- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Photodynamic Therapy in a 3D Model of Ovarian Cancer
Published: Vol 9, Iss 15, Aug 5, 2019 DOI: 10.21769/BioProtoc.3314 Views: 5620
Reviewed by: Khyati Hitesh ShahAnne-Marie Caroline OverstreetAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
Real-time IncuCyte® Assay for the Dynamic Assessment of Live and Dead Cells in 2D Cultures
Arlene K. Gidda [...] Sharon M. Gorski
Feb 5, 2025 2950 Views
NanoPDLIM2-Based Combination Therapy for Lung Cancer Treatment in Mouse Preclinical Studies
Thi Hoa Le [...] Zhaoxia Qu
Sep 5, 2025 2271 Views
Combining Microwave Ablation With CAR-T-Cell Therapy in Tumor-Bearing Mouse Models
Bihui Cao [...] Jia Shen
Oct 20, 2025 2363 Views
Abstract
Photodynamic therapy (PDT), is a clinically-approved light-based anti-cancer treatment modality in which a photoactivatable photosensitizer is irradiated with an appropriate wavelength of light to generate cytotoxic molecules to kill cancer cells. In this article, we describe an in vitro PDT protocol using a 3-dimensional (3D) model of ovarian cancer that was established on beds of Matrigel. PDT was performed using a liposomal formulation of verteporfin photosensitizer (Visudyne®). The cancer cells were genetically-labeled with the fluorescent protein mCherry to facilitate the evaluation of the treatment response. This protocol is advantageous because the mCherry fluorescence is an indicator of cell viability, eliminating the need for external dyes, which often exhibit limited penetration and diffusion into 3D organoids. Additionally, Visudyne PDT achieves significant tumor-killing efficacy in a 3D model for ovarian cancer.
Keywords: Photodynamic therapyBackground
Photodynamic therapy (PDT) is an FDA-approved light-based treatment modality that utilizes a photoactivatable molecule, termed a photosensitizer (PS). Upon irradiation with an appropriate wavelength of light, the PS generates reactive molecular species (RMS) within the cytosol. These RMS, often reactive oxygen species, induce photodamage and are toxic to the cell (Celli et al., 2010; Obaid et al., 2016). Certain formulations of PS preferentially accumulate in the malignant tissue and thus PDT often achieves dual selectivity as a result of confined PS accumulation in the tumor and its spatiotemporal control over the light irradiation. PDT is clinically approved for several indications, including esophageal cancer, lung cancer, age-related macular degeneration, and is being investigated for many more cancerous and noncancerous indications (Celli et al., 2010).
Ovarian cancer is a deadly gynecologic malignancy often associated with treatment-resistant metastases and poor outcomes in patients. While surgery and chemotherapy are the current standard of care for ovarian cancer, the efficacy of PDT to manage ovarian cancer is currently being investigated in pre-clinical studies by many research groups (Goff et al., 1996; Rizvi et al., 2010).
Like many other malignancies, ovarian cancer metastases display complex morphological and biochemical characteristics that are lost in traditional monolayer (2D) cell culture. 3D tumor model is an emerging technique for evaluating tumor properties and treatment response in vitro. 3D cultures of tumor cells create a more physiologically relevant tumor microenvironment than traditional 2D cell culture because they retain the biological and structural features of tumors that are found in vivo (Rizvi et al., 2016). Cells in monolayer experience contact inhibition and a homogeneous oxygen and nutrient environment, while 3D cultures display heterogeneity within and across spheroids and restore extracellular matrix-related cues that are important in treatment response. Like in vivo tumors, 3D cultured tumors proliferate into heterogeneous clusters with varying levels of oxygen and nutrients, making them useful tools for studying PS uptake and PDT efficacy within oxygen gradients. Increased PDT resistance has been found in 3D tumor models compared with 2D, providing a platform for testing alternative treatment regimens and combination therapies to overcome this resistance. The complex factors that contribute to treatment failure in patients necessitate an in vitro model that mimics the structural, biological, and extracellular cues that play a major role in treatment outcomes (Rizvi et al., 2016).
Here, we describe an in vitro PDT protocol using a 3D model of ovarian cancer. 3D nodules of human ovarian cancer cells (NIH: Ovcar5) that stably express a fluorescent reporter were grown on beds of Growth Factor-Reduced Matrigel®. PDT was performed using FDA-approved PS verteporfin (Visudyne®). The treatment response was evaluated by imaging of the fluorescent 3D nodules. The protocol described here has been adapted from a previously published article (Rizvi et al., 2019).
Materials and Reagents
- Pipette tips (200 and 1,000 μl, Fisher Scientific, catalog numbers: 02-707-407 and 02-707-413, respectively)
- LASER-safety glasses (Thorlabs, catalog number: LG3)
- Aspirating pipettes (Celltreat, catalog number: 229265)
- 5 and 10 ml serological pipettes (Corning, catalog numbers: 357543 and 357551, respectively)
- 15 ml centrifuge tubes (Corning, catalog number: 352095)
- 50 ml centrifuge tubes (Corning, catalog number: 430829)
- 24-well glass-bottom black-wall plate (Greiner Sensoplates, catalog number: 662892)
- T75 flask (Corning, catalog number: 353136)
- Aluminum foil (Fisher Scientific, 01-213-105)
- mCherry-expressing NIH: OVCAR5 cell line (Rizvi et al., 2019) (“mCherry-Ovcar5” thereafter)
- Visudyne® (Verteporfin for injection by Bausch & Lomb)
- Dulbecco’s Phosphate Buffered Saline (DPBS) without Ca2+ & Mg2+ (Corning, Cellgro, catalog number: 21-031-CV)
- Heat inactivated fetal bovine serum (FBS) (Gibco, catalog number: 26140079)
- 100x Penicillin-streptomycin solution (Corning, catalog number: 30001Cl)
- Trypsin-EDTA solution (Corning Cellgro, catalog number: 25-052-Cl)
- Corning Growth factor-reduced (GFR) Matrigel® Matrix (“Matrigel” thereafter) (Corning, catalog number: 354230)
- Matrigel matrix using Cell Recovery Solution (Corning, catalog number: 354253)
- RPMI 1640 medium (Corning, Cellgro, catalog number: 10-040-CV)
- DMSO (Sigma, catalog number: D2438)
- RPMI Growth Medium (see Recipes)
- 4% Matrigel (see Recipes)
- Visudyne® working solution (see Recipes)
Equipment
- Pipettes (P1000 and P200, Gilson, catalog numbers: F123602 and F123601, respectively)
- Biosafety cabinet for cell culture (Baker Company, catalog number: SG-200)
- Pipette-aid (Biotang, catalog number: 710932)
- Analytical balance (Sartorius, catalog number: ENTRIS2241SUS)
- Inverted light microscope (VWR, catalog number: 89404-462)
- 5x air objective (NA 0.16) lens (PerkinElmer, catalog number: HH14000402)
- Freezer (-20 °C) (Thermo Scientific, model: 20LFEETSA)
- LASER Source (690 nm) (Intense Ltd., model: Intense series 7400)
- Laser power meter (Ophir Vega, catalog number: 7Z01560)
- Laser shutter controller (INLINE-TTL Electronic Shutter)
- Centrifuge machine (Beckman, model: Allegra 6R)
- High-Content Imaging Instrument (Operetta CLS, PerkinElmer)
- Humidified cell incubator (Thermo Scientific, model: FormaTM 310)
- Cell counter (Beckman Coulter, model: Z1 Particle Counter)
Software
- Harmony 4.6 (PerkinElmer)
- GraphPad Prism 8
- (Optional) ImageJ or qVista software
Procedure
Note: Perform all the cell culture-related work inside a sterile biosafety cabinet following standard aseptic technique.
- Preparation of Matrigel Plate
- Thaw a vial of Matrigel overnight by submerging the vial in ice in a walk-in cold room.
- Place a box of sterile pipette tips (1,000 µl) and a glass-bottom black-wall 24-well plate on ice or ice pack for at least 15 min before handling Matrigel.
- Using a P1000 pipette, gently dispense 250 µl of Matrigel into each well without introducing any bubbles while keeping the plate on a flat ice pack.
- Incubate the plate at 37 °C in a cell culture incubator for 20-30 min to polymerize Matrigel.
- Add 500 µl/well of RPMI growth medium (see Recipes) containing 4% Matrigel to avoid further drying of the Matrigel beds.
- Keep the plate in the cell culture incubator until the mCherry-Ovcar5 cells are ready in Procedure B.
- Cell seeding and 3D culture
- Maintain a 2D culture of mCherry-Ovcar5 cells in a T75 flask with RPMI growth medium in a humidified cell culture incubator at 37 °C with 5% CO2.
- Check the confluency of mCherry-Ovcar5 cells using an inverted light microscope. Use a plate that has 50%-80% confluency.
- Replace the existing culture medium from the flask with 10 ml DPBS and incubate for 10 min.
- Discard DPBS and add 2-3 ml of Trypsin-EDTA solution to the flask.
- Incubate for 3-5 min at 37 °C.
- Observe the plate under a light microscope to check if the cells come off the surface completely.
- Quench trypsin by adding 10 ml RMPI growth medium and resuspend the cells.
- Transfer the cell suspension into a 15 ml tube.
- Centrifuge the tube at 450 x g for 5 min at room temperature.
- Discard the supernatant and resuspend the cell pellet with 10 ml RPMI growth medium.
- Determine the cell concentration using Beckman Coulter Counter by setting the cell size between 8-20 µm.
- Prepare 13 ml cell suspension in RPMI growth medium to achieve a concentration of 10,000 cells/500 µl.
- Add 500 µl of the cell suspension into each well of the 24-well plate resulting in the final concentration of 10,000 cells in 1 ml medium with 2% Matrigel.
- Incubate the plate for 7 days to allow 3D tumor nodule formation and replace the medium with fresh RPMI growth medium with 2% Matrigel on Day 4.
- In vitro Photodynamic therapy
- On Day 7, observe the plate under an inverted light microscope to ensure the formation of the tumor nodules on the Matrigel beds (see Figure 1A).
- Prepare 13 ml of Visudyne working solution (see Recipes) by diluting Visudyne stock in RPMI growth medium to achieve 1 µM PS concentration.
- Protect the PS solution in the dark and perform the following steps in the dark or low-light environment.
- Set a very slow aspiration speed and connect a P200 pipette tip (without cotton/filter) to an aspiration pipette and gently aspirate the existing medium from each well without disrupting the 3D nodules.
- Add 500 µl of Visudyne working solution into each well except ‘no treatment’ and ‘light-only’ control wells. Add 500 µl of RPMI growth medium into each of the control wells.
- Incubate for 90 min at 37 °C in a cell culture incubator.
- Replace Visudyne-containing medium with fresh RPMI growth medium.
- During the PS incubation, prepare the Laser source and controller by adjusting light density (irradiance) to 50 mW/cm2 using a Laser power meter.
- To perform PDT, sequentially irradiate each well with 690 nm light directly administered vertically through the bottom of the plate at different doses of light (for example 0-10 J/cm2) by varying the exposure time using a custom-made programmable TTL shutter.
Note: Do not apply any Laser in the ‘no treatment’ wells whereas apply the highest dose of Laser into the ‘light-only’ control wells. - Incubate the plate for additional 4 days following the light irradiation.
- Imaging
- On Day 11, image the plate to quantify residual viable tumor by analyzing mCherry fluorescence (see Figure 1B).
- Place the plate on Operetta CLS, a high-content imaging instrument with confocal capability while maintaining the temperature at 37 °C with 5% CO2.
- Use the excitation and emission filters of 530-560 nm and 570-650 nm, respectively to image mCherry-expressing cells.
- Set image acquisition parameters in Z-stack mode by keeping the visually best fit focal plane in the middle of 10-14 z-planes with 50 µm step size.
- Adjust the LED power and exposure time based on the “No Treatment” control groups so that the camera sensor is not saturated leaving a wide dynamic range for capturing the highest and lowest.
- Acquire images using a 5x air objective (NA 0.16) lens in a 2-by-2 mosaic-format (512 x 512 pixels each with 10% overlap).
Data analysis
- Use Harmony 4.6 (PerkinElmer) software that comes with Operetta CLS for image analysis.
- Stitch the image-mosaics from the same plane first and then generate 2-dimensional “maximum intensity projection” (MIP) to get a global image for each well.
- Set a threshold for mCherry fluorescence based on the “no treatment control” to determine the “live tumor area.”
- Export the values for residual tumor area as a cvs. file and prepare presentable graphs using other software (such as Microsoft® Excel or GraphPad Prism) (see Figure 1C)
Representative Results
Representative outcomes from a PDT treatment of 3D ovarian tumor nodules are shown in Figure 1.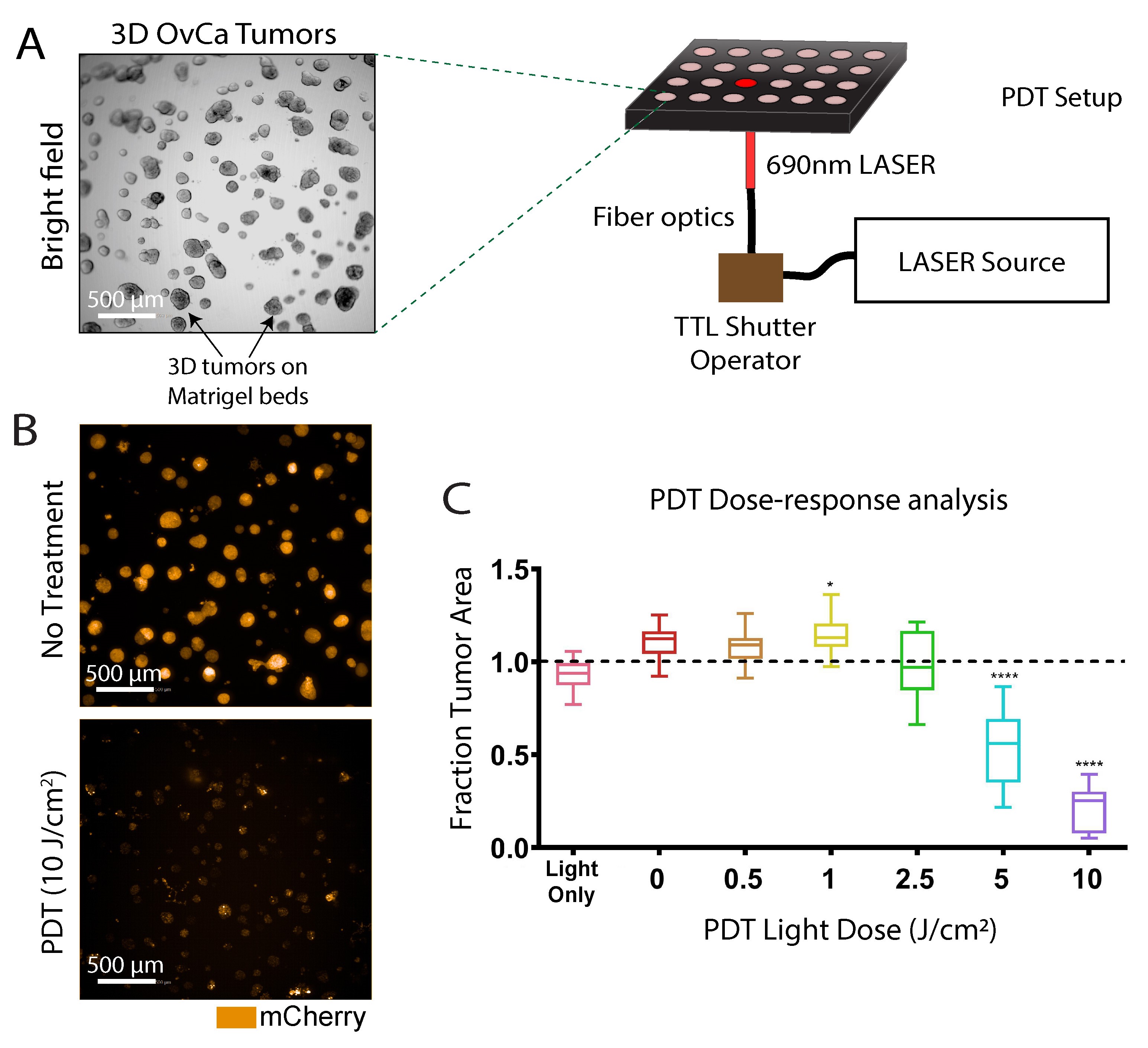
Figure 1. Photodynamic therapy of 3D ovarian cancer nodules. A. A schematic representation of how PDT is performed on the 3D ovarian cancer nodule grown on beds of Matrigel. Scale bar: 500 µm. B. Representative fluorescence images of 3D tumor nodules following 10 J/cm2 PDT treatment are shown along with ‘no treatment’ control. Images were taken 4 days after PDT treatment. Scale bars: 500 µm. C. Quantification of mCherry-expressing tumor area was performed using a high-content imaging system and presented as ‘fraction tumor area’ (residual live tumor area) in the graph after normalizing to ‘no treatment’ control. Images were adapted and reproduced from a published article (Rizvi et al., 2019) with written permission from the publisher.
Notes
- Always use LASER safety glasses to protect eyes during PDT irradiation.
- PDT can be performed outside of a biosafety cabinet while keeping the cellular content sterile in a closed-lid plate.
- PSs are generally light-sensitive. Thus, PS incubation should be done in the dark. The Multi-well plate can be wrapped in aluminum foil during the incubation process to avoid accidental light exposure.
- The treatment response analysis can also be determined if non-fluorescent cells are used. In that case, conventional viability dyes, such as Calcein-AM and propidium iodide can be used as described previously (Anbil et al., 2013).
- Image analysis can alternatively be done using ImageJ or qVista software (Celli et al., 2014).
- There are a plethora of PSs and their liposomal and antibody-conjugated formulations. The PS formulation used in a PDT treatment determines the incubation time, light dose, and the wavelength of light irradiation.
- PDT can also be performed in 2D monolayer culture which may require optimization of the light dose. In general, the LD50 for 2D culture is lower compared with 3D culture.
- PDT-treated cells can be harvested for downstream applications (such as qPCR, Flow cytometry, etc.) by depolymerizing Matrigel matrix using Cell Recovery Solution (Corning, 354253) following the manufacturer’s protocol.
Recipes
- RPMI Growth Medium
- Add 10% (v/v) FBS, 100 IU/ml Penicillin and 100 µg/ml Streptomycin into 500 ml of RPMI 1640 medium
- Pre-heat the growth medium at 37 °C for all the procedures unless otherwise mentioned specifically
- 4% Matrigel
Add 400 μl Matrigel into 10 ml RPMI growth medium - Visudyne working solution (1 μM verteporfin)
- First, determine the PS concentration of the Visudyne stock using the following method and then prepare Visudyne working solution by diluting the stock in RPMI growth medium to achieve 1 μM PS concentration: Set ‘blank’ of your UV-Vis instrument with DMSO by scanning from 200 nm to 800 nm
- Dilute your Visudyne in an appropriate amount of DMSO that yields an absorbance value at 687nm between 0.1 and 1
- Measure the absorbance of the diluted Visudyne from 200 nm-800 nm
- Calculate your Visudyne concentration using the following equation:
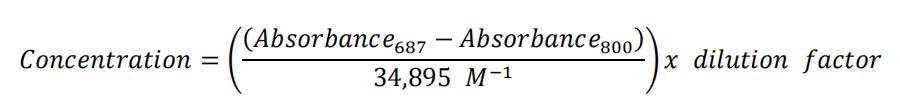
Acknowledgments
This protocol was first reported in a previous publication (Rizvi et al., 2019). We would like to thank the funding sources and the authors of the original publication. The figures were reproduced with written permission from the publisher. The authors would like to thank Saad Mohammad and Joseph Swain for helpful discussion.
Competing interests
The authors declare no competing financial interests.
References
- Anbil, S., Rizvi, I., Celli, J. P., Alagic, N., Pogue, B. W. and Hasan, T. (2013). Impact of treatment response metrics on photodynamic therapy planning and outcomes in a three-dimensional model of ovarian cancer. J Biomed Opt 18(9): 098004.
- Celli, J. P., Rizvi, I., Blanden, A. R., Massodi, I., Glidden, M. D., Pogue, B. W. and Hasan, T. (2014). An imaging-based platform for high-content, quantitative evaluation of therapeutic response in 3D tumour models. Sci Rep 4: 3751.
- Celli, J. P., Spring, B. Q., Rizvi, I., Evans, C. L., Samkoe, K. S., Verma, S., Pogue, B. W. and Hasan, T. (2010). Imaging and photodynamic therapy: mechanisms, monitoring, and optimization. Chem Rev 110(5): 2795-2838.
- Goff, B. A., Blake, J., Bamberg, M. P. and Hasan, T. (1996). Treatment of ovarian cancer with photodynamic therapy and immunoconjugates in a murine ovarian cancer model. Br J Cancer 74(8): 1194-1198.
- Obaid, G., Broekgaarden, M., Bulin, A. L., Huang, H. C., Kuriakose, J., Liu, J. and Hasan, T. (2016). Photonanomedicine: a convergence of photodynamic therapy and nanotechnology. Nanoscale 8: 12471-12503.
- Rizvi, I., Bulin, A. L., Briars, E., Anbil, S. and Hasan, T. (2016). Chapter 11: Mind the gap: 3D models in photodynamic therapy. In: Photodynamic Medicine. pp. 197-221.
- Rizvi, I., Celli, J. P., Evans, C. L., Abu-Yousif, A. O., Muzikansky, A., Pogue, B. W., Finkelstein, D. and Hasan, T. (2010). Synergistic enhancement of carboplatin efficacy with photodynamic therapy in a three-dimensional model for micrometastatic ovarian cancer. Cancer Res 70(22): 9319-9328.
- Rizvi, I., Nath, S., Obaid, G., Ruhi, M. K., Moore, K., Bano, S., Kessel, D. and Hasan, T. (2019). A Combination of visudyne and a lipid-anchored liposomal formulation of benzoporphyrin derivative enhances photodynamic therapy efficacy in a 3D model for ovarian cancer. Photochem Photobiol 95(1): 419-429.
Article Information
Copyright
© 2019 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Nath, S. and Moore, K. (2019). Photodynamic Therapy in a 3D Model of Ovarian Cancer. Bio-protocol 9(15): e3314. DOI: 10.21769/BioProtoc.3314.
Category
Cancer Biology > General technique > Cancer therapy
Cell Biology > Cell viability > Cell death
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link











