- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Quantitation of TLR4 Internalization in Response to LPS in Thioglycollate Elicited Peritoneal Mouse Macrophages by Flow Cytometry
Published: Vol 9, Iss 18, Sep 20, 2019 DOI: 10.21769/BioProtoc.3369 Views: 4838
Reviewed by: Alka MehraMartin V KolevMarieta Ruseva

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

Novel Experimental Approach to Investigate Immune Control of Vascular Function: Co-culture of Murine Aortas With T Lymphocytes or Macrophages
Taylor C. Kress [...] Eric J. Belin de Chantemèle
Sep 5, 2025 3587 Views

Detection of Autophagy in Human Peripheral Blood Mononuclear Cells Using Guava® Autophagy and Flow Cytometry
Melanie Scherer [...] Jörg Bergemann
Sep 20, 2025 1445 Views
Protocol for the Isolation and Analysis of Extracellular Vesicles From Peripheral Blood: Red Cell, Endothelial, and Platelet-Derived Extracellular Vesicles
Bhawani Yasassri Alvitigala [...] Lallindra Viranjan Gooneratne
Nov 5, 2025 1510 Views
Abstract
Internalization of cell surface Toll-like Receptor 4 (TLR4) is a convenient read-out to measure LPS dependent activation of the TRIF adaptor pathway. We here provide a protocol to quantify the LPS dependent internalization of TLR4 using thioglycollate-elicited peritoneal macrophages by flow cytometry.
Keywords: TLR4Background
Toll-like Receptor 4 (TLR4) is unique among the Toll-like receptors in that it can deliver qualitatively different signals from two distinct cellular locations (reviewed in Takeda and Akira, 2004). Cell surface TLR4/MD2 is initially ligated by extracellular bacterial lipopolysaccharide (LPS) and engages the intracellular adaptor MAL and MyD88. Subsequently a fraction of cell surface LPS bound TLR4/MD2 will undergo CD14 and clathrin dependent internalization and translocation to an early endosomal compartment (Zanoni et al., 2011). In this endosomal compartment TLR4, engages the signaling adaptors TRAM and TRIF. Much of the regulation of TLR4 internalization and transport remains to be discovered. A flow cytometric method for quantifying loss of TLR4 surface expression, as a metric for endosomal translocation was first described in Bone Marrow Derived Macrophages (BMDMs) by Jonathan Kagan and colleagues (Kagan et al., 2008). Here we describe a modified version of this protocol to enable quantitation of TLR4 internalization in thioglycollate-elicited peritoneal macrophages on which we have published (Rajaiah et al., 2015; Perkins et al., 2018).
Materials and Reagents
- Consumables
- Pipette tips
- Ice buckets
- Pipettes
- 10 ml syringes (BD, catalog number: 309604)
- 18 G x 1 needles (BD, catalog number: 305195)
- 50 ml conical tubes (Denville, catalog number: C1062-P)
- FACS tubes (round), sterile, with caps (Corning, Falcon, catalog number: 352054)
- FACS tubes (round), non-sterile, without caps
- Cell strainers, 70 μm (Fisherbrand, catalog number: 22363548)
- 0.22 μm filters for filtering culture media and FACS buffer (Sarstedt, catalog number: 83.1823.001)
- Reagents
- Ultra pure LPS, purified from E. coli K235 by the hot phenol method as previously described (McIntire et al., 1967)
- Unconjugated rat IgG2a, λ anti-mouse CD16/32 (FC receptor) antibody, clone 93 (Biolegend, catalog number: 101302)
- PE-conjugated anti-mouse CD284 (TLR4) antibody, clone SA15-21 (Biolegend, catalog number: 145404)
- PE-conjugated Isotype control antibody, Rat IgG2a, κ, clone RTK2758 (Biolegend, catalog number: 400507)
- Optional: other antibodies for cellular identification (e.g. Alexafluor-conjugated anti mouse F480; Biolegend, catalog number: 123119)
- Optional: paraformaldehyde solution (PFA), 4% in PBS (Affymetrix, catalog number: 19943)
- Sterile Saline (Baxter, catalog number: 0338-0048-03)
- Optional: ACK lysis buffer (Quality Biological, Inc., catalog number: 118-156-721)
- Sterile Phosphate Buffered Saline (PBS) (Corning, catalog number: 21-040-CV)
- Culture Media (see Recipes)
- RPMI 1640 (Corning, catalog number: 15-040-CV)
- Fetal Bovine Serum (FBS) (Atlanta Biologicals, catalog number: S11550)
- Penicillin/Streptomycin (Gibco, catalog number: 15140-122)
- L-glutamine (Gibco, catalog number: 25030-081)
- FACS buffer (see Recipes)
- Sterile Phosphate Buffered Saline (PBS) (Corning, catalog number: 21-040-CV)
- EDTA, 0.5 M (Quality Biological, Inc., catalog number: 351-027-101)
- Fetal Bovine Serum (FBS) (Atlanta Biologicals, catalog number: S11550)
Equipment
- Light microscope with hemocytometer slide
- 37 °C incubator with CO2 control designated for cell culture work (no bacterial/viral work)
- Biosafety Cabinet designated for cell culture work (no bacterial/viral work) Sterile Culture Hood
- Vortex Genie
- Refrigerated Swing Bucket Centrifuge with inserts for 50 ml conical tubes and flow cytometry tubes
- Flow Cytometer (e.g., Becton Dickinson LSR II)
- Computer for analysis
Software
- FCS Express 6.0 Flow (De Novo Software; Glendale, CA USA)
- GraphPad Prism 7 (Graph Pad Software; La Jolla, CA USA)
Procedure
Day 1
- Collect macrophages from previously thioglycollate injected animals by peritoneal lavage using 2 x 10 ml of sterile saline, using the 10 ml syringe with an 18 G needle attached. We usually pool peritoneal exudates from 2-5 littermates of the same gender to ensure sufficient yield for all controls and time points. A typical protocol can be found here: https://www.jove.com/video/1488/isolation-of-mouse-peritoneal-cavity-cells.
- Pellet cells by centrifugation at 93-218 x g (1,000 rpm on a Thermo Scientific TX-750 4 rotor) for 10 min, and resuspend in culture media.
- Count cells using a hemocytometer (or cell counter) and add 1 x 106-1.5 x 106 Thioglycollate elicited peritoneal macrophages (TEPMs) to each sterile FACS tube in 1.5 ml culture media (see Recipes below). A typical experiment includes three time points in unstimulated and three timepoints in stimulated macrophages. For each timepoint, you can set up tubes as following:
1) A tube for unstained control, 2) A tube for isotype control, 3) A tube for unstimulated control with TLR4 antibody stain, 4) A tube for LPS stimulation with TLR4 antibody stain, 5) single-color controls if cells are to be co-stained with other cell markers, and 6) A tube for dual color stain if using. Lightly recap FACS tubes to allow gas exchange in the incubator. - Allow TEPM to rest overnight in a 37 °C incubator with 5% CO2.
Day 2
- The following morning, add 100 ng/ml ultrapure E. coli K235 LPS to each tube to be used to measure TLR4 internalization (excepting unstimulated controls). Lightly recap FACS tubes and return to the 37 °C incubator during stimulation.
- At appropriate time points, transfer each tube of control or LPS stimulated TEPMs to an ice bath and add two volumes ice-cold culture media or FACS buffer (see Recipes below) to arrest TLR4 internalization.
- Pellet cells in a pre-cooled swing bucket by centrifuging at 112-264 x g (1,100 rpm on TX-750 4 rotor) for 5-10 min.
Note: For this and all subsequent washes: decant media, resuspend cells by gently running the flow tubes over an empty tube rack, and immediately return to ice bucket. If more time points will be collected, resuspend these cells in ice-cold culture media and keep on ice until the last samples are removed from the incubator. - Wash cells twice more with ice-cold FACS buffer. Resuspend in 100 μl final volume of ice-cold FACS buffer.
- Add 2 μg αFC CD16/32–(approx. 4 μl of Biolegend #101302) Ab and incubate for 15 min on ice. This prevents FC mediated nonspecific antibody binding.
- Add 0.5 μg PE labeled anti-TLR4 Ab or Isotype control (optional: co-stain with other antibodies to aid in identification of cellular subsets) and incubate for 30 min on ice.
- Wash cells 3 x with ice-cold FACS buffer.
- Wash 1 x with ice-cold PBS.
- Either proceed to Flow Cytometry run immediately or fix cells using 1% PFA (final concentration) in 100 μl PBS. Resuspend cells in 0.5-1.0 ml PBS and pass samples through a 70 μm cell strainer into a clean flow tube (not required to be sterile) to remove clumps just prior to running on a flow cytometer.
Data analysis
- Flow cytometry analysis
- For flow cytometry analysis, select single cells by forward scatter (FSC) profile (Height and Area of peaks). Set “stop gate” to ≥ 20,000 single events for each sample. Leave “record gate” to record all events.
- For analysis, gate events on FSC-A and SSC-A profile to exclude dead cells and cellular debris.
- TLR level can be visualized in a histogram for PE intensity.
- Data (all events) are exported as FCS3.0 files.
- Data analysis for the measurement of TLR4 endocytosis.
- Open data files in FCS Express 6.0 Flow to fine tune “single cell” and “live macrophage” gates.
- Record the median fluorescence intensity (MFI) for each sample.
- Subtract the MFI of the unstained cells (autofluorescence) from the MFI of each sample.
- Average the MFI of the “0 min time point” TLR4-stained samples (do this for each genotype, if multiple strains of mice were used)–and normalize the MFI data to the average (Time 0 min = 100%).
- The internalization rate is calculated as the % of TLR4 lost from the surface over time.
- Expected results: In wild type mice, the level of TLR4 surface expression in unstimulated cells should remain constant over the course of the experiment, and be well above the level of the isotype control-stained cells. LPS-stimulated cells should lose surface expression at a steady rate (at least over the first 90 min). For example please see Figure 1.
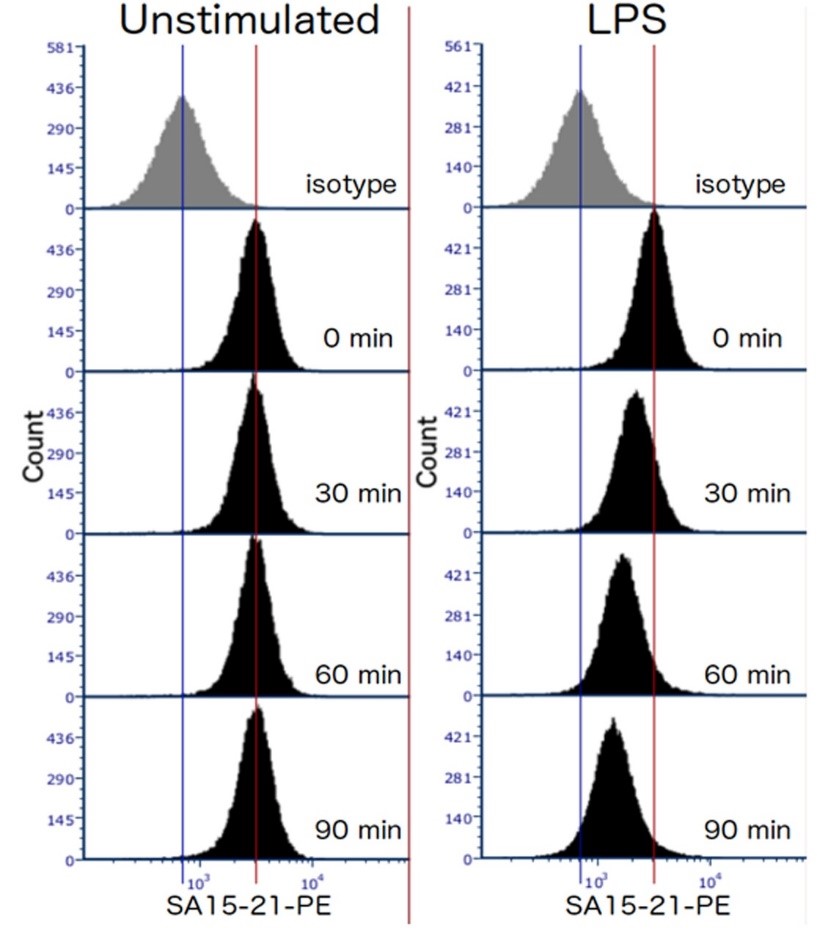
Figure 1. Representative data showing TLR4 internalization in response to 100 ng/ml E. coli LPS in TEPMs at the indicated time points. TEPMs were stained with anti-mouse TLR4 antibody (clone SA15-21) and analyzed by flow cytometry.
Notes
- We strongly recommend testing different lots of thioglycollate before starting these experiments, as endotoxin contamination is common and will pre-activate macrophages.
- The yield of thioglycollate-elicited macrophages can vary based not only on the lot of thioglycollate, but also on the experience level of the lab member performing the peritoneal lavage, and the age and strain of the mice. The following table (Table 1) shows the range of peritoneal lavage cell yields from our laboratory for 6-8 weeks old mice of commonly used strains.
Table 1. Typical yields of thioglycollate elicited macrophages. For each mouse genotype, expected yields of macrophages obtained following peritoneal lavage are provided. For C57BL/6J animals, average yields obtained by new investigators are also indicated.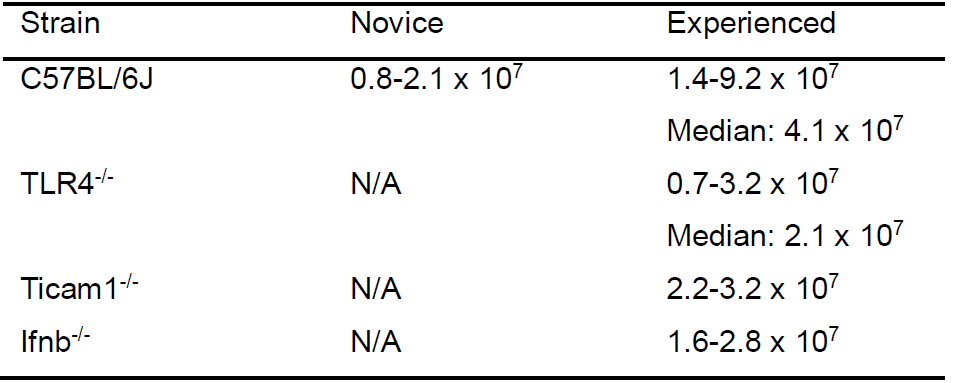
- If the cell pellets from the peritoneal lavage look bloody, we perform red blood cell lysis using ACK buffer, followed by two washes with sterile PBS before resuspending in culture media.
- It is important to allow the macrophages to rest overnight after isolation in order to achieve consistent kinetics of internalization from experiment to experiment.
- Inspect each tube before adding LPS. Any tubes that have acidified or become cloudy in the overnight incubation are probably contaminated and should be discarded.
- The success of this experiment depends on keeping the cells cold to prevent further internalization of the TLR4 complex. At cold temperatures, the cell membranes change physical properties from fluid to gel (Thewalt and Bloom, 1992; Spink et al., 1996). Protein complexes required for internalization will still form inside the membrane and as soon as membranes liquidize, internalization will proceed. Therefore, please be sure to pre-cool centrifuges and have an ice bucket and ice-cold FACS buffer or media available for rapid-cooling of cells at the time they are removed from the incubator. Minimize the time that the lid of the centrifuge is open, avoid touching the bottom of the flow cytometry tubes, and return tubes to the ice bucket immediately when removing from centrifuge. Without touching the bottom, you can hold up the tubes against a light to visually inspect the tubes after their centrifugation: Pellets of cold cells have higher opacity. Experiments can also be performed in a cold-room.
- This assay is compatible with surface staining of other cell markers. If intracellular staining is required, we recommend setting up a parallel set of samples for permeabilization.
Recipes
- Culture Media
- RPMI 1640 + 2% FBS + 2 mM L-glutamine + 1% Pen/Strep
- Filter sterilize through 0.22 μm filter
- Store at 4 °C for up to three weeks
- FACS buffer
- Ice-cold PBS + 0.5% FBS + 2 mM EDTA
- Store at 4 °C for up to three weeks
Acknowledgments
Supported by the US National Institutes of Health (AI123371 and AI125215 to S.N.V.). This protocol was adapted or modified from previous work (Kagan et al., 2008; Zanoni et al., 2011; Rajaiah et al., 2015).
Competing interests
We declare there are no competing interests related to this work.
References
- Kagan, J. C., Su, T., Horng, T., Chow, A., Akira, S. and Medzhitov, R. (2008). TRAM couples endocytosis of Toll-like receptor 4 to the induction of interferon-β. Nat Immunol 9(4): 361-368.
- McIntire, F. C., Sievert, H. W., Barlow, G. H., Finley, R. A. and Lee, A. Y. (1967). Chemical, physical, biological properties of a lipopolysaccharide from Escherichia coli K-235. Biochemistry 6(8): 2363-2372.
- Perkins, D. J., Richard, K., Hansen, A. M., Lai, W., Nallar, S., Koller, B. and Vogel, S. N. (2018). Autocrine-paracrine prostaglandin E2 signaling restricts TLR4 internalization and TRIF signaling. Nat Immunol 19(12): 1309-1318.
- Rajaiah, R., Perkins, D. J., Ireland, D. D. and Vogel, S. N. (2015). CD14 dependence of TLR4 endocytosis and TRIF signaling displays ligand specificity and is dissociable in endotoxin tolerance. Proc Natl Acad Sci U S A 112(27): 8391-8396.
- Spink, C. H., Manley, S. and Breed, M. (1996). Thermodynamics of transfer of cholesterol from gel to fluid phases of phospholipid bilayers. Biochim Biophys Acta 1279(2): 190-196.
- Takeda, K. and Akira, S. (2004). TLR signaling pathways. Semin Immunol 16(1): 3-9.
- Thewalt, J. L. and Bloom, M. (1992). Phosphatidylcholine: cholesterol phase diagrams. Biophys J 63(4): 1176-1181.
- Zanoni, I., Ostuni, R., Marek, L. R., Barresi, S., Barbalat, R., Barton, G. M., Granucci, F. and Kagan, J. C. (2011). CD14 controls the LPS-induced endocytosis of Toll-like receptor 4. Cell 147(4): 868-880.
Article Information
Copyright
© 2019 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Richard, K., Vogel, S. N. and Perkins, D. (2019). Quantitation of TLR4 Internalization in Response to LPS in Thioglycollate Elicited Peritoneal Mouse Macrophages by Flow Cytometry. Bio-protocol 9(18): e3369. DOI: 10.21769/BioProtoc.3369.
Category
Cell Biology > Cell-based analysis > Flow cytometry
Immunology > Immune cell function > Macrophage
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link









