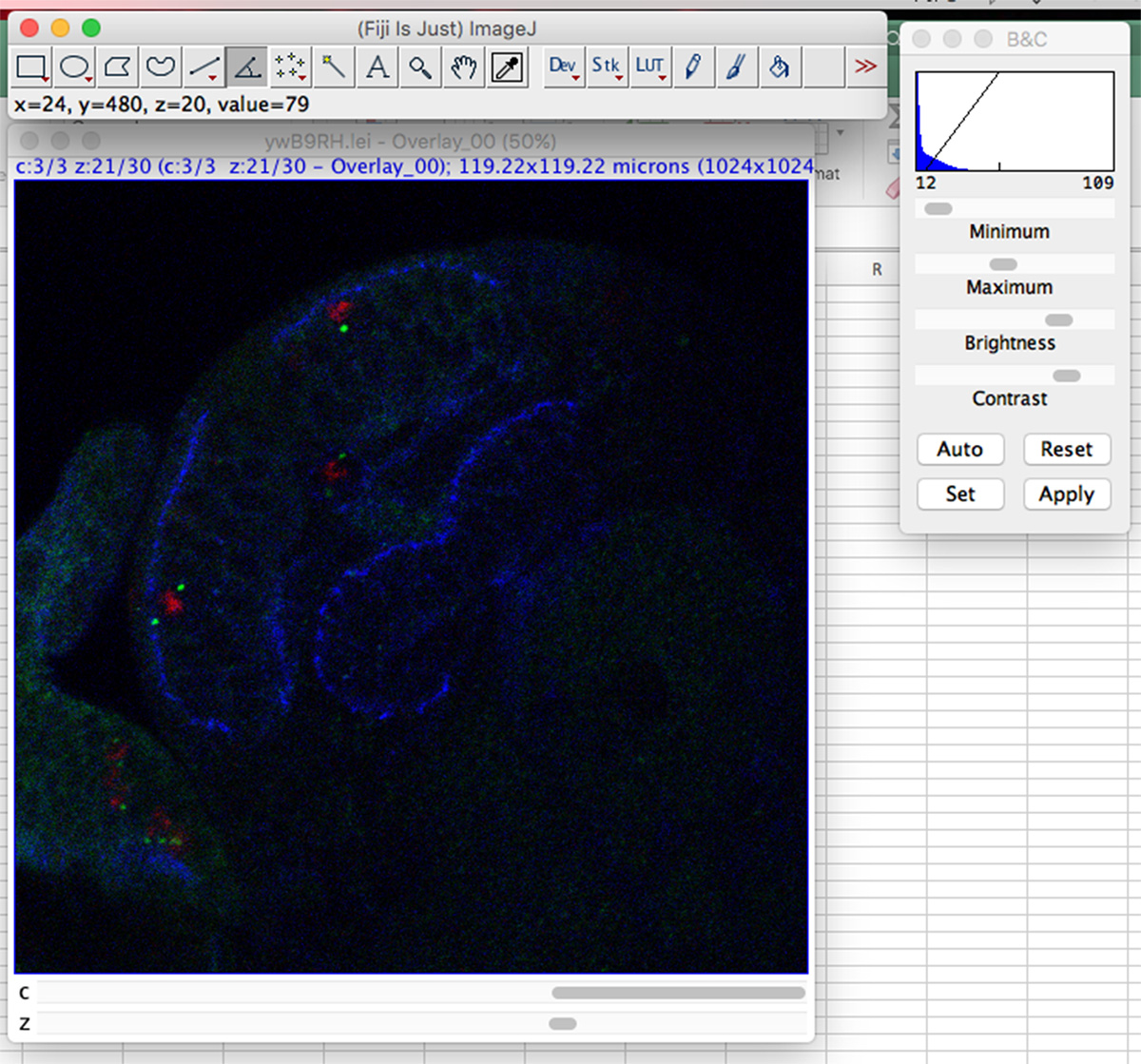
Figure 2. A Z-stack of Drosophila early third instar larval brain hemisphere. PH3, γ-Tubulin and Phalloidin staining are shown in red, green and blue, respectively.
Note: It is important to analyze only the portion of neuroepithelium where the cells look more perpendicular to the longitudinal neuroepithelial plane.
- Spindle angle measurement
- Select the “angle” tool and draw a line linking both centrosomes, press the mouse button and draw an angle respect to a second line running parallel to the apical surface of the neuroepithelium, as it is shown in Figure 3. Then press <m> and a window with the measurement results will appear.
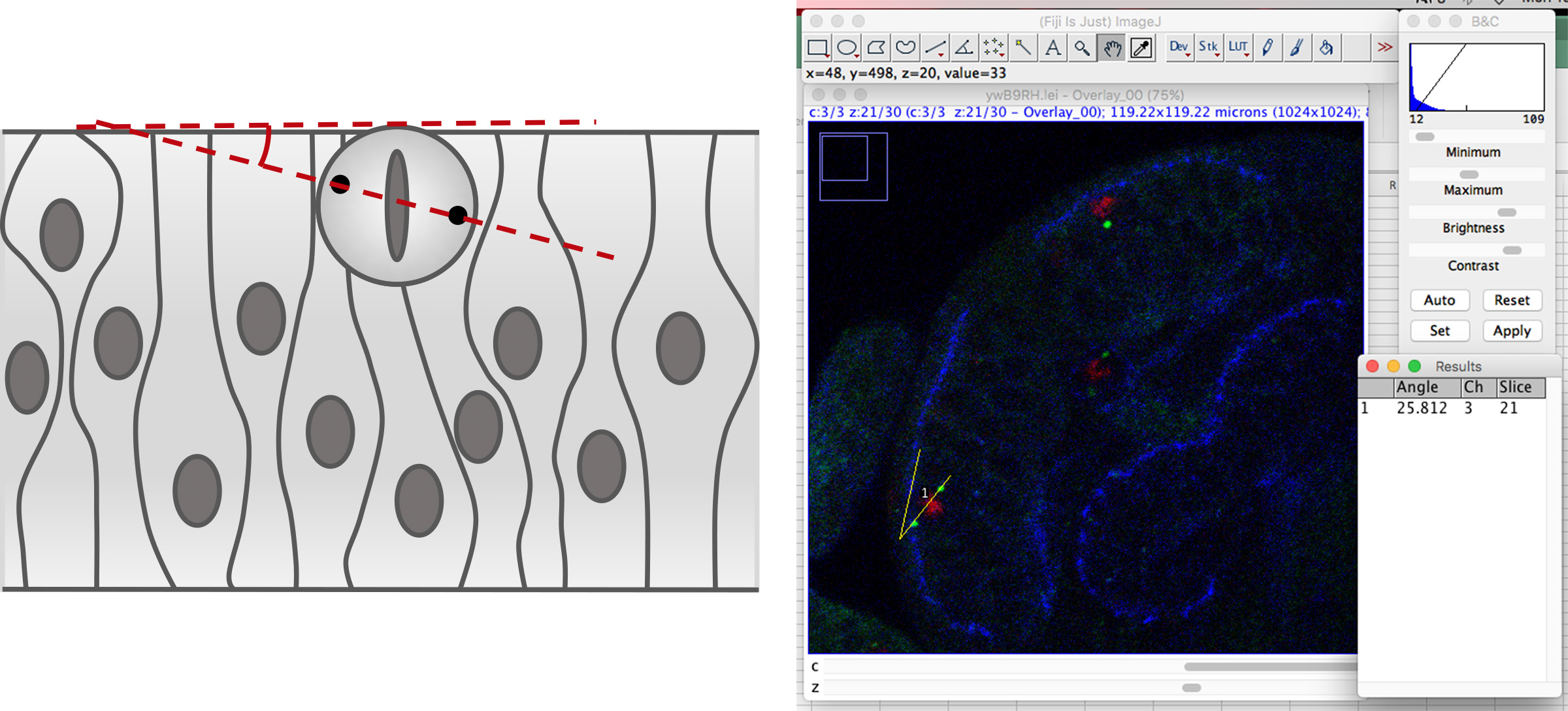
Figure 3. Measurement of the mitotic spindle angle
Note: These measurements frequently require correction in the xy plane to allow that both spindle poles are present in a single Z-plane. To avoid overcorrection, only mitosis with centrosomes in the same or in two consecutive Z planes are considered for quantification. - Collect all the measurements.
- Save the results as an excel file.
- Open the document in Excel. Copy the column with the angle results and paste it in a new column (Column A in the example shown in Figure 4). Arrange the data in order (Column B in the example) and proceed with the following steps. Step 1: Transform the values above 90 degrees by applying: y = 180 - x (Column C in the example), where x is the value obtained in the measurement. Step 2: Divide all the data into categories (every 15 degrees, for example).
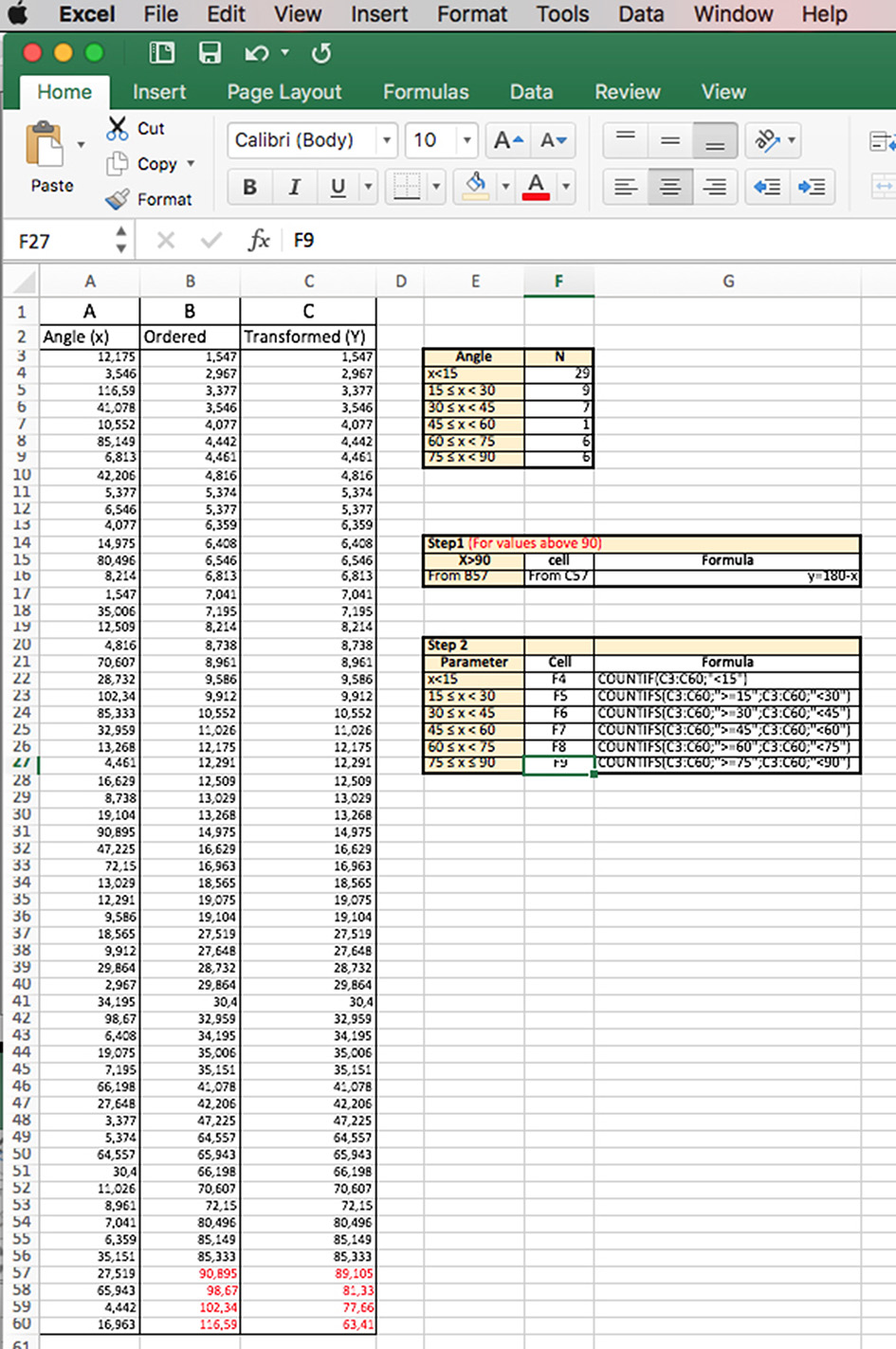
Figure 4. Excel sheet workup for data analysis of the mitotic spindle orientation
With the data arranged in this way, perform different statistic tests to see whether two populations have the same distribution of mitotic angles respect to the plane. It is recommendable to understand well and define properly the behavior of your control, as well as to define whether your data follow a Parametric distribution or not.
Note: Angles respect to the apical surface equal or minor than 30 degrees are considered wild type.
- Select the “angle” tool and draw a line linking both centrosomes, press the mouse button and draw an angle respect to a second line running parallel to the apical surface of the neuroepithelium, as it is shown in Figure 3. Then press <m> and a window with the measurement results will appear.
- Measurement of mitotic cell positioning
- Select the “straight line” tool and, in each mitotic cell, draw two lines: one from the apical surface to the middle of the metaphasic plate (and press <m>), and another from the apical to the basal side of the epithelium where the mitotic cell is located (and press <m>). These lines correspond to “a” and “b”, respectively, in Figure 5.
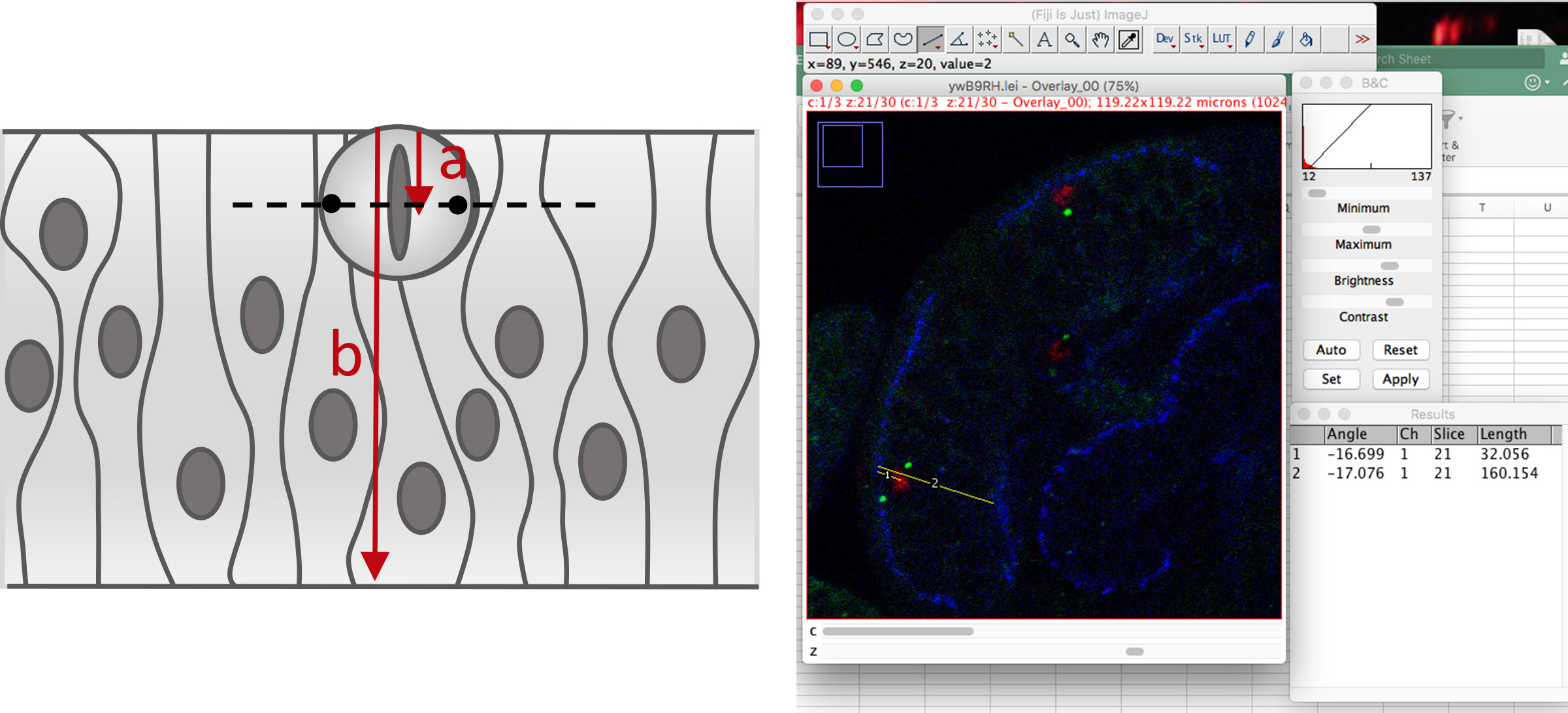
Figure 5. Measurement of mitotic cells relative positioning within the neuroepithelium - Collect all the measurements.
- Save the results as an excel file.
- Open the document in Excel.
- Calculate (Column D in the example in Figure 6) the Relative Position (%) of the mitotic plate respect to the apical side of the epithelium for every cell, by applying:
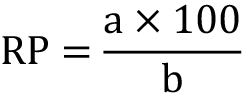
where “a” is the distance from the apical surface to the mitotic plate and “b” is the whole width of the epithelia at this position (see also Figure 5). - Order all the data and see how the mitotic nuclei arrange along the apico-basal axis of the NE.
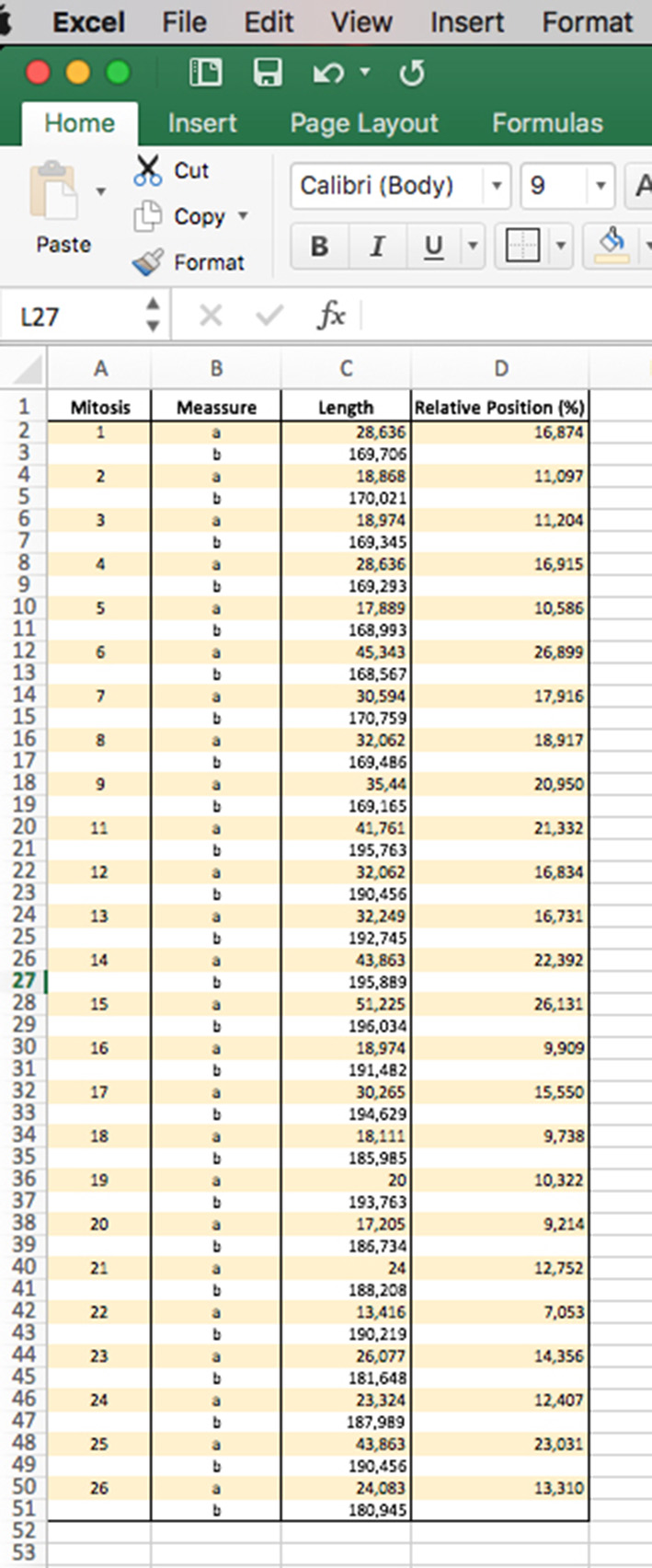
Figure 6. Excel sheet workup for data analysis of the relative mitotic cell distance - With the data arranged in this way, represent the distribution of the mitotic cells along the apico-basal axis of the neuroepithelium. Then, perform different statistic tests to analyze possible differences between two samples.
- Select the “straight line” tool and, in each mitotic cell, draw two lines: one from the apical surface to the middle of the metaphasic plate (and press <m>), and another from the apical to the basal side of the epithelium where the mitotic cell is located (and press <m>). These lines correspond to “a” and “b”, respectively, in Figure 5.
Having measured both parameters (mitotic spindle orientation and mitotic cell distance) for a given genotype, correlations to see whether one parameter has an effect on the other can be performed. For example, in Franco and Carmena (2019), we determined that 80% of the cells in the control divide in the most apical part (0-40%) of the NE. Even though we could not find a straight correlation between failures in positioning and spindle orientation, we could conclude that basal mitosis was more prone to show defects in mitotic spindle alignment.
Recipes
- PBS
10 mM Na2HPO4
2.68 mM KCl
140 mM NaCl
pH 7.4
Sterilize by autoclaving - PBT
PBS with 0.3% Triton - Blocking Reagent
PBT with 0.1% BSA - 4% Paraformaldehyde (PFA) in PBT-0.1T
Dilute 16% PFA in PBS with 0.1333 % Triton
Acknowledgments
Our lab was supported by the Spanish grants from the Ministry of Economy and Competitiveness (MINECO) BFU2012-33020 and BFU2015-64251, and by FEDER (European Regional Development Fund). The Instituto de Neurociencias in Alicante is a “Severo Ochoa” Center of Excellence. The original research paper where this protocol was used is Franco and Carmena, 2019.
Competing interests
The authors declare no competing interests.
References
- Bergstralh, D. T. and St Johnston, D. (2014). Spindle orientation: what if it goes wrong? Semin Cell Dev Biol 34: 140-145.
- Franco, M. and Carmena, A. (2019). Eph signaling controls mitotic spindle orientation and cell proliferation in neuroepithelial cells. J Cell Biol 218(4): 1200-1217.
- Ichikawa, T., Nakazato, K., Keller, P. J., Kajiura-Kobayashi, H., Stelzer, E. H., Mochizuki, A. and Nonaka, S. (2013). Live imaging of whole mouse embryos during gastrulation: migration analyses of epiblast and mesodermal cells. PLoS One 8(7): e64506.
- Juschke, C., Xie, Y., Postiglione, M. P. and Knoblich, J. A. (2014). Analysis and modeling of mitotic spindle orientations in three dimensions. Proc Natl Acad Sci U S A 111(3): 1014-1019.
- Lazaro-Dieguez, F., Ispolatov, I. and Musch, A. (2015). Cell shape impacts on the positioning of the mitotic spindle with respect to the substratum. Mol Biol Cell 26(7): 1286-1295.
- Meyer, E. J., Ikmi, A. and Gibson, M. C. (2011). Interkinetic nuclear migration is a broadly conserved feature of cell division in pseudostratified epithelia. Curr Biol 21(6): 485-491.
- Noatynska, A., Gotta, M. and Meraldi, P. (2012). Mitotic spindle (DIS)orientation and DISease: cause or consequence? J Cell Biol 199(7): 1025-1035.
- Norden, C. (2017). Pseudostratified epithelia-cell biology, diversity and roles in organ formation at a glace. J Cell Sci 130(11): 1859-1863.
- Pease, J. C. and Tirnauer, J. S. (2011). Mitotic spindle misorientation in cancer--out of alignment and into the fire. J Cell Sci 124(Pt 7): 1007-1016.
- Sauer, F. C. (1935). Mitosis in the neural tube. J Comp Neurol 62(2): 337-405.
- Strzyz, P. J., Matejcic, M. and Norden, C. (2016). Heterogeneity, cell biology and tissue mechanics of pseudostratified epithelia: coordination of cell divisions and growth in tightly packed tissues. Int Rev Cell Mol Biol 325: 89-118.
- Tsuda, S., Kitagawa, T., Takashima, S., Asakawa, S., Shimizu, N., Mitani, H., Shima, A., Tsutsumi, M., Hori, H., Naruse, K., Ishikawa, Y. and Takeda, H. (2010). FAK-mediated extracellular signals are essential for interkinetic nuclear migration and planar divisions in the neuroepithelium. J Cell Sci 123(Pt 3): 484-496.
Article Information
Copyright
© 2019 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Readers should cite both the Bio-protocol article and the original research article where this protocol was used:
- Franco, M. and Carmena, A. (2019). Measurement of Mitotic Spindle Angle and Mitotic Cell Distance in Fixed Tissue of Drosophila Larval Brains. Bio-protocol 9(22): e3432. DOI: 10.21769/BioProtoc.3432.
- Franco, M. and Carmena, A. (2019). Eph signaling controls mitotic spindle orientation and cell proliferation in neuroepithelial cells. J Cell Biol 218(4): 1200-1217.
Category
Developmental Biology > Cell signaling > Mitotic spindle orientation
Developmental Biology > Cell growth and fate > Proliferation
Cell Biology > Cell-based analysis > Mitotic spindle orientation
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link









