- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
A High-throughput qPCR-based Method to Genotype the SOD1G93A Mouse Model for Relative Copy Number
Published: Vol 9, Iss 12, Jun 20, 2019 DOI: 10.21769/BioProtoc.3276 Views: 7307
Reviewed by: Gal HaimovichKanika GeraAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
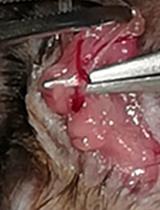
Operant Self-medication for Assessment of Spontaneous Pain Relief and Drug Abuse Liability in Mouse Models of Chronic Pain
David Cabañero [...] Rafael Maldonado
Mar 5, 2022 3051 Views
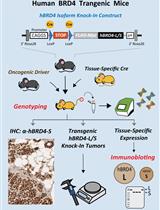
Conditional Human BRD4 Knock-In Transgenic Mouse Genotyping and Protein Isoform Detection
Michael Paul Lewis [...] Cheng-Ming Chiang
Apr 5, 2022 3624 Views
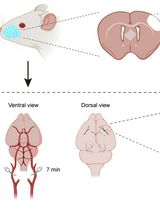
Developing a Ministroke Model in Mouse Barrel Cortex
Song Wang [...] Yihan Wu
Mar 20, 2025 2345 Views
Abstract
The most commonly used mouse model in ALS preclinical research expresses multiple copies of the human SOD1 (G93A) transgene. During the course of breeding, successive generations of mice can lose copies of the transgene. Because shorter lifespan of these mice is dependent on transgene copy number, it is essential to ensure that no low-copy, and therefore longer-lived, mice are included in preclinical studies. Existing techniques for SOD1G93A mouse genotyping are broadly based on creating a standard curve using a reference gene and deducing the relative amount of SOD1 by comparison with the standard curve. This type of technique is used in Alexander et al. (2004), Vieira et al. (2017) and Maier et al. (2018). However, it is not described in detail (see Note 1). This paper provides a detailed protocol for determining the relative copy number of the human SOD1 transgene. Briefly, the protocol involves first the extraction of high-quality genomic DNA from mouse ear tissue, creation of a genomic DNA concentration-based standard curve, and qPCR analysis of up to 88 samples at once alongside the standard curve with Gapdh as a reference gene. Analysis involves the normalization of each unknown sample using the standard curve followed by determination of the copy number of the sample relative to the cohort median. This protocol has been optimized to produce high-quality genomic DNA and consistent results, and the relative copy number cutoffs have been optimized and validated empirically by comparison of relative copy number and mouse lifespan.
Keywords: ALS mouse modelBackground
The SOD1G93A mouse model (B6SJL-Tg(SOD1*G93A)1Gur/J) is currently the most commonly used mouse model for preclinical testing of therapies for amyotrophic lateral sclerosis (ALS), a progressive neurodegenerative disease with most patients living between two and five years after diagnosis (Morrice et al., 2018). The human SOD1 transgene is present in multiple copies and mouse lifespan inversely correlates with copy number. Alexander et al. (2004) found that mice with 24 copies of the SOD1(G93A) gene lived to approximately 129 days, whereas mice with 34 copies lived to 99 days and mice with 20 copies lived to 150 days (Alexander et al., 2004). Additionally, 3.6% of high-copy transgenic male breeders had offspring with low copy numbers over the course of four years, signifying that it is not enough to only test stud males for copy number (Alexander et al., 2004). Although higher copy does correlate with shorter lifespan, spontaneous recombination tends toward dropped copies rather than increased copy number (Alexander et al., 2004; Zwiegers et al., 2014). Therefore, we focused our assay on identifying low-copy mice only.
In order to be sure that any measurable effects in a preclinical drug study are attributable to the treatment and not to an inherent difference in transgene copy, it is absolutely essential that no low-copy mice are included. While there are no SOD1 (G93A) genotyping methods published in a dedicated method or protocol journal, there are papers which briefly describe genotyping in the methods section. For example, Alexander et al. (2004) describe a qPCR-based method that uses change in cycle threshold (∆Ct) of SOD1 (G93A) and a reference gene to quantify copy number difference (Alexander et al., 2004).
In our hands, we found that published genotyping methods prior to 2016 were not sensitive enough to exclude all low-copy animals. Figure 1 shows the age at death of all mice included in a non-treatment group of survival studies from August 2015 to January 2019. The x-axis is the start date of each study, and the data set for each start date is the death ages of the mice included in the untreated group of that study. The finalized genotyping protocol detailed in this paper was instated on May 9, 2016, and the first mouse genotyped with this protocol was included in the study with the start date of June 7, 2016. The graph was analyzed in GraphPad Prism using a Box-and-Whiskers Tukey Test, which highlights any value that is either greater than the 75th percentile plus 1.5 times the interquartile range (IQR) or less than the 25th percentile minus 1.5 times the IQR. These outliers are shown as large blue dots. Since the genotyping protocol was instated, the number of outliers and their distance away from the median tend to decrease.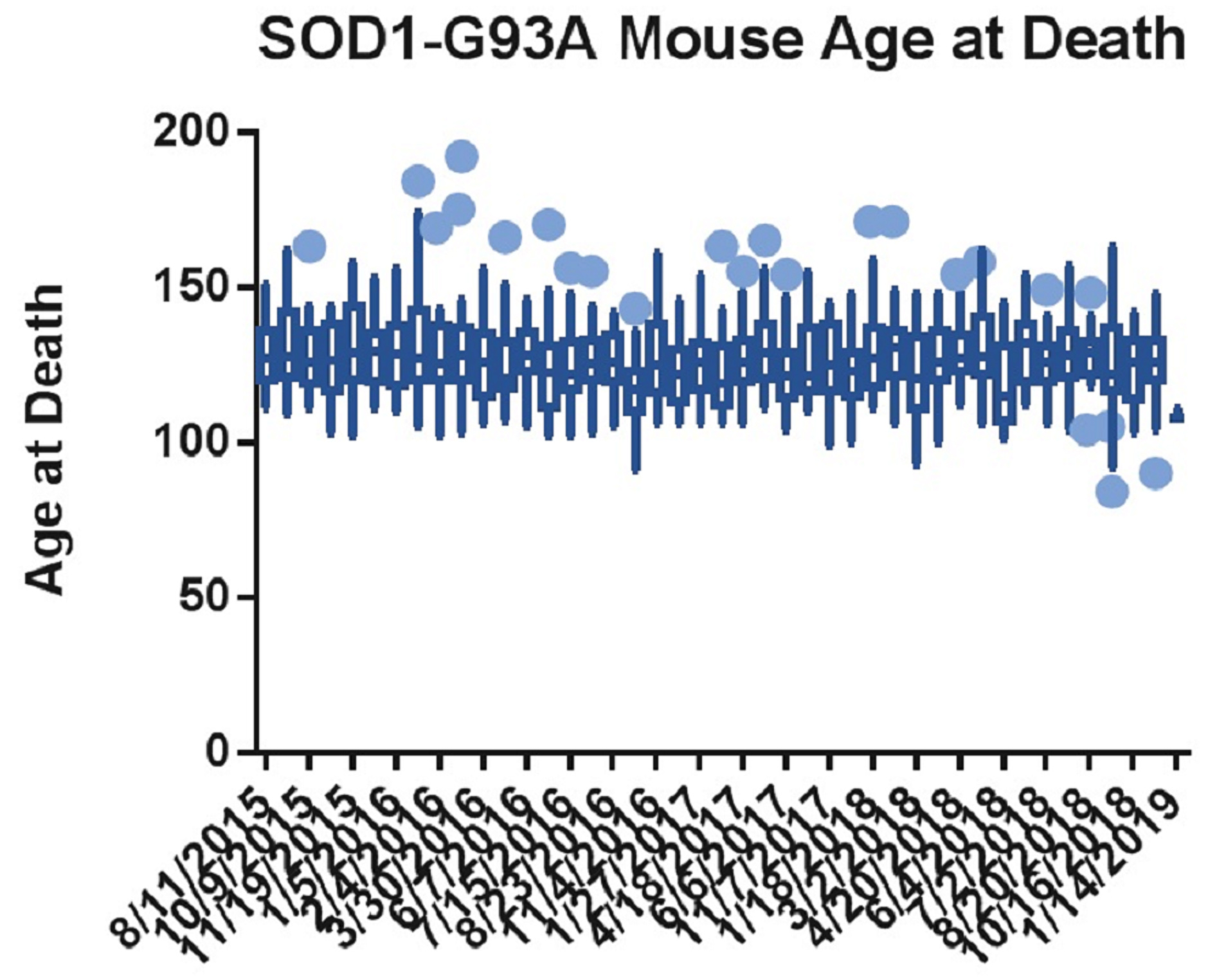
Figure 1. GraphPad Prism analysis of mouse death ages from August 2015 to January 2019. These mice were in untreated groups of survival studies. The dates shown on the x-axis indicate the start date of each study, and the data set associated with each date represents death ages of the mice in the untreated group of each study. Tukey’s Test on box-and-whisker plots has been applied, with the blue dots showing which data points are outside the 75th percentile plus 1.5 times the interquartile range (IQR) or the 25th percentile minus 1.5 times the IQR.
This genotyping protocol has been used in several published studies since 2016, including Vieira et al. (2017), and provides a qPCR-based method to ascertain relative copy number in a cohort of up to 88 mouse ear tissue samples without the need to determine absolute gene copy. The protocol relies on a concentration-based Gapdh standard curve to normalize the samples. After normalization, relative copy numbers are assigned based on log difference between each sample and the cohort median. Along with developing a reliable method to screen for low-copy mice, we optimized tissue collection to avoid cross-contamination and optimized genomic DNA extraction for consistent yield and high quality (see Note 2). Additionally, we also performed a dilution qPCR of potential reference gene primer and probe sets to determine which correlated best with the custom SOD1 target primer/probe set, and to determine the concentration range of our standard curve (Figure 2). As shown with the green triangles, the copy-number Gapdh primer/probe set from Thermo Fisher had the most comparable slope and Ct values to the SOD1 primer/probe sets, and both performed best between 2.5 and 0.07 ng/μl genomic DNA. To determine the safest relative copy number cutoff, we performed a survival study correlating copy number with lifespan. Mice with 16 or more copies showed typical lifespans for this mouse model, while mice with 15 or fewer copies were long-lived (Figure 3). Using this information we determined the safest cutoff to be 15 copies and fewer.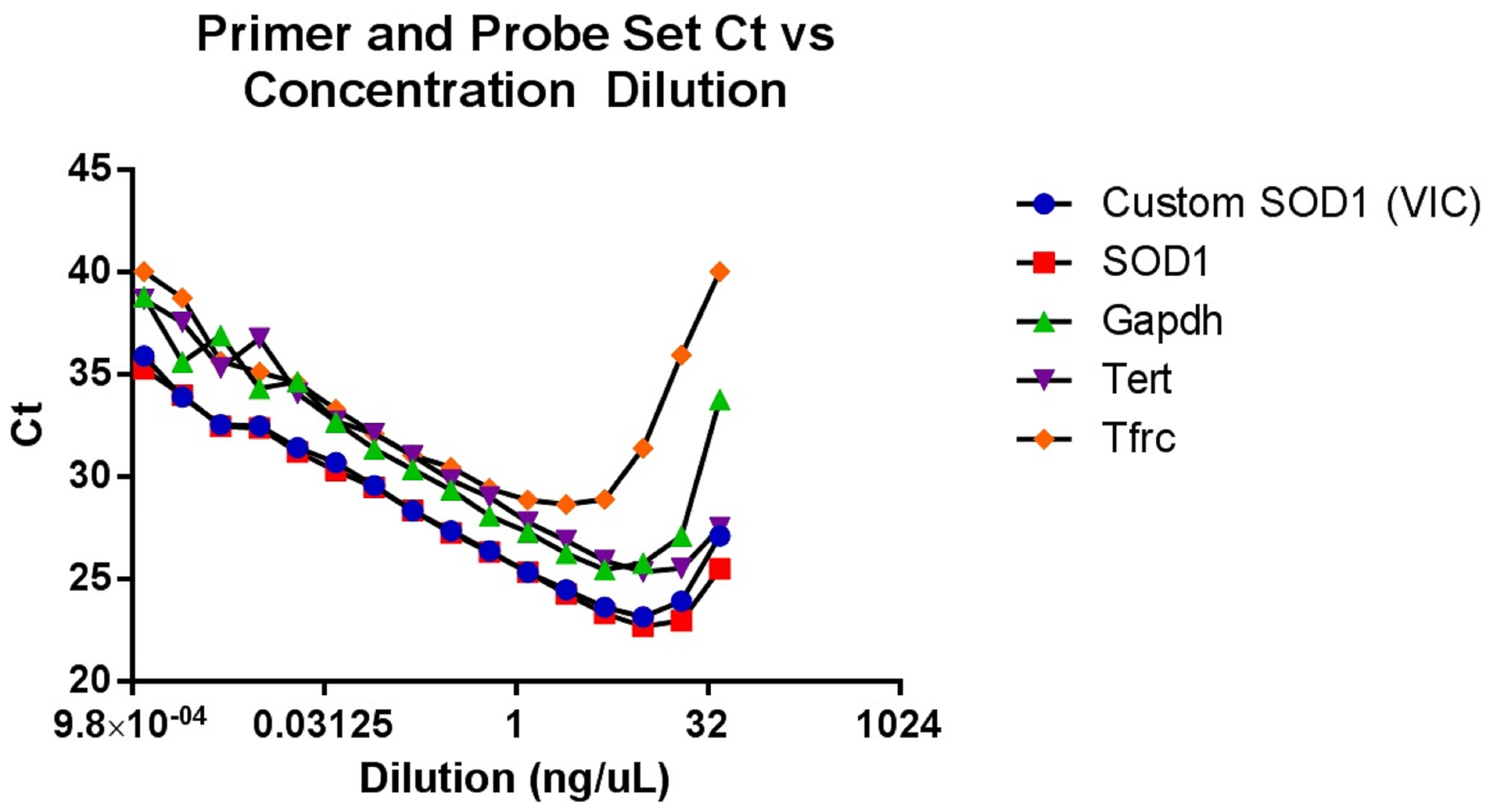
Figure 2. A qPCR experiment performed with serial two-fold dilutions of SOD1G93A genomic DNA from mouse ear tissue, probed with our custom SOD1 primer/probe set as well as several off-the-shelf primer/probe assays from Thermo Fisher. Our custom SOD1 primer/probe set compares well with the off-the-shelf SOD1 assay, and the Gapdh reference assay is closest to the SOD1 target assays in both Ct and in slope of the linear region. The graph is most linear between 0.07 and 2.5 ng/μl of genomic DNA, which gave us the optimal range for our experimental standard curve.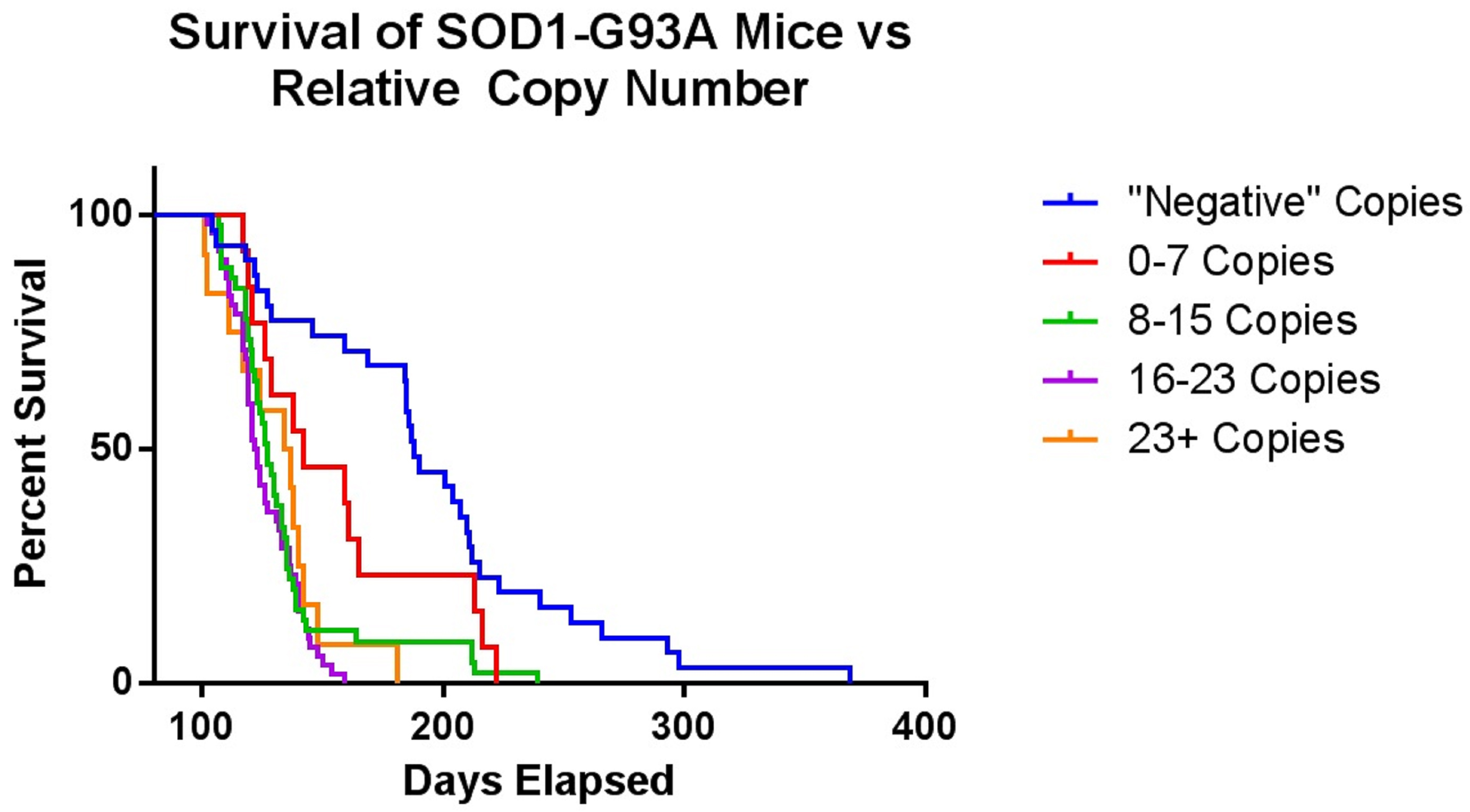
Figure 3. This graph shows the survival of SOD1G93A mice compared to their relative copy number as assigned by the genotyping protocol. The mice within a tolerable range of death ages were those in the 16-23+ copy number groups. Therefore, we selected 15 as the cutoff for relative copy number.
Materials and Reagents
Note: All storage is at room temperature unless otherwise specified.
- QIAcube HT Plasticware (QIAGEN, catalog number: 950067)
- Adhesive PCR Plate Seals (Thermo Fisher, Thermo Scientific, catalog number: AB0558)
- Sterile Alcohol Prep Pads (Fisher Scientific, Fisherbrand, catalog number: 22-363-750)
- 15 ml Centrifuge Tube (Corning, catalog number: 430766)
- 96-well UV-Transparent Microplates (Corning, catalog number: 3635)
- 384-well qPCR plate (Thermo Fisher, Applied BioSystems, catalog number: 4309849)
- Custom SOD1 MGB probe and primers (Thermo Fisher, Custom Genomics, -20 °C)
- Probe: SOD1HumanV2-55T (VIC-ACTCTCTCCAACTTTG) with VIC fluorescent reporter chemistry, 6000 pmole (wet, desalt)
- Forward primer: SOD1HumanV2-27F (GTAAATCAGCTGTTTTCTTTGTTCAGA), 80,000 pmole (dry, desalt)
- Reverse primer: SOD1HumanV2-104R (TTCACTGGTCCATTACTTTCCTTTAA), 80,000 pmole (dry, desalt)
- QIAamp 96 QIAcube HT Kit (QIAGEN, catalog number: 51331)
- PBS, Phosphate Buffered Saline, 10x Solution (Fisher BioReagents, Fisher Scientific, catalog number: BP3994)
- Alcohol, Reagent (Denatured Alcohol), 70% (v/v) Aqueous Solution (Ricca Chemical Company, Fisher Scientific, catalog number: 25467032, room temperature flammables cabinet)
- Neosporin First Aid Antibiotic Ointment (Hanna’s Pharm Supply Co, catalog number: 302-571-8761)
- Cotton-Tipped Applicators (Fisher Scientific, Puritan, catalog number: 22-029-571)
- 50 ml Reagent Reservoir (Corning, Costar, catalog number: 4870)
- DL-Dithiothreitol solution BioUltra for molecular biology, 1 M in H2O (Millipore Sigma, Sigma-Aldrich, catalog number: 43816), 4 °C
- Ethanol, Absolute (200 Proof), Molecular Biology Grade (Fisher Scientific, catalog number: BP2818100), room temperature flammables cabinet
- Nuclease-Free Water (Thermo Fisher, Invitrogen, catalog number: AM9932)
- TaqMan Genotyping Master Mix (Thermo Fisher, Applied Biosystems, catalog number: 4371355), 4 °C
- TaqMan Gapdh Copy Number Assay (Thermo Fisher Assay ID Mm00186822cn), -20 °C
- Sample Prep solution (see Recipes)
- Tissue Digestion solution (see Recipes)
- Primer Dilution solution (see Recipes)
- SOD1G93A Copy Number Genotyping qPCR Master Mix (see Recipes)
- Primer/Probe solution (see Recipes)
Equipment
- Two sterilized vessels for tissue harvest ethanol washes
- Surgery-quality tools to harvest ear tissue (forceps and scissors, or ear punch)
- Shaker capable of 18 h of 200 rpm and 56 °C incubation (Eppendorf, New Brunswick, catalog number: M1352-0010) (see Note 3)
- Qiagen QIAcube HT (QIAGEN, catalog number: 9001793) (see Note 3)
- SpectraMax M5 Multi-Mode Microplate Reader (Molecular Devices, catalog number: M5) (see Note 3)
- NanoDropTM 8000 Spectrophotometer (Thermo Fisher, Thermo Scientific, catalog number: ND-8000-GL) (see Note 3)
- Allegra X-22R Benchtop Centrifuge (Beckman Coulter, catalog number 392188) (see Note 3)
- QuantStudioTM 7 Flex Real-Time PCR System, 384-well, desktop (Thermo Fisher, catalog number: 4485701) (see Note 3)
Software
- Excel 2013 (Microsoft)
- Prism 6 (GraphPad, www.graphpad.com)
Procedure
- Tissue collection
Note: Although this protocol specifies extracting and analyzing genomic DNA from ear samples, 20 mg mouse tail tip samples may also be used.- Preparation
- Create a plate map with samples organized top to bottom in columns (i.e., Sample 1 is in spot A1, Sample 2 is in spot B1, etc.). Remember that in every experiment, 8 sample spots need to be reserved for a standard curve, so the most mice that can be analyzed in a single qPCR is 88 (Figure 4A).
- Create Sample Prep solution (Recipe 1) with appropriate excess (at least 10).
- Using an S-Block from the Qiagen QIAcube Plasticware kit, load 160 μl of the Sample Prep solution into every well of every column needed for the experiment (Figure 4B).
Note: The QIAcube HT works in column format, and every well of the columns that are used (regardless of whether they contain a sample) needs to be filled in order for the automated protocol to work properly. - Seal the top of the S-block with an adhesive plate seal.
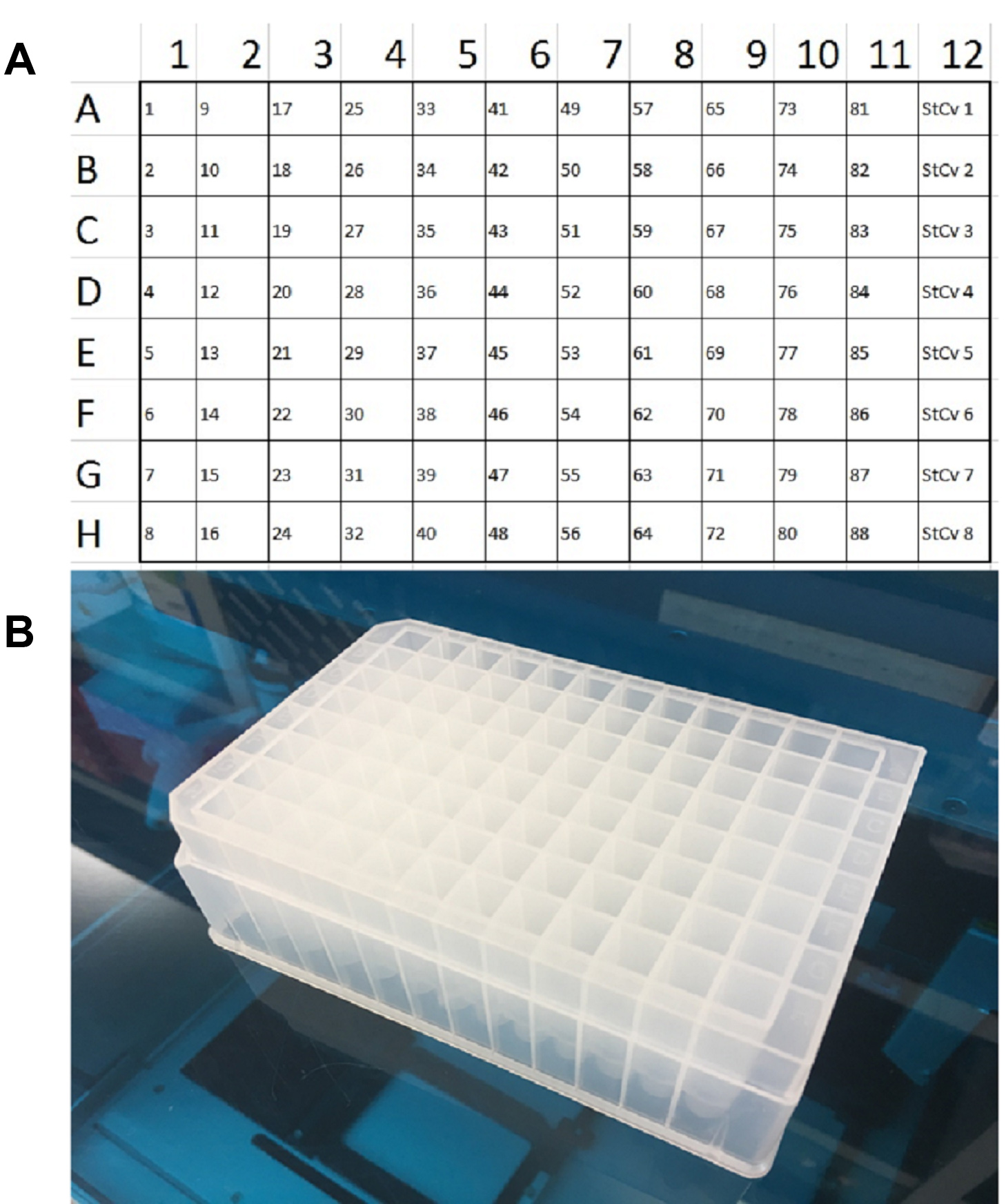
Figure 4. Sample collection plate example. A. Sample collection plate map for an 88-sample experiment, with placeholder wells for the standard curve. B. Qiagen S-Block.
- Sample Collection
- Fill two clean vessels with enough 70% reagent alcohol to wash tools.
- Pre-clean tools with 70% reagent alcohol.
- Collect an ear sample from each mouse that is 20 mg or less using scissors or ear punch (Figure 5); using forceps, place sample into the appropriate well according to the plate map.
- Apply antibiotic ointment to mouse ear using cotton-tipped applicator.
Note: The antibiotic swab does not usually need to be cleaned between uses, but if the mouse bleeds, we replace the soiled swab. - Swish forceps and scissors in first alcohol wash, wipe with alcohol prep pad to remove any solid material such as hairs, and then swish in second alcohol wash. Leave tools in the second wash until ready to use again; do not mix up washes because the first wash should function as a “dirty wash” and the second should function as a “clean wash”.
- Repeat with remaining mice, changing the alcohol prep pad and cleaning gloves often.
- Cover S-block with adhesive when you’re done and during any extended breaks.
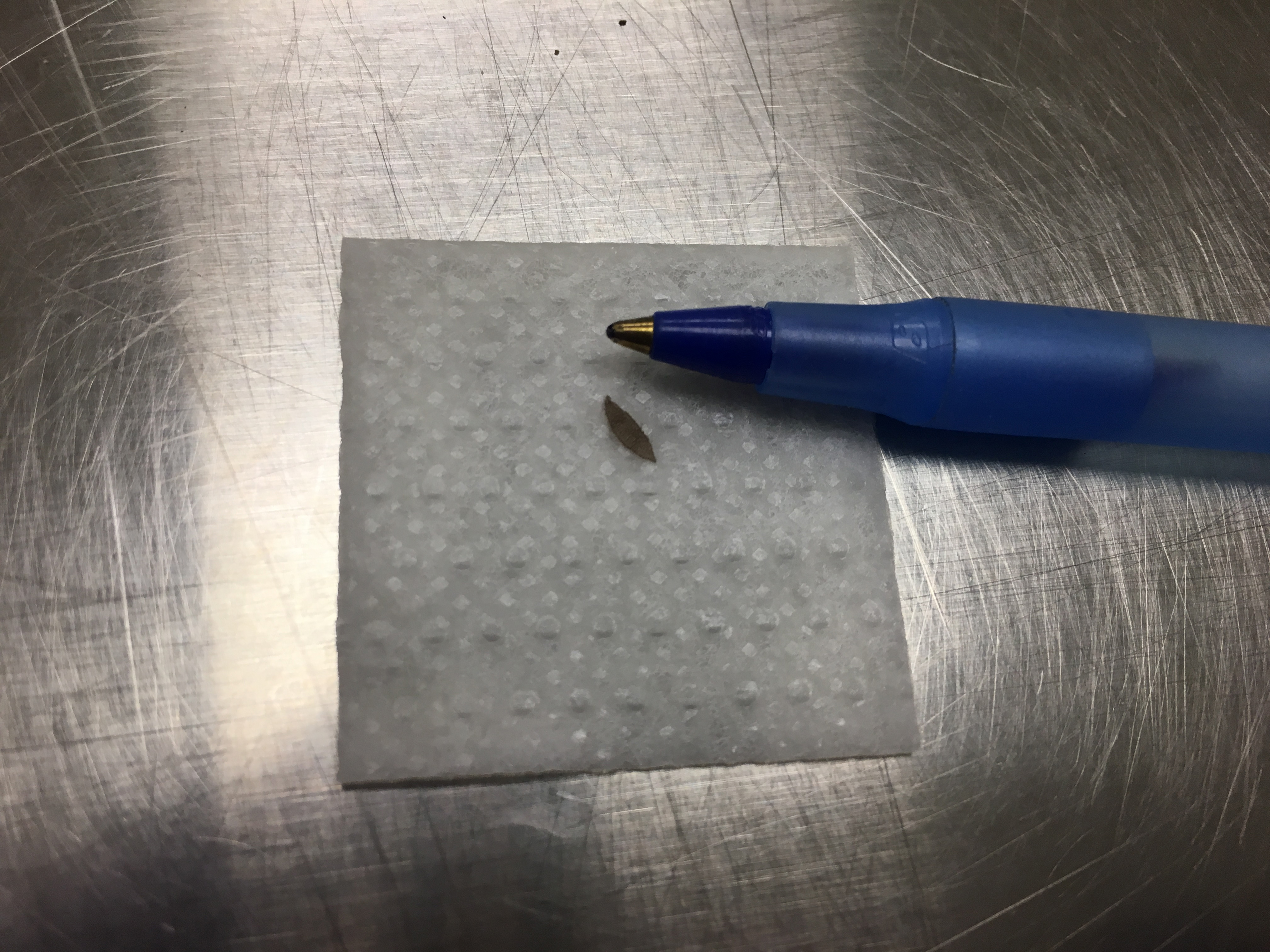

Figure 5. An ear tissue sample that is approximately 20 mg
- Sample digestion
- In reagent reservoir, create Tissue Digestion solution (Recipe 2) with appropriate excess (at least 10), mix well, and aseptically load 40 μl into sample wells in S-block; load recipe into any “blank” wells too (wells in a used column that do not have a sample).
- Ensure that every sample is submerged.
- Seal S-block with a thermally stable adhesive plate cover; add extra security with lab tape so that the adhesive cover does not pop off during overnight digestion (samples will evaporate if this happens).
- Secure in a thermal shaker set to 56 °C; shake at 200 rpm overnight or up to 18 h.
- Preparation
- Genomic DNA extraction
- QIAcube HT setup
- After overnight sample digest, open QIAcube HT’s QIAamp 96 protocol.
- Follow the setup wizard in the QIAamp 96 QIAcube HT protocol to input sample number and make the following customizations.
- Make elution volume 150 μl (Figure 6A).
Note: In our hands, 150 μl elution produces sample yield between 10-50 ng/μl, which we found to be the best range for sample dilution. You may need to change elution volume to optimize yield range. - Remove Top Elute (change volume to 0) (Figure 6B).
Note: With experimentation, we found that using Top Elute is messy and does not improve sample yield or quality.
- Make elution volume 150 μl (Figure 6A).
- Set up deck according to the specifications on the software program.
- Deionize the deck with an ionizing gun or fan prior to running the protocol.
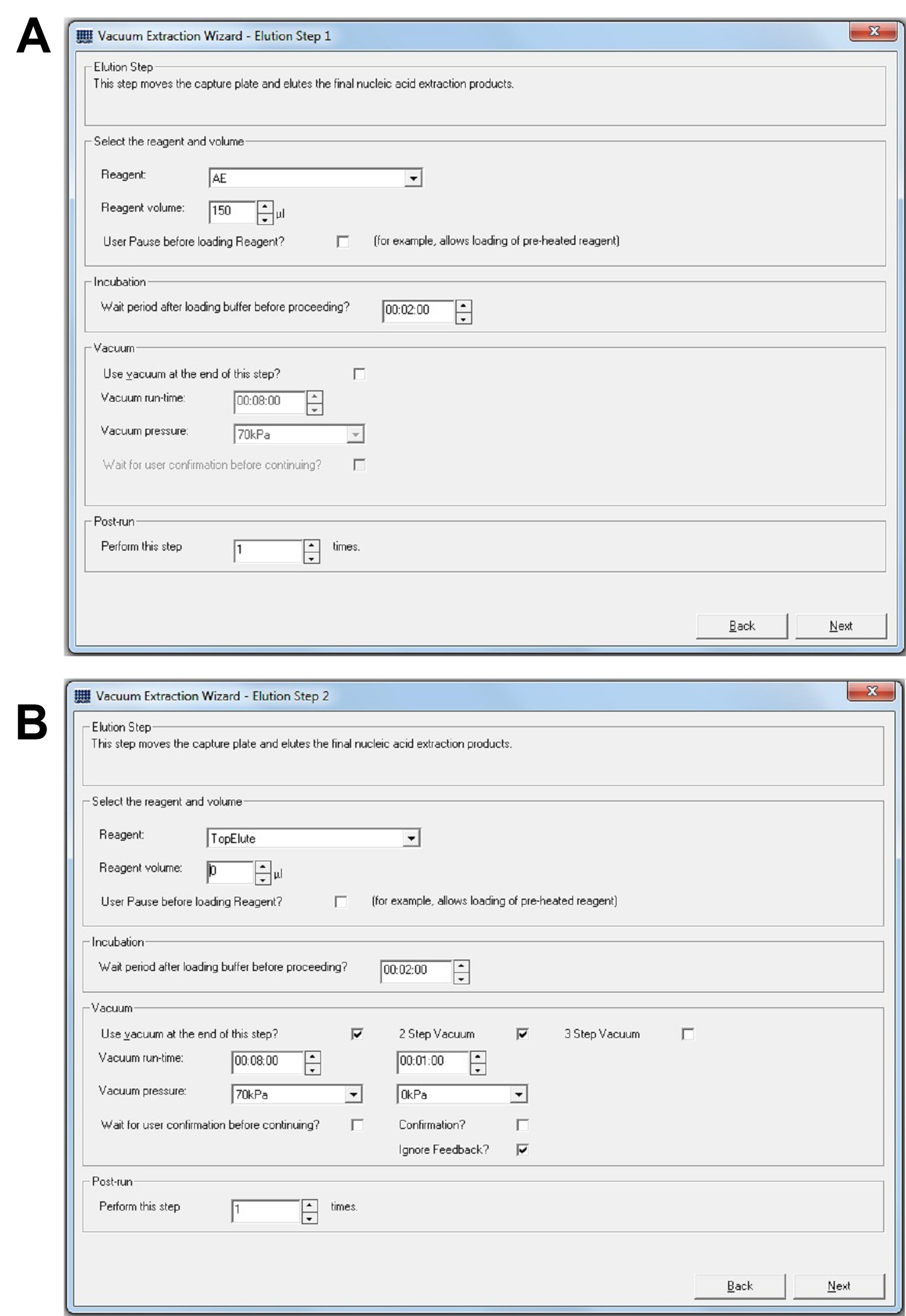
Figure 6. QIAcube setup. A. Set Elution Buffer to 150 μl. B. Set Top Elute volume to 0 μl.
- Storage: once the protocol is finished, cover the elution plate with adhesive tape and lid, label, and store at 4 °C until ready to use.
Note: The kit instructs the user to use the rubber tube caps, but we find that to be cumbersome and have an increased chance of liquid flicking into another well.
- QIAcube HT setup
- Genomic DNA Dilution
- Use a plate reader or MultiDrop to take 260 nm, 280 nm, and 320 nm readings of the DNA (if using a plate reader, dilute gDNA 1:10 in water in a UV plate).
- Quantify the concentration and 260/280 ratio; ensure that 260/280 values are approximately 1.8.
Note: Another ratio that indicates sample purity is 260/230. Ideal 260/230 values are at least 1.0, but we have not found that 260/230 values have a significant effect on qPCR results so it is not further addressed in this protocol. - Find the samples with the highest concentration; choose one dilution factor for the whole plate that ensures that the highest concentration sample does not exceed 1 ng/μl and the lowest concentration sample is not lower than 0.02 ng/μl.
- Dilute all samples in PCR-clean water in a new plate and mix well.
- Standard curve setup
- Combine and mix several samples’ worth of high-quality genomic DNA made from several different SOD1G93A mice to a total of approximate 100 μl of mixed sample.
- Find the concentration (ideally, use a NanoDrop for most accurate results); take a few replicates to make absolutely sure.
- Dilute the sample down to 2 ng/μl in 80 μl volume with PCR-clean water; save remaining mixed gDNA stock at -20 °C for future standard curves.
- Starting with 70 μl of 2 ng/μl gDNA, make 8 serial two-fold dilutions in PCR-clean water in one column of a PCR plate; this creates a standard curve where the most-concentrated sample is 2 ng/μl and the least is 0.016 ng/μl (Figure 7).
Note: You can make more than one standard curve at a time if you use them a lot, and store them at 4 °C. - Cover with adhesive plate sealer and store at 4 °C. Use aseptic technique every time you use the standard curve.
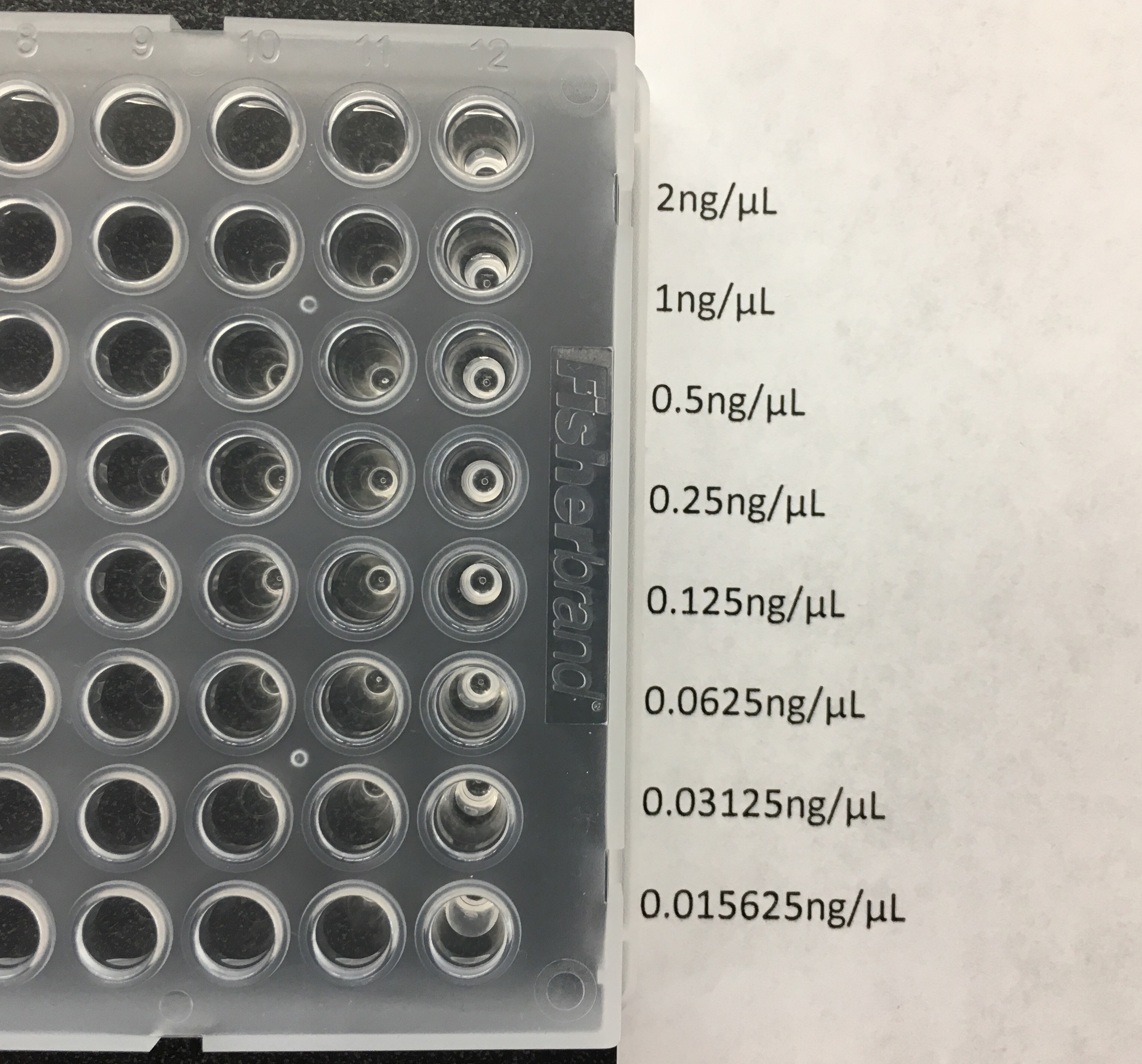
Figure 7. Standard curve setup in a 96-well mixer plate
- qPCR setup
- Primer setup
- Dilute dry primers according to the “Primer Dilution solution” (Recipe 3); leave on the bench for 10 min, vortex, and spin down.
- Aliquot into 100 μl aliquots, label, and store at -20 °C.
- Aliquot wet probes into 20 μl aliquots, label, and store at -20 °C.
- qPCR Master Mix
- Make master mix according to the “SOD1G93A Copy Number Genotyping qPCR Master Mix solution” (Recipe 4), using appropriate excess (be sure to include standard curve in sample number calculations and make enough for two genes, SOD1 and Gapdh).
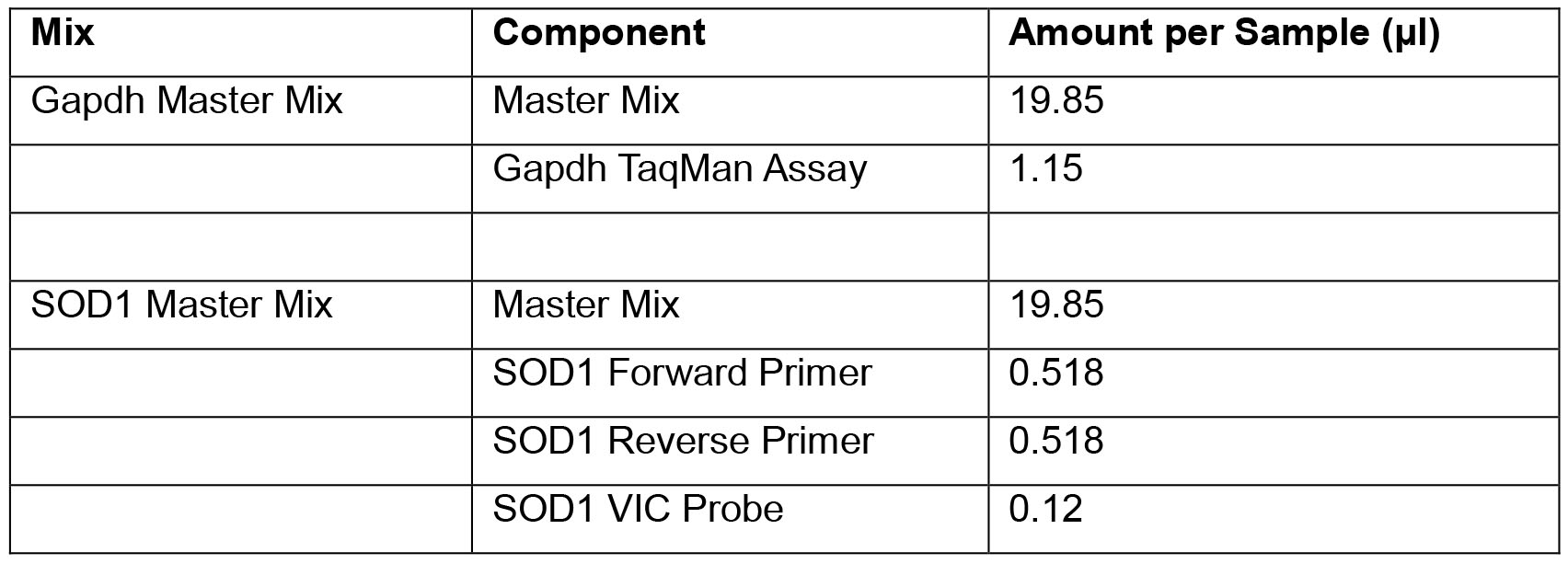
- Create SOD1 and Gapdh mixes according to the “Primer/Probe solution” (Recipe 5), using appropriate excess. Remember when calculating excess that you can use at most half of the number of excess samples that you used for the master mix for the SOD1 and Gapdh mixes. Mix gently; store at 4 °C covered with foil until ready to use (not more than 1-2 h).
- Label 2 96-well PCR plates “SOD1” and “Gapdh”, respectively.
- Aliquot 4 μl of diluted unknown gDNA into each plate.
- Add 4 μl of standard curve after the last column of samples on the Gapdh plate.
- Aliquot 20 μl of SOD1 master mix to each sample well of the SOD1 plate; pipette mix very well but do not introduce bubbles. Repeat with Gapdh plate and master mix.
- Spin plates at 180 x g for 3 min.
- Starting with the SOD1 plate, pipette 9.8 μl duplicates across a 384-well qPCR plate with an 8-channel pipette so that the samples occupy rows A, C, E, G, etc. Repeat with Gapdh directly under the SOD1 rows so that Gapdh samples occupy rows B, D, F, H, etc.; make sure to include standard curve (Figure 8).
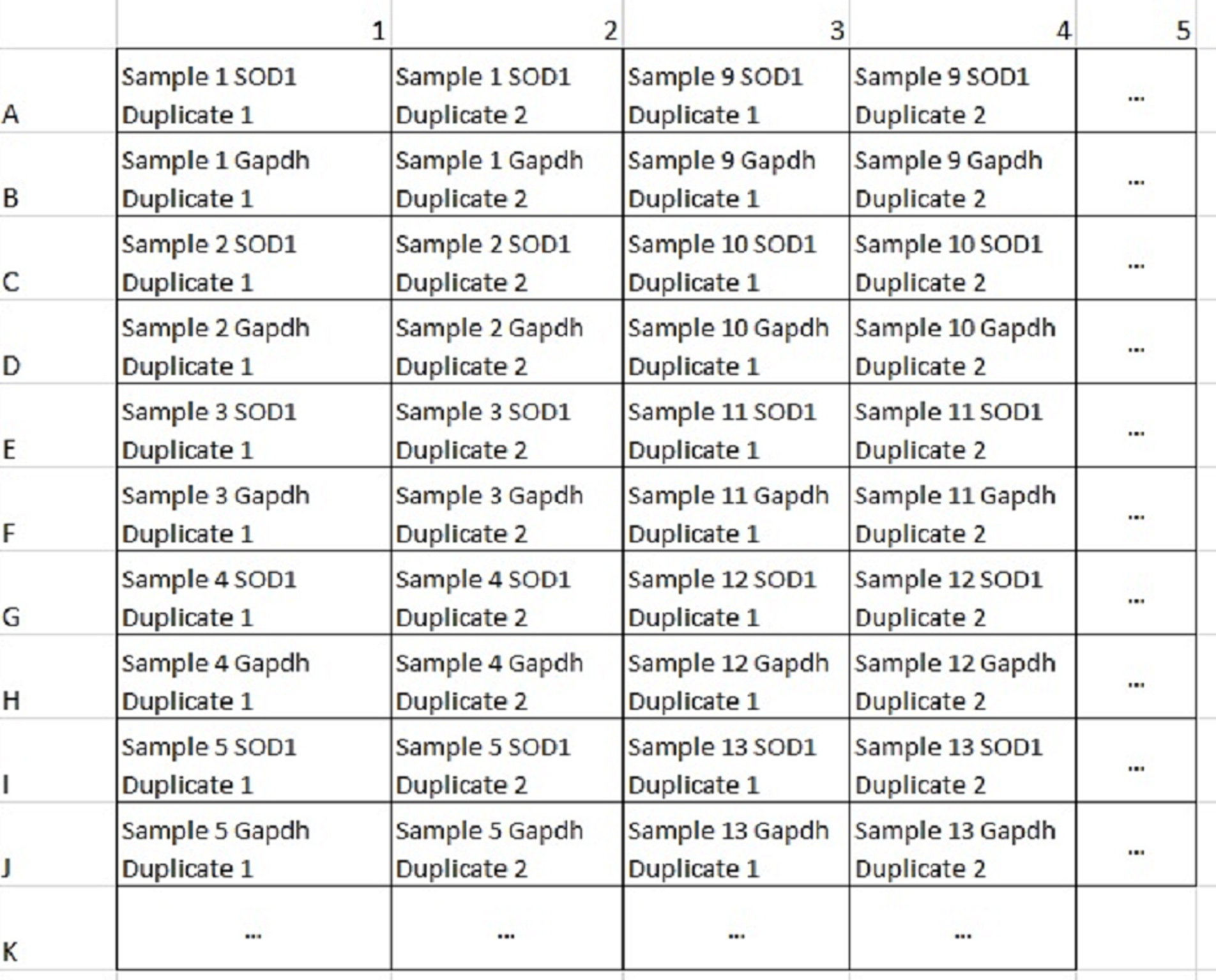
Figure 8. A few example columns of a qPCR plate map in a 384-well plate. Gapdh standard curve samples would be added to the final two columns in duplicate after all samples have been added. - Spin at 400 x g for 3 min.
- Seal the plate with an optical plate sealer and run a standard-length TaqMan qPCR (Figure 9).
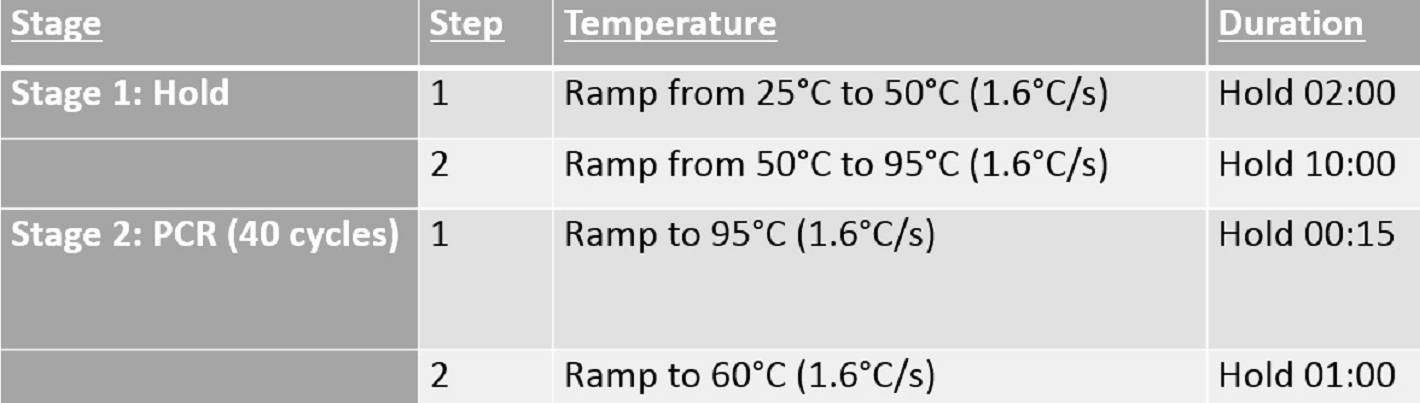
Figure 9. Standard-length genotyping qPCR cycling conditions for qPCR machine setup
- Primer setup
Data analysis
- Organize data
- Export the following data from the qPCR machine into Microsoft Excel: Raw Ct, Sample Name, Target Name.
- Find the average raw Ct of each set of duplicates.
- Find interpolated concentrations of unknowns using standard curve
- Open an XY spreadsheet in GraphPad Prism; choose “Enter X values” and “Enter and plot a single Y value for each point”.
- Label the X column “Concentration, ng/μl” and the first Y column “Gapdh Ct”.
- In the first 8 rows of the “Concentration”, put the concentration values of the standard curve (2, 1, 0.5, 0.25, 0.125, 0.0625, 0.03125, 0.015625).
- In the first 8 rows of the “Gapdh Ct” column, paste the raw average Ct values for your Gapdh standard curve.
- Click “Analyze” and choose “Nonlinear Regression” (If you need to choose again, choose semilog line–X is log, Y is linear”). Make sure “Interpolate unknowns from standard curve” is checked.
- Ensure that R-squared value is at least 0.99 in the Table of Results tab.
- Starting at Row 9, paste in raw average Gapdh Cts under “Gapdh Ct” column; a new Result tab will appear called “Interpolated X Values”.
- Normalize SOD1 Cts using interpolated concentrations
- Create a table in Excel with the following headings: Sample, Target Ct, Gapdh Concentration, Normalization Value, Normalization Coefficient, Log, Normalized SOD1 Ct.
- Use Figure 10A to put appropriate values into each column.
- Find relative copy number
- In a separate sheet, paste the Samples and their Normalized Target Ct values. Rank them from lowest to highest Ct using the Sort function, and assign them rank numbers starting at 1. If there are wild-types in your samples (Cts approaching 40), do not include them in this table.
- Create a table in a separate Excel document with the following headings: Rank, Sample, Ct, Median, Ct-Median, Fold Change, Relative Copies.
- Use Figure 10B to put appropriate values into each column using your new ranked table.
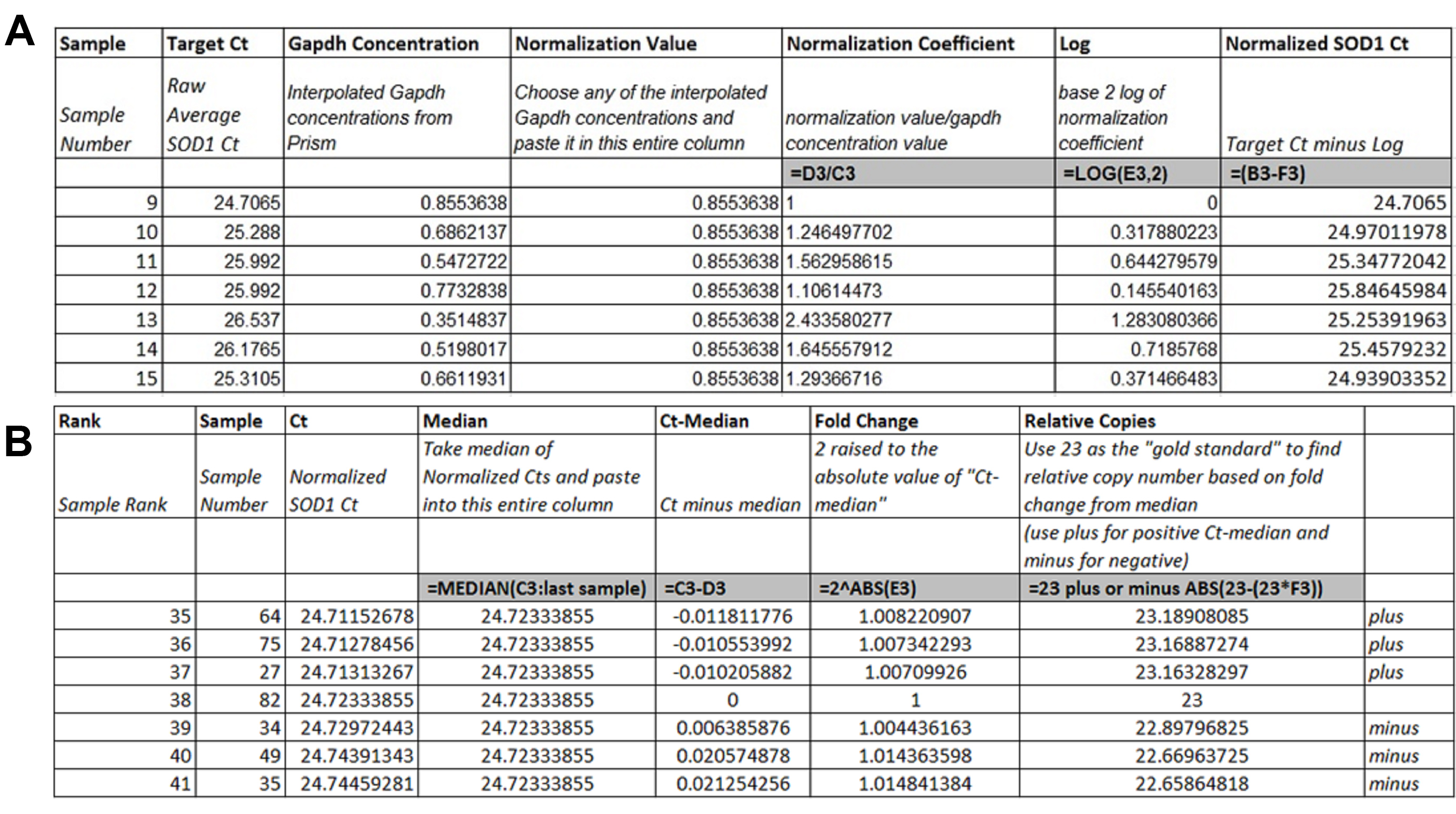
Figure 10. Excel table setup for Ct normalization and relative copy number calculation. A. Normalization of SOD1 Cts using interpolated Gapdh concentration. B. Relative copy number calculation.
- Make genotype calls
- As shown in the Background section, we found that 16 “relative copies” and above causes mice to have a normal lifespan. We make the “low copy” call for mice with 15 relative copies and fewer.
Note: While we found that 15 copies were a safe threshold, you can be as strenuous as you want. For larger and higher-powered efficacy studies it may be worth it to change the threshold to 18 or 20 to make absolutely sure that no lower copy mice are put into the study. - If you are analyzing stud tails, be more conservative because this affects the future of your colony. We use 18 as the cutoff for stud tails.
- It is worth keeping track of stud males for each cohort; if there is a large number of low copies in a cohort, they may all be the progeny of one low copy male.
- As shown in the Background section, we found that 16 “relative copies” and above causes mice to have a normal lifespan. We make the “low copy” call for mice with 15 relative copies and fewer.
- Representative Data
Please see the attached Excel document “Representative Data Spreadsheet 08152018” for a representative set of data, including Excel formulas.
Notes
- Although no genotyping protocol is described in this paper, the mice were genotyped using this SOD1G93A genotyping protocol.
- There are many ways to arrive at high-quality genomic DNA; this method is written about our specific method, which has been optimized and customized for minimal cross-contamination,
- This protocol was written with this piece of equipment in mind; however, you can use another piece of equipment with comparable function.
Recipes
- Sample Prep solution
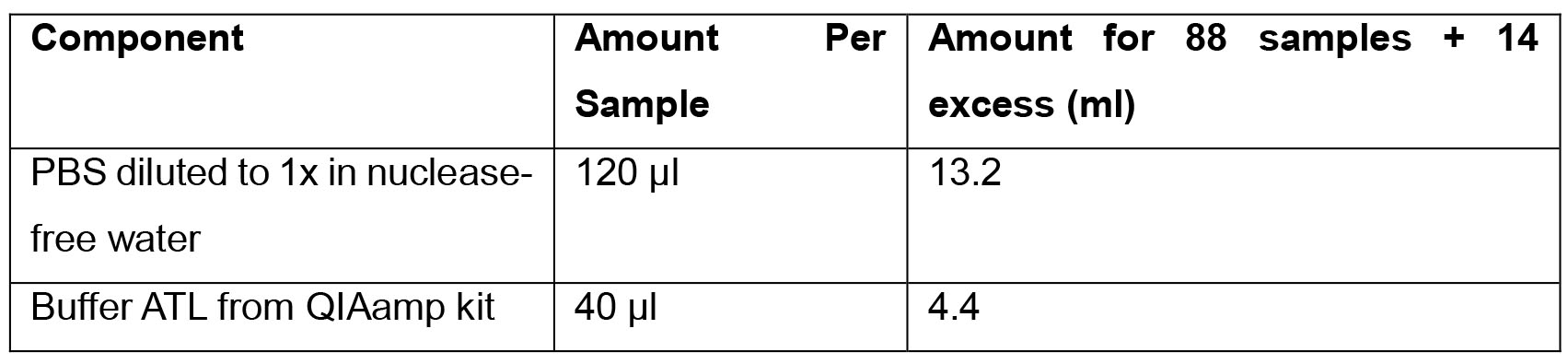
- Tissue Digestion solution
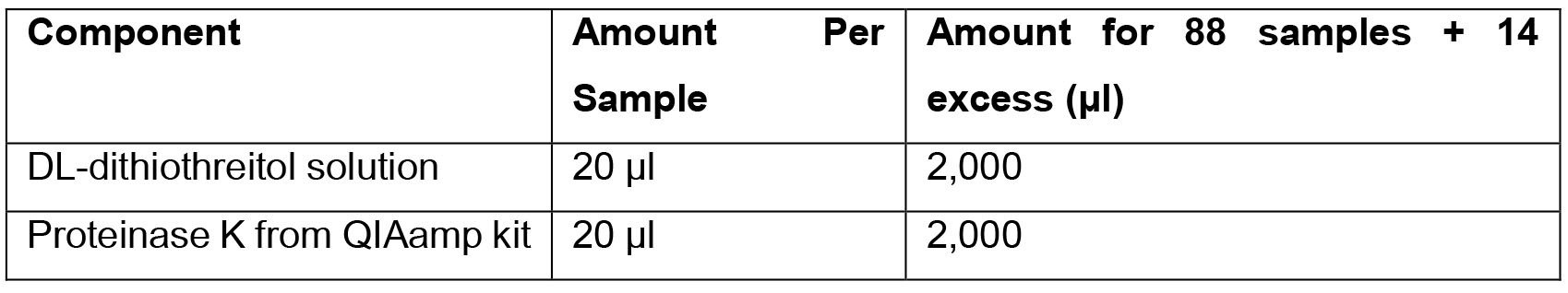
- Primer Dilution solution
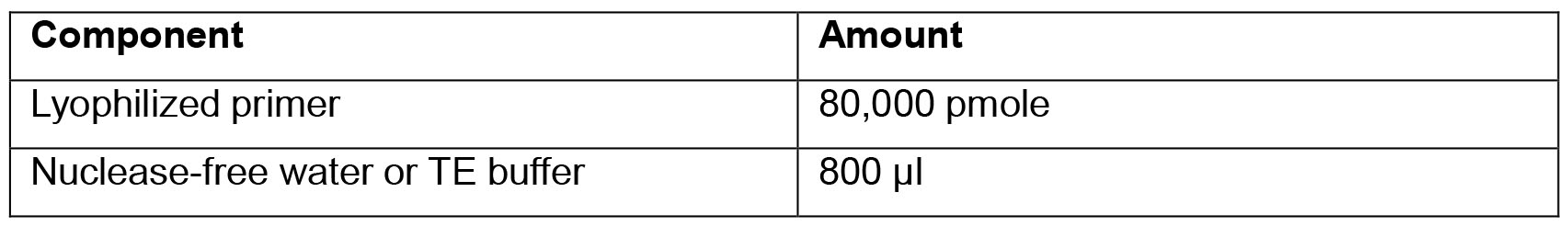
- SOD1G93A Copy Number Genotyping qPCR Master Mix
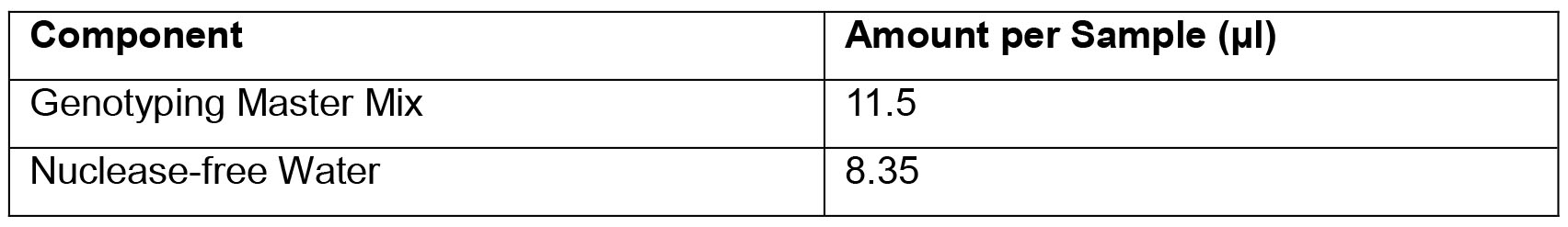
- Primer/Probe solution
Acknowledgments
We acknowledge Matt Ferola and Carlos Maya for diligent and experienced mouse husbandry; we acknowledge Shawn Sullivan for his contribution to the acquisition of historical data. We thank Augie’s Quest for generous support of these studies. We thank people living with ALS for inspiring this research.
Competing interests
The authors declare no competing interests.
Ethics
All experiments were conducted in accordance with the protocols described by the National Institutes of Health Guide for the Care and Use of Animals and were approved by ALS TDI’s institutional animal care and use committee (IACUC). The IACUC ID is 2016-011 and the validity period is December 2016 through December 2019.
References
- Alexander, G. M., Erwin, K. L., Byers, N., Deitch, J. S., Augelli, B. J., Blankenhorn, E. P. and Heiman-Patterson, T. D. (2004). Effect of transgene copy number on survival in the G93A SOD1 transgenic mouse model of ALS. Brain Res Mol Brain Res 130(1-2): 7-15.
- Maier, M., Welt, T., Wirth, F., Montrasio, F., Preisig, D., McAfoose, J., Vieira, F. G., Kulic, L., Spani, C., Stehle, T., Perrin, S., Weber, M., Hock, C., Nitsch, R. M. and Grimm, J. (2018). A human-derived antibody targets misfolded SOD1 and ameliorates motor symptoms in mouse models of amyotrophic lateral sclerosis. Sci Transl Med 10(470).
- Morrice, J. R., Gregory-Evans, C. Y. and Shaw, C. A. (2018). Animal models of amyotrophic lateral sclerosis: A comparison of model validity. Neural Regen Res 13(12): 2050-2054.
- Vieira, F. G., Hatzipetros, T., Thompson, K., Moreno, A. J., Kidd, J. D., Tassinari, V. R., Levine, B., Perrin, S. and Gill, A. (2017). CuATSM efficacy is independently replicated in a SOD1 mouse model of ALS while unmetallated ATSM therapy fails to reveal benefits. IBRO Rep 2: 47-53.
- Zwiegers, P., Lee, G. and Shaw, C. A. (2014). Reduction in hSOD1 copy number significantly impacts ALS phenotype presentation in G37R (line 29) mice: implications for the assessment of putative therapeutic agents. J Negat Results Biomed 13: 14.
Article Information
Copyright
© 2019 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Tassinari, V. R. and Vieira, F. G. (2019). A High-throughput qPCR-based Method to Genotype the SOD1G93A Mouse Model for Relative Copy Number. Bio-protocol 9(12): e3276. DOI: 10.21769/BioProtoc.3276.
Category
Neuroscience > Sensory and motor systems > Animal model
Molecular Biology > DNA > Genotyping
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link









