- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Cell Wall Compositional Analysis of Rice Culms
Published: Vol 9, Iss 20, Oct 20, 2019 DOI: 10.21769/BioProtoc.3398 Views: 5736
Reviewed by: Ali ParsaeimehrJinping ZhaoAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

Quantification of Ethylene Production in Leaf and Bud Tissue of the Subtropical Tree Crop Litchi (Litchi chinensis Sonn.) Using Gas Chromatography and Flame Ionization Detection
Regina B. Cronje and Arnoldus J. Jonker
Mar 20, 2023 1589 Views
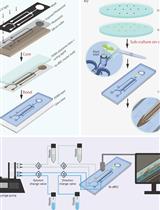
Bi-directional Dual-flow-RootChip for Physiological Analysis of Plant Primary Roots Under Asymmetric Perfusion of Stress Treatments
Claudia Allan [...] Claudia-Nicole Meisrimler
Aug 5, 2023 1988 Views

Enzymatic Starch Quantification in Developing Flower Primordia of Sweet Cherry
Nestor Santolaria [...] Afif Hedhly
Apr 5, 2025 1934 Views
Abstract
The plant cell wall is a complicated network that is mainly constituted of polysaccharides, such as cellulose, hemicellulose and pectin. Many noncellulosic polysaccharides are further acetylated, which confers these polymers flexible physicochemical properties. Due to the significance of cell wall in plant growth and development, the analytic platform has been the focus for a long time. Here, we use internodes/culms, an important organ to provide mechanical support for rice plants, as an experimental sample to explore the method for cell wall composition analysis. The method includes preparation of cell wall residues, sequential extraction of polysaccharides, and measurement of cellulose. The procedure for acetate examination is also described. This method is applicable to determine the composition of individual cell wall polymers and the modifier acetates, and is suitable to identify cell wall relevant mutants based on the advantages in high throughput, precision and repeatability.
Keywords: XylanBackground
The plant cell wall represents one of the most complicated cellular structure in nature and is essential for plant growth and adaptations to environments. Besides presenting multiple polysaccharide components and phenolic compounds, acetylation is a prevalent modification on most cell-wall polymers, which alters the physicochemical properties and increases the complexity of cell wall structure. Establishment of the effective analytic platform for cell wall composition is always a challenging task. The previous analytic method often uses alkali to extract cell wall residues, but removes acetate. Recent works have revealed that acetate patterns on xylan determine the folding of this polymer and impact the binding to cellulose or lignin, indicating its importance in cell wall formation and plant growth control (Grantham et al., 2017; Kang et al., 2019; Zhang et al., 2019). The method that can simultaneously examine the composition of diverse cell-wall polysaccharides and their acetyl modifications needs to be developed. It becomes realizable as solvent dimethyl sulfoxide has been found extracting xylan without trimming acetyl esters (Goncalves et al., 2008). Rice culms are representative for cell wall composition methodology analysis because this organ is rich of secondary wall-bearing fiber cells and also contains multiple cell types. In addition to abundant materials, acetylation level varies on different wall polymers and during the culm development. By using rice culms as analytic samples, we developed a protocol for cell wall composition and acetyl modification analyses with some changes from the previous method (Foster et al., 2010). This protocol offers a widely used way to examine the composition of diverse cell wall polymers and determine the acetate content in different rice varieties and other crops.
Materials and Reagents
- 96-well flat bottom assay plate (Greiner bio-one, catalog number: 655180)
- UV capable 96-well flat bottom assay plate (Corning, catalog number: 3635)
- Glass bottle
- Eppendorf tubes (1.5 ml) (Eppendorf, catalog number: 0030120.086)
- Sarstedt tubes 2 ml (Sarstedt, D-51588)
- 50 ml plastic centrifuge tube (Corning CentriStar)
- Glass microfiber filters (Whatman, catalog number: 1820-025)
- Rice mature plants
- Endopolygalacturonase M2 (Megazyme, catalog number: PGALUSP, 4 °C)
- Pectin methyl esterase (Sigma-Aldrich, catalog number: P5400-1KU, −20 °C)
- α-amylase (Megazyme, catalog number: E-BLAAM, 4 °C)
- ddH2O
- Acetatic Acid Assay Kit (Megazyme, catalog number: K-ACET, 4 °C)
- Acetone
- DMSO (Sigma-Aldrich, catalog number: D5879)
- 70% (v/v) aqueous ethanol
- Chloroform/methanol (1:1, v/v) solution
- Updegraff reagent (Acetic acid: nitric acid: water, 8:1:2 v/v)
- 72% Sulfuric acid (Prepared with concentrated Sulfuric acid GR)
- 1 mg/ml glucose stock (Prepared from D-(+)-glucose) (Sigma-Aldrich, catalog number: G8270, −20 °C)
- Anthrone reagent (2 mg/ml Anthrone in concentrated sulfuric acid) (Sigma-Aldrich, catalog number: 319899)
- Trifluoroacetic acid (TFA) (Sigma-Aldrich, catalog number: T6508)
- Ammonium formate (Aldrich, catalog number: 516961)
- 11% peracetic acid solution (prepared from 35% peracetic) (Aladdin, catalog number: P112625)
- Ethanol: methanol: water solution (7:2:1, adjust the pH to 3.0 with HCOOH)
- 1% ammonium oxalate (Sigma-Aldrich, catalog number: 09898)
- 37% hydrogen chloride
- MES/Tris buffer (pH 8.1-8.3) (see Recipes)
- 2 M trifluoroacetic acid (see Recipes)
- 1 N sodium hydroxide (see Recipes)
- 1 N hydrogen chloride (see Recipes)
- 50 mM ammonium formate (pH 4.5) (see Recipes)
Equipment
- Freeze dryer (Beijing Songyuanhuaxing Technology Develop Co. Ltd., model: LGJ-12)
- Ball mill (QIAGEN, TissueLyser II, catalog number: 85300)
- (Optional) Vortex shaker
- Basket centrifuge (Eppendorf, model: 5810R)
- Centrifuge (Eppendorf, model: 5430) (to fit Eppendorf 1.5 ml tubes)
- Thermomixer comfort (Eppendorf)
- Dri-Block heaters (Techne, model: DB200/3)
- Microplate reader (PerkinElmer, Enspire)
- Concentrator (Eppendorf, concentrator plus)
- (Optional) Drying oven
- (Optional) Shaking incubator
- Sieves (Mesh size of 0.15 mm)
- pH meter (Mettler Toledo)
- Semi-micro scales (dual resolutions starting at 0.01 mg)
Procedure
- Preparation of destarched alcohol-insoluble cell-wall residues (AIR)
- Pool the whole 2nd internodes (numbered from the top down) of 5-20 rice mature plants.
- Freeze the fresh samples in liquid nitrogen and then lyophilize them in a freeze dryer (The rice internodes were lyophilized for 48 h to ensure complete dryness).
- Grind tissues to a particle size no more than 0.15 mm using ball mill and sieve through mesh with a size of 0.15 mm.
- Weigh approximately 1 g of the ground plant biomass into a 50 ml plastic centrifuge tube.
- Add 30 ml of 70% (v/v) aqueous ethanol, mix thoroughly using a vortex mixer and leave in a thermomixer comfort set at 37 °C and 200 rpm for 12 h.
- Centrifuge at 1,500 x g for 10 min in a basket centrifuge and discard the supernatant.
- Repeat Steps A5-A6 once.
- Add 30 ml of the chloroform/methanol (1:1 v/v) solution, mix thoroughly using a vortex mixer and leave in a shaking incubator for 30 min at 37 °C and 200 rpm.
- Centrifuge at 1,500 x g for 10 min at room temperature and discard the supernatant.
- Repeat Steps A8-A9 twice.
- Add 15 ml of acetone, shake the tube to re-suspend the pellet.
- Centrifuge at 1,500 x g for 10 min and discard the supernatant.
- Repeat Steps A11-A12 twice.
- Let the biomass samples dry in a drying oven at 40 °C without shaking for approximately 16 h.
- Treat the residues with 100 U α-amylase in 40 ml MES/Tris buffer (pH 8.1) at 97 °C for 35 min, then 60 °C for 1 h.
- Centrifuge at 1,500 x g for 10 min and discard the supernatant.
- Wash the pellet with 30 ml ddH2O three times and with 15 ml acetone twice, with centrifugation (1,500 x g for 10 min) and supernatant removal after each wash.
- Let the biomass samples dry in an oven at 40 °C for approximately 16 h to get destarched AIR.
- Analysis of the crystalline cellulose content
- Weigh 2 mg AIR material in five replicates into 2 ml Sarstedt tubes.
- Add 250 µl of 2 M trifluoroacetic acid (TFA) to each sample and make ensure no material is splashed up onto the tube walls.
- Cap tightly and incubate for 90 min at 121 °C in Dri-Block heaters.
- Cool the heating blocks and samples on ice.
- Centrifuge at 11,000 x g for 10 min, then transfer the supernatant to a new Sarstedt tube for optional noncellulosic polysaccharides composition analysis (Foster et al., 2010) and keep the pellet for crystalline cellulose assay.
- Add 1 ml of Updegraff reagent (Acetic acid: itric acid: water, 8:1:2 v/v) to the pellets left over from the Step B5, cap tightly and vortex.
- Heat in Dri-Block heaters at 100 °C for 30 min.
- Cool samples on ice.
- Centrifuge samples at 11,000 x g for 10 min.
- Discard supernatant ensuring no pellet material is discarded.
- Wash once with 1 ml water and four times with 1 ml acetone, centrifuge and discard supernatant as done above.
- Air dry the pellet in the Dri-Block heaters at 35 °C.
- Add 175 µl 72% Sulfuric acid and incubate at room temperature for 60 min.
- Add 825 µl ddH2O, vortex and centrifuge samples at 11,000 x g for 5 min.
- Analyze the glucose content of the supernatant using the anthrone assay in a 96-well flat bottom assay plate.
- Add 10 µl of sample and 90 µl of ddH2O for a total volume of 100 µl in each sample well.
- Prepare standards using 1 mg/ml glucose stock (stock at -20 °C). Make 0, 2, 4, 6, 8, and 10 µg standards by pipetting 0, 2, 4, 6, 8, and 10 µl into the appropriate well, adding 100, 98, 96, 94, 92, and 90 µl ddH2O accordingly.
- Add 200 µl freshly prepared Anthrone Regent.
- Heat the plate for 30 min at 80 °C with shaking in a thermomixer comfort. Cool to room temperature.
- Shake the plate thoroughly in a thermomixer comfort and then read absorption at 625 nm within 1 h.
- Extraction of pectin from cell wall residues
- Weigh about 6 mg destarched AIR for 5 replicates into 1.5 ml Eppendorf tubes.
- Add 1 ml of 50 mM ammonium formate (pH 4.5) buffer, then add 2 U of endopolygalacturonase M2 and 0.04 U of pectin methyl esterase. Mix and incubate at 37 °C for 16 h with shaking at 300 rpm in a thermomixer comfort.
- Centrifuge at 3,000 x g for 10 min and transfer the pectin-rich supernatants to a new tube. Keep the remains as pectin-free samples.
- Freeze the supernatants using liquid nitrogen and then lyophilize them in a freeze dryer, finally get the gelatinous pectin.
- Isolation of the acetyl-xylan from AIR
- Weigh about 400 mg of destarched AIR into a 50 ml centrifuge tube.
- Add 30 ml 1% ammonium oxalate and incubate at 85 °C for 2 h. Centrifuge at 1,500 x g for 15 min in a basket centrifuge. Discard the supernatant to remove pectin.
- Add 30 ml 11% peracetic acid solution, incubate in a water bath at 85 °C for 30 min for delignification.
- Centrifuge at 1,500 x g for 10 min. Discard the supernatant and wash the pellets with 30 ml ddH2O three times and with 15 ml acetone once, through centrifugation (1,500 x g for 10 min) and supernatant removal.
- Let the pellets dry in an oven set at 40 °C for approximately 16 h.
- Add 30 ml DMSO and then incubate at 70 °C for 12 h to conduct extraction, centrifuge at 1,500 x g for 10 min, transfer the supernatant to a new glass bottle.
- Repeat Step D6 with 12 ml DMSO, and combine the supernatants containing the extractives.
- Filter the extracts through glass microfiber filters.
- Precipitate the extracts with 5 volume of ethanol: methanol: water solution (7:2:1, pH 3.0) at 4 °C for 12 h.
- Centrifuge at 1,500 x g for 15 min. Discard the supernatant and wash the pellets four times each with 10 ml anhydrous ethanol with centrifugation (1,500 x g for 10 min) and supernatant removal after each wash.
- Dry the pellets under vacuum in a concentrator at room temperature.
- Determining the content of acetyl esters
- Weigh about 1 mg destarched AIRs/pectin/xylan for 5 replicates into 1.5 ml Eppendorf tubes.
- Add 100 µl 1 N sodium hydroxide to tubes and shake for 1 h at 28 °C and 200 rpm to release the bound acetate.
- Add 100 µl of 1 N hydrogen chloride to neutralize samples.
- Centrifuge at 15,000 x g for 10 min, and transfer 10 µl supernatant aliquot to a UV capable 96-well flat bottom assay plate and immediately quantify released acetic acids according to the instruction of Acetic Acid Assay Kit.
- Add 94 µl of ddH2O to each sample well in the UV capable 96-well flat bottom assay plate.
- Add 42 µl of freshly prepared mixture of kit-supplied Solutions 1 and 2 (2.5:1, 30 µl + 12 µl each), mix and incubate at 25 °C for 3 min with shaking at 300 rpm in a thermomixer comfort.
- Read the absorption at 340 nm (A0).
- Add 12 µl of freshly diluted kit-supplied solution 3 (1:10, v/v, in ddH2O), mix and incubate at 25 °C for 4 min with shaking at 300 rpm in a thermomixer comfort.
- Read the absorption at 340 nm (A1).
- Add 12 µl of freshly diluted kit-supplied solution 4 (1:10, v/v, in ddH2O), mix and incubate at 25 °C for 12 min with shaking at 300 rpm in a thermomixer comfort.
- Read the absorption at 340 nm (A2).
- At the same time, it is necessary to make a blank control and a standard curve in parallel. To make a standard curve, add 5, 10, 15, 30, 50 µl acetic acid standard (solution 5) equal to 0.5, 1, 1.5, 3, 5 µg, and adjust the volume of ddH2O in Step E5 to 99, 94, 89, 74, 54 µl, respectively.
Data analysis
- Generation of standard curve and calculation of cellulose content
To calculate cellulose amount, the standard curve needs to be plotted out first (The absorbance of glucose standards versus the concentration of glucose standards). Then calculate cellulose content (µg glucose/mg AIR) of sample with standard curve. - For acetylation content analysis: Δacetic acid = (A2 - A0)sample - (A1 - A0)2sample/(A2 - A0)sample - [(A2-A0)blank - (A1 - A0)2blank/(A2 - A0)blank].
Locate the sample value in the standard curve, and get acetic acid content in the sample aliquot. Calculate sample contents by multiplying with a factor of (100 + 100)/10/weight (µg/mg AIR). - For statistical analysis, typically use at least five biological replicates from a pool of internodes. Results are reported as means and standard deviation and statistical significance is assessed by Student’s t-test or one-way ANOVA followed by Tukey’s multiple comparison test.
Notes
- If a freeze-dryer is not available, it could be alternative to prepare AIR directly from frozen tissue that is homogenized with a mortar and pestle immersed in liquid nitrogen or a ball mill pre-immersed in liquid nitrogen.
- For acetylation analysis, the acetic acid released should be assayed as soon as possible, do not leave too long before plate reading.
Recipes
- MES/Tris buffer (pH 8.1)
- Add 488 mg MES and 355 mg Tris in 40 ml double distilled water
- Adjust the pH to 8.1 with sodium hydroxide and finally dilute to 50 ml
- 2 M trifluoroacetic acid
Dilute 15.31 ml TFA with ddH2O to a final volume of 100 ml - 1 N sodium hydroxide
Add 2 g of sodium hydroxide to 50 ml of ddH2O - 1 N hydrogen chloride
Mix 4 ml 37% hydrogen chloride with 44 ml ddH2O - 50 mM ammonium formate (pH 4.5)
- Add 0.15 g of ammonium formate to 50 ml ddH2O
- Adjust the pH to 4.5 with formic acid
Acknowledgments
This protocol was adapted from previous work (Zhang et al., 2019). This work was supported by the National Natural Science Foundation of China (grant No. 31530051, 31571247, and 91735303), the Youth Innovation Promotion Association, Chinese Academy of Sciences (grant No. 2016094). This protocol is based on the following previously published reports: Updegraff (1969), Harholt et al. (2006), Goncalves et al. (2008), Li et al. (2009), Foster et al. (2010), Bromley et al. (2013), de Souza et al. (2014), Zhang et al. (2017).
Competing interests
The authors declare no conflict of interest.
References
- Bromley, J. R., Busse-Wicher, M., Tryfona, T., Mortimer, J. C., Zhang, Z., Brown, D. M. and Dupree, P. (2013). GUX1 and GUX2 glucuronyltransferases decorate distinct domains of glucuronoxylan with different substitution patterns. Plant J 74(3): 423-434.
- de Souza, A., Hull, P. A., Gille, S. and Pauly, M. (2014). Identification and functional characterization of the distinct plant pectin esterases PAE8 and PAE9 and their deletion mutants. Planta 240(5): 1123-1138.
- Goncalves, V. M., Evtuguin, D. V. and Domingues, M. R. (2008). Structural characterization of the acetylated heteroxylan from the natural hybrid Paulownia elongata/Paulownia fortunei. Carbohydr Res 343(2): 256-266
- Foster, C.E., Martin, T.M., Pauly, M. (2010). Comprehensive compositional analysis of plant cell walls (Lignocellulosic biomass): Part II. Carbohydrates. J Vis Exp 37: e1837.
- Grantham, N. J., Wurman-Rodrich, J., Terrett, O. M., Lyczakowski, J. J., Stott, K., Iuga, D., Simmons, T. J., Durand-Tardif, M., Brown, S. P., Dupree, R., Busse-Wicher, M. and Dupree, P. (2017). An even pattern of xylan substitution is critical for interaction with cellulose in plant cell walls. Nat Plants 3(11): 859-865.
- Harholt, J., Jensen, J. K., Sorensen, S. O., Orfila, C., Pauly, M. and Scheller, H. V. (2006). ARABINAN DEFICIENT 1 is a putative arabinosyltransferase involved in biosynthesis of pectic arabinan in Arabidopsis. Plant Physiol 140(1): 49-58.
- Kang, X., Kirui, A., Dickwella Widanage, M. C., Mentink-Vigier, F., Cosgrove, D. J. and Wang, T. (2019). Lignin-polysaccharide interactions in plant secondary cell walls revealed by solid-state NMR. Nat Commun 10(1): 347.
- Li, M., Xiong, G., Li, R., Cui, J., Tang, D., Zhang, B., Pauly, M., Cheng, Z. and Zhou, Y. (2009). Rice cellulose synthase-like D4 is essential for normal cell-wall biosynthesis and plant growth. Plant J 60(6): 1055-1069.
- Updegraff, D. M. (1969). Semimicro determination of cellulose in biological materials. Anal Biochem 32(3): 420-424.
- Zhang, B., Zhang, L., Li, F., Zhang, D., Liu, X., Wang, H., Xu, Z., Chu, C. and Zhou, Y. (2017). Control of secondary cell wall patterning involves xylan deacetylation by a GDSL esterase. Nat Plants 3: 17017.
- Zhang, L., Gao, C., Mentink-Vigier, F., Tang, L., Zhang, D., Wang, S., Cao, S., Xu, Z., Liu, X., Wang, T., Zhou, Y. and Zhang, B. (2019). Arabinosyl deacetylase modulates the Arabinoxylan acetylation profile and secondary wall formation. Plant Cell 31(5): 1113-1126.
Article Information
Copyright
© 2019 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Zhang, L., Zhang, B. and Zhou, Y. (2019). Cell Wall Compositional Analysis of Rice Culms. Bio-protocol 9(20): e3398. DOI: 10.21769/BioProtoc.3398.
Category
Plant Science > Plant physiology > Plant growth
Developmental Biology > Cell growth and fate > Cell wall
Cell Biology > Cell structure > Cell wall
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link









