- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Assessing Yeast Cell Survival Following Hydrogen Peroxide Exposure
Published: Vol 9, Iss 2, Jan 20, 2019 DOI: 10.21769/BioProtoc.3149 Views: 11907
Reviewed by: Emily CopeLionel SchiavolinAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

A Rapid Induction Overexpression System for the Fission Yeast Schizosaccharomyces pombe
Angad Garg
Oct 20, 2021 2704 Views
Spot Assay and Colony Forming Unit (CFU) Analyses–based sensitivity test for Candida albicans and Saccharomyces cerevisiae
Satya Ranjan Sahu [...] Narottam Acahrya
Nov 5, 2023 2827 Views

A Miniaturized Percoll Gradient Method for Isolation of Quiescent Cells of Yeast
Cenxuan Zu [...] Shaolan Zou
Apr 20, 2025 1552 Views
Abstract
In the presence of oxidative stress, cellular defense systems that can detoxify reactive oxygen species are activated through multiple signaling cascades and transcriptional reprogramming. The budding yeast Saccharomyces cerevisiae has served as an excellent model for genetically-identifying factors important for the response to oxidative stress. Here, we describe two assays for testing yeast gene deletion strains or strains overexpressing a gene of interest for viability following oxidative stress induced by hydrogen peroxide treatment. These include a plate-based spot assay for visualizing cell growth and a quantitative colony counting assay. As stress response assays can be highly variable depending on cell growth conditions, these protocols have been optimized for obtaining highly-reproducible results between experiments. We demonstrate the use of these protocols for genetic tests of a putative chromatin regulator implicated in regulating the transcriptional response to oxidative stress.
Keywords: YeastBackground
The unicellular eukaryote Saccharomyces cerevisiae has been an exceptional model for revealing large networks of genes critical to the survival of cells in diverse types of environmental stress (Gasch et al., 2000; Weiner et al., 2012; Ho and Gasch, 2015), largely due to the excellent genetic tools available in this system and the ease with which stress responses can be evaluated. In particular, genetic studies in yeast have uncovered the regulatory mechanisms that control the cells’ response to oxidative stress (Morano et al., 2012). Environmental factors, including radiation and the presence of redox-cycling agents, are a source of oxidative stress which cause increased reactive oxygen species (ROS). These ROS include the superoxide anion, H2O2, and hydroxyl radicals, which are highly reactive with proteins, lipids, and nucleic acids, causing oxidative damage to all of these macromolecules (Schieber and Chandel, 2014). Aerobic respiration also generates low levels of endogenous ROS by the leakage of electrons from the electron transport chain, and mitochondrial dysfunction, as occurs with stress or aging, increases ROS levels further, which triggers the induction of cellular defense mechanisms such as activated expression of antioxidant enzymes which detoxify ROS (Schieber and Chandel, 2014). Pathologies including cancer, cardiovascular disease, autoimmune diseases, and aging are associated with deregulated oxidative stress defense systems and altered levels of intracellular ROS. Therefore, identifying and characterizing the network of proteins responsible for protecting cells during oxidative stress is key to defining the pathophysiology underlying these diseases.
To test the role of a given gene in the response to oxidative stress, it is advantageous to analyze yeast strains which are deleted or otherwise mutated for the gene and which overexpress the protein encoded by the gene. If a particular gene is required to protect cells during oxidative stress, it is expected that its deletion would render cells sensitive to the stress, and its overexpression would cause resistance to the stress. The assays described below test the survival of yeast cells following treatment with hydrogen peroxide, which has served as a model compound for inducing oxidative stress in yeast (Morano et al., 2012; Kwolek-Mirek and Zadrag-Tecza, 2014). Specifically, these protocols were used to test the role of the chromatin regulator Set4 during oxidative stress (Tran et al., 2018). We used a set4∆ yeast strain, and a strain carrying a plasmid for the inducible overexpression of SET4 in the presence of β-estradiol, as previously described (McIsaac et al., 2013). The raw data shown here were obtained to generate the results published in Tran et al. (2018) that demonstrate a role for Set4 in protecting cells during oxidative stress.
We present both a semi-quantitative plate spot assay and a quantitative colony forming unit (cfu) survival assay to interrogate the survival of cells following hydrogen peroxide treatment. We found it useful to be able to visualize the growth differences in cells on the plates and validate these differences quantitatively using the cfu assay. Both protocols here appeared to show much more consistent results than plate spot assays in which hydrogen peroxide is added to the medium in the plate prior to spotting the yeast on the plate (Kwolek-Mirek and Zadrag-Tecza, 2014). In addition, we applied methods to optimize the growth conditions and hydrogen peroxide concentration to account for the observations that yeast cells acquire resistance to hydrogen peroxide when grown in nutrient-poor media, such as synthetic complete or dropout media, or following the diauxic shift, which occurs when cells transition from primarily glucose metabolism via glycolysis to aerobic respiration of ethanol (Guan et al., 2012). When these conditions are not accounted for, results are highly variable and may represent confounding factors in the experiment, rather than the consequences of the deletion or overexpression of the gene being investigated. These assays are not resource-intensive and can be used to test multiple mutant or overexpressing strains simultaneously. Overall, these protocols represent a sensitive and highly-reproducible approach to assaying the response of yeast cells to oxidative stress.
Materials and Reagents
- Pipette tips
- P2 tips (Gilson, catalog number: F171200)
- P200 tips (Gilson, catalog number: F171300)
- P1000 tips (Gilson, catalog number: F171500)
- Toothpicks and inoculating sticks (sterilized) (VWR, catalog number: 12000-806)
- Petri dish (Corning, catalog number: 351029)
- Assay Plate, 96-well, U bottom (Falcon, catalog number: 62406-015)
- 1.5 ml microfuge tubes (Fisherbrand, catalog number: 05-408-129)
- yEG001
- yEG322
- yEG315
- yEG372
- yEG375
- 50 ml conical centrifuge tubes (Falcon, catalog number: 14-432-22)
- Yeast strains (Table 1)
- Vector pMN3 (empty vector) (McIsaac et al., 2013)
- Vector pMN3-SET4 (Tran et al., 2018)
- Sterile, distilled water
- Yeast extract (Research Products International, catalog number: Y20025)
- Peptone (Research Products International, catalog number: P20250)
- Dextrose (Thermo Scientific, catalog number: BP350-1)
- Yeast nitrogen base (US Biologicals, catalog number: Y2025)
- Drop-out Mix Synthetic Minus Uracil (US Biologicals, catalog number: D9535)
- Ethanol (Sigma-Aldrich, catalog number: E7023)
- Hydrogen peroxide (Fisher Scientific, catalog number: H325-100)
- β-estradiol (Millipore Sigma, catalog number: E2758-1G)
- Agar (US biological, catalog number: A0930)
- YPD (Yeast extract-Peptone-Dextrose) medium (see Recipes)
- Synthetic Complete without Uracil (SC-URA) medium (see Recipes)
- 10 mM β-estradiol (see Recipes)
Table 1. Yeast strains used in this protocol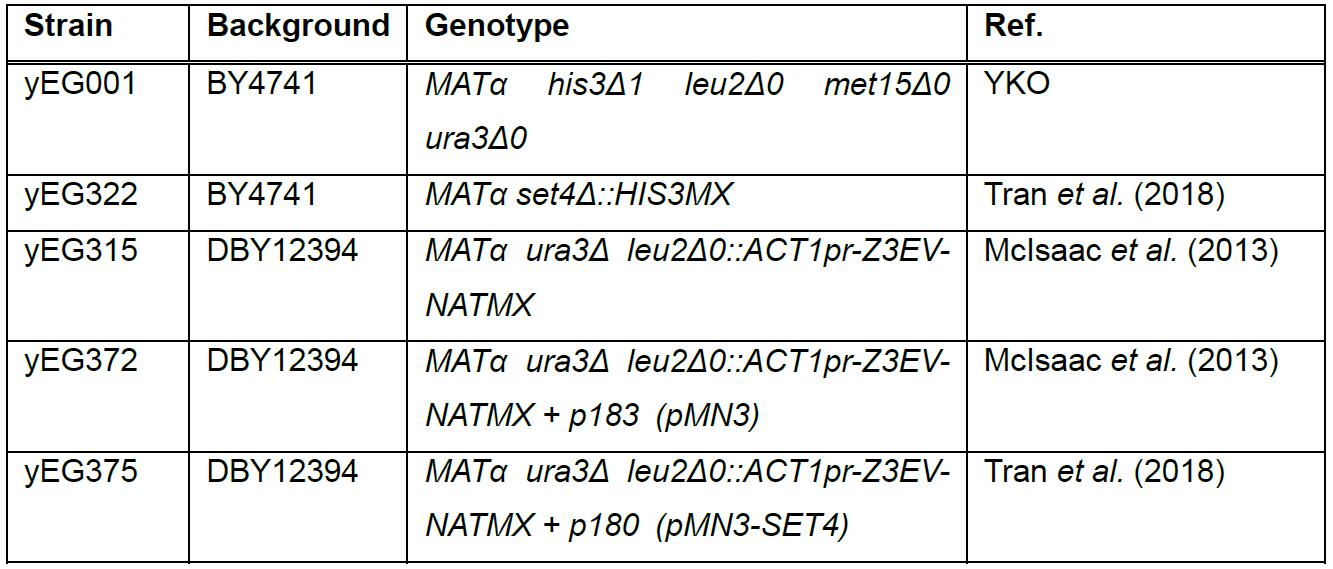
Equipment
- 125 ml (or 50 ml, if available) Erlenmeyer flasks (VWR, catalog number: 10536-912)
- Glass cell spreaders (VWR, catalog number: 89166-788)
- Single-channel pipettors
- P2 pipette (Gilson, catalog number: F144054M)
- P20 pipette (Gilson, catalog number: F144056M)
- P200 pipette (Gilson, catalog number: F144058M)
- P1000 pipette (Gilson, catalog number: F144059M)
- Glass culture tubes (15 ml) (Corning, catalog number: 982016X)
- Multichannel Pipettors (Pipetman Neo P8 x 20N and Pipetman Neo P8 x 200N)
- Incubator
- Rollordrum Tc-7 Tissue Culture Rotator (New Brunswick Scientific)
- Incubator Shaker (New Brunswick Scientific G-25, temperature required: 30 °C; shaker dimensions: 22 in. x 45.5 in. x 30.5 in)
- Vortexer (VWR Standard Heavy Duty Vortex Mixer, catalog number: 97043-562)
- Spectrophotometer (Thermo Scientific, model: NanoDrop 2000c, catalog number: ND-2000c)
- MultiDoc-it Imaging system (UVP)
- Centrifuge (Beckman Coulter Allegra X-14R, catalog number: A99465)
- Microfuge (Beckman Coulter Microfuge 20R)
- Autoclave (Steris AMSCO Century SG-120 Scientific Gravity Sterilizer)
Software
- Excel, Microsoft Office
- Prism GraphPad
Procedure
- Assessing survival of wild-type and mutant yeast cells by spot assay
- Streak wild-type and mutant yeast strains (Table 1) to single colonies on YPD plates. Incubate for 2 days at 30 °C.
- Using a sterile toothpick, select individual wild-type, and mutant colonies. Resuspend the colony in 100 µl YPD in a centrifuge tube. Measure the OD600 using a UV-VIS spectrophotometer (e.g., NanoDrop 2000c) and calculate to determine the volume required to achieve a final cell concentration of OD600 equals 0.0006 in 30 ml YPD. Dilute this volume of culture into 30 ml YPD and grow overnight (~16-18 h for a colony from a plate stored at 4 °C, or ~12-14 h for a colony from a plate coming from the 30 °C incubator) to mid-log phase (OD600 ~0.4-0.5) at 30 °C with shaking at 220 rpm.
- Measure the OD600 of the cultures. Determine the volume of culture equivalent to ~12 OD600 units. For example, 12 OD600 units of a culture with an OD600 of 0.5 equals 24 ml culture (divide 12 by 0.5). Transfer the calculated volume to a 50 ml conical centrifuge tube and centrifuge at 3,900 x g for 3 min.
- Remove supernatant, and resuspend cells in 20 ml pre-warmed YPD.
- Mix cultures well by vortexing, and add 10 ml of culture to two sterile flasks for each strain being tested.
- To one of the flasks, add H2O2 to 4 mM final concentration in 10 ml culture. Commercially-available H2O2 is usually 30% (w/w), density of 1.1 g/ml, and has a molecular weight of 34.01 g/mol. Therefore, the stock concentration is 9.79 M, and 4.08 µl of the stock gives a final concentration of 4 mM in a 10 ml culture volume. The calculation for determining the molarity of the stock solution is shown below:
- Stock solution: 30 g of H2O2 per 100 g H2O
- 100 g H2O/1.1 g/ml = 90.9 ml of solution for 30 g H2O2
- (30 g H2O2/90.9 ml) x 100 = 33.3 g H2O2 in 100 ml H2O
- 33.3 g H2O2/34.01 g/mol = 0.979 mol in 100 ml = 9.79 M
- Place all flasks in the shaker and continue incubating at 30 °C for 30 min.
- After the 30 min incubation, remove 1 ml from each culture and place in a 1.5 ml microfuge tube. Centrifuge at 12,200 x g for 2 min.
- Aspirate supernatant and resuspend the cell pellet in 1 ml sterile, distilled water.
- Set up a 10-fold dilution series of cells in a 96-well plate. You will have a separate row of cells for each strain. In the first well, add 200 μl of undiluted cells. In the next five wells, add 180 μl of sterile water. For each strain, take 20 μl of the undiluted cells and add it to the next well (containing 180 μl of water) using a multichannel pipette. Pipet up and down to mix sample. Then take 20 μl of this dilution and add it to the next well. Pipet up and down to mix and transfer 20 μl to the next well. Repeat to finish the dilution series.
- Using a multichannel pipette, spot 5 μl from each well on to a YPD plate. Start with the most dilute samples and end with the undiluted samples.
- Let the spots dry on the plates, and then transfer to the incubator at 30 °C.
- Cells from untreated cultures should be visible by day 2, whereas cells from cultures treated with H2O2 may take up to 5 days to be visible. Take pictures of the plates every day on a black background with white light using a gel documentation or other imaging system. Representative images are shown in Figure 1. A flow-chart outlining the steps in this protocol is provided in Figure 2.
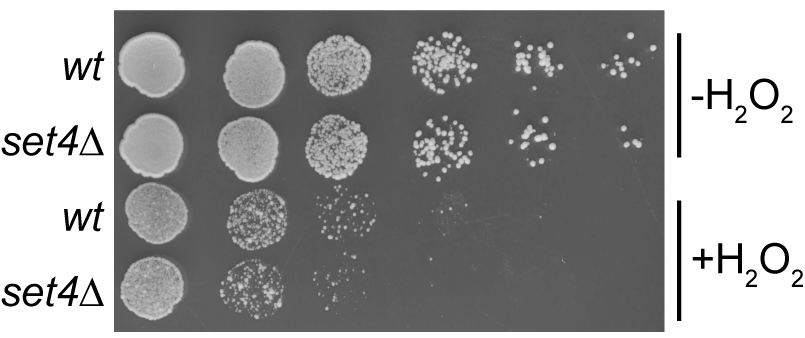
Figure 1. Representative image of a spot assay YPD plate showing serial dilutions of wt and set4∆ cells either untreated or treated with 4 mM H2O2 prior to spotting on the plate. Image was acquired after two days incubation at 30 °C. This type of data corresponds to the experimental outline described in Procedure A.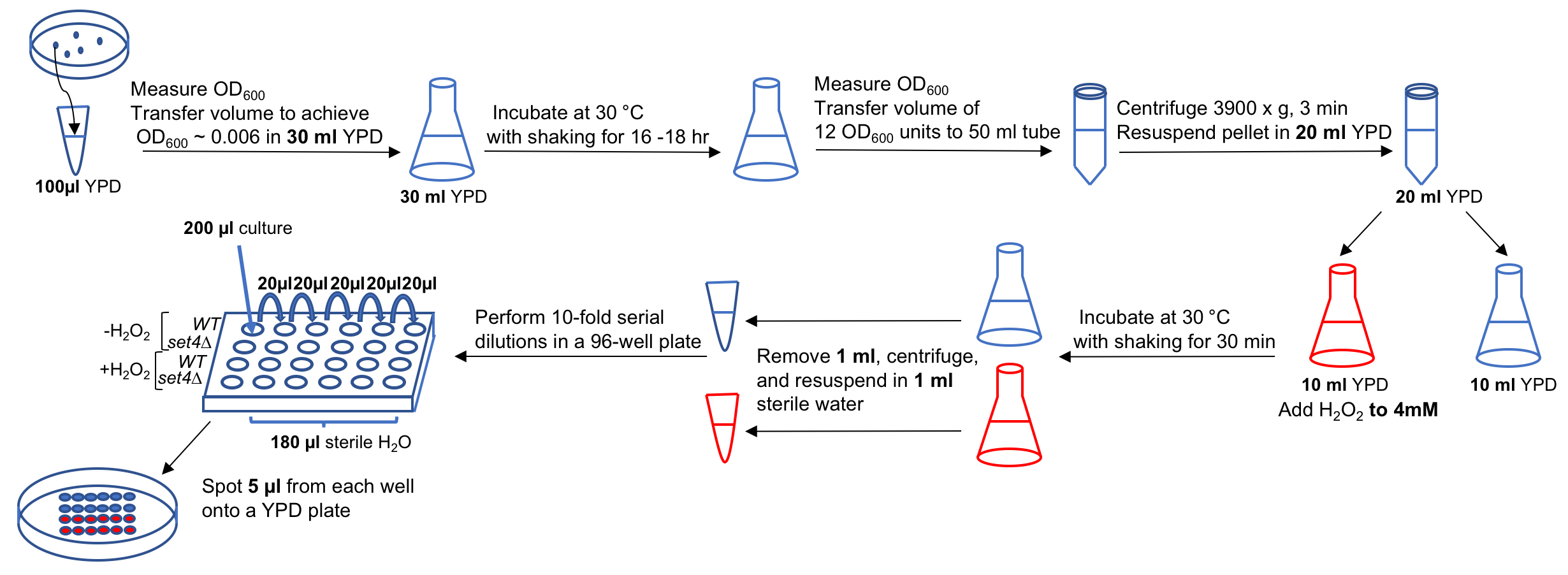
Figure 2. Flow-chart describing experimental outline for Procedure A. Many of the steps shown in the flow-chart are similar to those used in Procedure C, in which SET4 (or a gene of interest) is overexpressed.
- Assessing survival of wild-type and mutant yeast cells by counting colony forming units (cfu)
- Follow Steps A1-A2 to grow yeast strains, except grow only 10 ml of each culture instead of 30 ml.
- When the cultures have reached mid-log phase (OD600 ~0.4-0.5), remove 10 µl of each culture to perform serial dilutions, and follow Step B3 to treat the remaining culture with H2O2. While the culture is incubating (as described below), dilute the 10 µl into 990 µl sterile, distilled water, and then perform a second 1:100 serial dilution. Plate 100 µl of the diluted cells onto a YPD plate using a glass plate spreader, representing a 1:1 x 105 dilution (or a dilution factor, DF, of 105) of the original culture.
- While performing dilutions and plating cells, treat the cultures with H2O2 to 4 mM final concentration (4.08 μl of the H2O2 stock in the 10 ml culture volume). Continue shaking at 30 °C for 30 min.
- After the incubation, remove 10 µl of each culture and dilute once into 990 µl sterile, distilled water. Plate 100 µl of the diluted cells onto a YPD plate using a glass plate spreader, representing a 1:1 x 103 dilution (or a dilution factor, DF, of 103) of the original culture.
- Incubate the plates at 30 °C for 2 days. Count colonies on each of the plates and record values to calculate percent survival, as shown in Table 2 and described in the Data Analysis section. A flow-chart outlining the steps in this protocol is shown in Figure 3.
Table 2. Raw data and percent survival calculations for assessing survival of yeast overexpressing SET4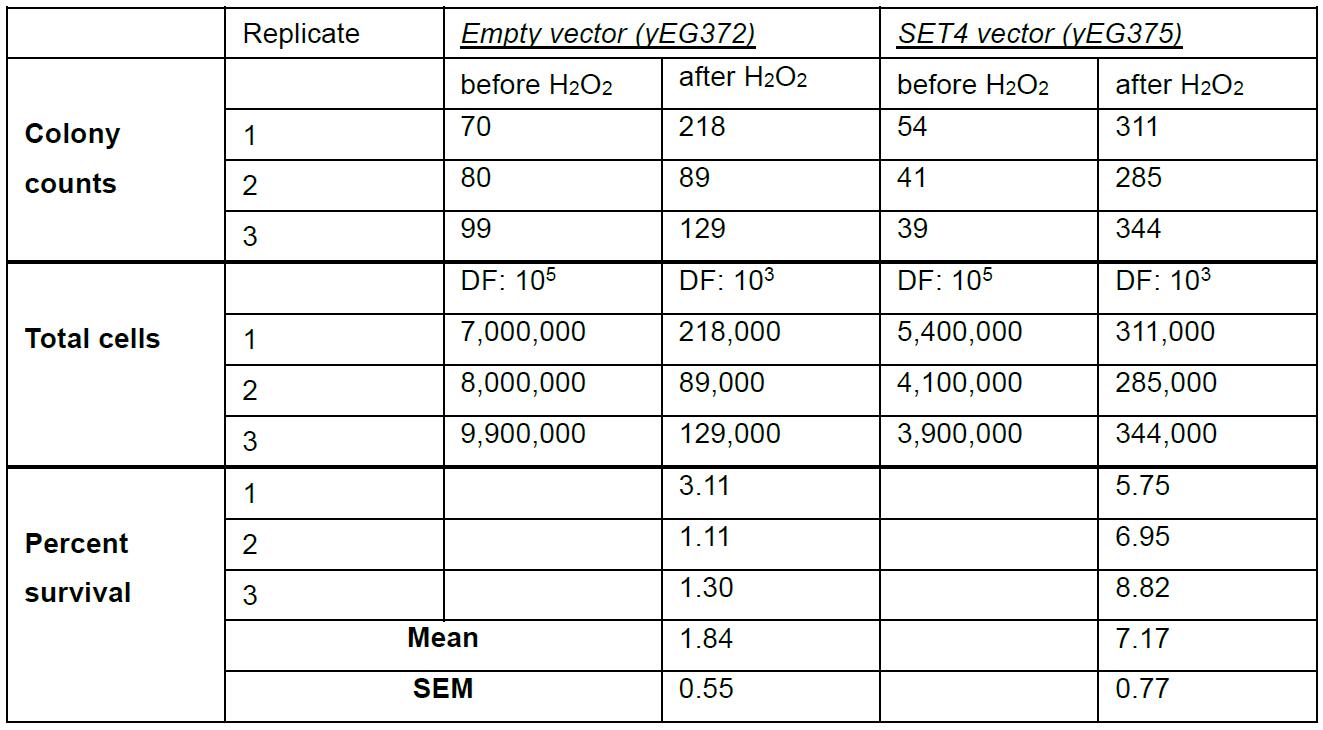
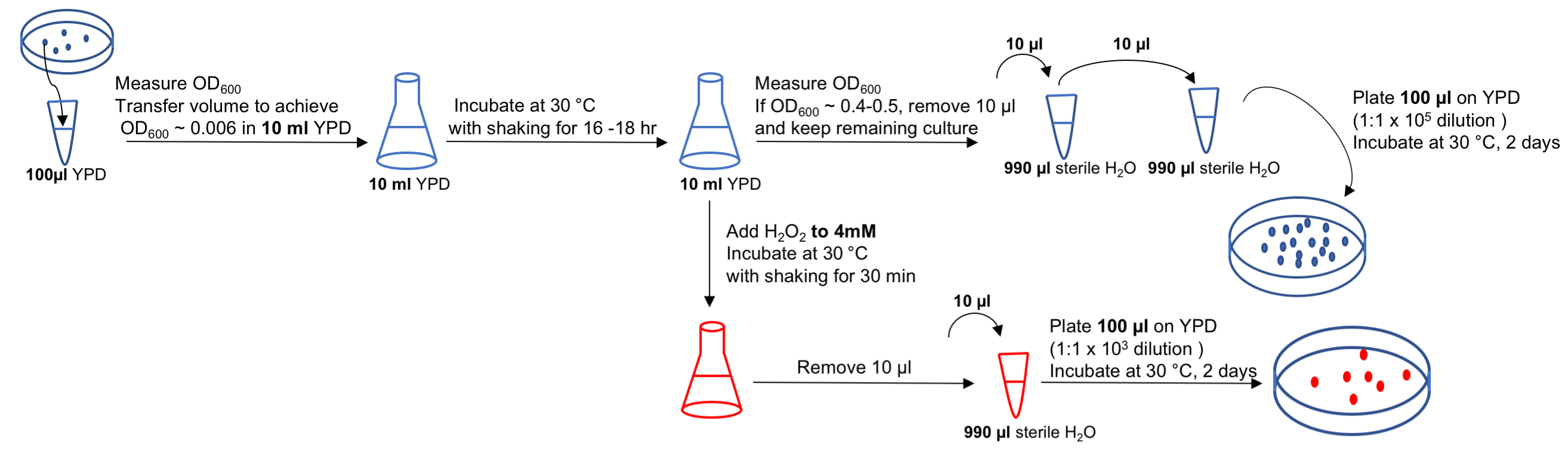
Figure 3. Flow-chart describing experimental outline for Procedure B. Many of the steps shown in the flow-chart are similar to those used in Procedure D, in which SET4 (or a gene of interest) is overexpressed.
- Assessing survival of yeast overexpression strains by spot assay
- For testing survival of cells overexpressing a gene of interest, yeast strain yEG315 was used and transformed (Gietz and Schiestl, 2007) with the vector pMN3, which drives overexpression of a cloned gene in the presence of β-estradiol (strains and plasmids provided by Scott McIsaac; Table 1 [McIsaac et al., 2013]).
- Streak yeast containing the empty vector or overexpression vector onto SC-URA plates to single colonies. Incubate for 2 days at 30 °C.
- Pick a single colony using a sterile inoculating stick and inoculate 3 ml of SC-URA medium in a 15 ml glass culture tube and grow overnight with shaking at 30 °C.
- The next day, measure the OD600 using a spectrophotometer. Dilute the cultures to OD600 ~0.1-0.2 in 20 ml SC-URA medium.
- Freshly prepare 10 mM β-estradiol in ethanol. Further dilute to 0.1 mM in ethanol and add 1 µM β-estradiol to each culture.
- Incubate the cultures with shaking at 30 °C. Grow to mid-log phase (OD600 ~0.5-0.7; approximately 4-6 h).
- Measure the OD600 of the cultures and calculate the volume of culture equivalent to ~12 OD600 units, as described in Procedure A.
- Transfer the calculated volume to a 50 ml conical centrifuge tube and centrifuge at 3,900 x g for 3 min.
- Resuspend in 20 ml pre-warmed SC-URA medium.
- Swirl to mix cultures well, and remove 10 ml of culture to a new flask so that you have two flasks for each strain being tested.
- To one of the flasks, add H2O2 to 20 mM final concentration (20.43 µl of the H2O2 stock in the 10 ml culture volume).
- Place all flasks in the shaker and continue incubating at 30 °C for 30 min.
- After the 30 min incubation, remove 1 ml from each culture and place in a 1.5 ml microfuge tube. Centrifuge at 12,200 x g for 2 min.
- Aspirate supernatant and resuspend the cell pellet in 1 ml sterile, distilled water.
- Follow Steps A10-A13 to set up 10-fold serial dilutions, spot cells on SC-URA plates, and take images of the plates after 2-3 days of growth at 30 °C.
- Assessing survival of yeast overexpression strains by counting colony forming units (cfu)
- Follow Steps C1-C4, except grow only 10 ml cultures in SC-URA (rather than 20 ml).
- Freshly prepare 10 mM β-estradiol in ethanol. Further dilute to 0.1 mM in ethanol and add 50 nM β-estradiol to each culture.
- When the cultures have reached mid-log phase (4-6 h at 30 °C; OD600 ~0.5-0.7), remove 10 µl of each culture and perform serial dilutions. Dilute the 10 µl into 990 µl sterile, distilled water, and perform a second 1:100 serial dilution. Plate 100 µl of the second dilution onto an SC-URA plate using a glass plate spreader, representing a 1:1 x 105 dilution of the original culture.
- While performing dilutions and plating cells, treat the cultures with a final concentration of 20 mM H2O2 (20.43 µl of the H2O2 stock in the 10 ml culture volume) for 30 min. Continue shaking at 30 °C.
- After the incubation, remove 10 µl of each culture and dilute once into 990 µl sterile, distilled water. Plate 100 µl of the diluted cells onto an SC-URA plate using a glass plate spreader, representing a 1:1 x 103 dilution of the original culture.
- Incubate the plates at 30 °C for 2 days. Count colonies on each of the plates and record values to calculate percent survival, as shown in Table 2.
Data analysis
For spot assay tests, plates were imaged using a gel documentation system to qualitatively assess survival following hydrogen peroxide treatment. Examples of these data are shown in Figures 2A and 2E in Tran et al. (2018). To quantitatively measure survival using the colony forming unit (cfu) assay, the total number of colonies on each plate was multiplied by the dilution factor (DF). Percent survival for each replicate was determined by dividing the total colony counts of the H2O2-treated cells by the total colony counts of the untreated cells and multiplying by 100, as outlined in Table 2. The average of the replicates was determined, and the statistical significance was evaluated using an unpaired t-test. These data were analyzed with GraphPad Prism. The raw data and calculations shown in Table 2 are represented graphically in Figure 2F in Tran et al. (2018).
Notes
For both sets of protocols described here, the recommended culture volume to grow to log-phase and perform the H2O2 treatment is a minimum of 10 ml in an Erlenmeyer flask. Although this is a much larger volume than is required for either the spot assay or the cfu assay, we found this volume of culture growth to provide more reproducible results. When similar assays were performed with cells grown in culture tubes and smaller volumes of medium, results were more variable, possibly due to decreased aeration of the cultures.
Another consideration for the H2O2 treatment steps is the different media types used here, either rich YPD medium or synthetic medium. Cells grown in synthetic medium (complete or dropout) develop inherent stress resistance and therefore exhibit higher survival rates upon H2O2 treatment. Therefore, a higher concentration of H2O2 is required when cells are grown in synthetic medium (such as in Procedures C and D), rather than rich medium. We found that 4 mM H2O2 in YPD and 20 mM H2O2 in synthetic medium produced comparable survival rates in wild type cells. However, depending on precise media composition and growth conditions, testing a range of H2O2 concentrations in a pilot experiment may be useful for assay optimization.
Recipes
- YPD (Yeast extract-Peptone-Dextrose) medium (for 1 L)
10 g yeast extract
20 g peptone
20 g dextrose (glucose)
Add distilled water to 1 L, mix well, and autoclave to sterilize
To make YPD plates, add 25 g agar prior to autoclaving. Pour into Petri plates after autoclaved material has cooled enough to be handled - Synthetic Complete without Uracil (SC-URA) medium (for 1 L)
1.92 g uracil drop-out mix
6.7 g yeast nitrogen base
20 g dextrose (glucose)
Add distilled water to 1 L, mix well, and autoclave to sterilize
To make SC-URA plates, add 25 g agar prior to autoclaving. Pour into Petri plates after autoclaved material has cooled enough to be handled - 10 mM β-estradiol
1.1 mg β-estradiol
400 µl ethanol
Make fresh just before use. Dilute further as described in protocol if necessary for accurate pipetting
Acknowledgments
This protocol was developed based on previously published results by the authors (Tran et al., 2018). This work was supported in part by NIH grant R01GM124342 to E.M.G. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Competing interests
The authors declare no competing interests.
References
- Gasch, A. P., Spellman, P. T., Kao, C. M., Carmel-Harel, O., Eisen, M. B., Storz, G., Botstein, D. and Brown, P. O. (2000). Genomic expression programs in the response of yeast cells to environmental changes. Mol Biol Cell 11(12): 4241-4257.
- Gietz RD, Schiestl RH (2007). Quick and easy yeast transformation using the LiAc/SS carrier DNA/PEG method. Nat Protoc 2(1): 35-37.
- Guan, Q., Haroon, S., Bravo, D. G., Will, J. L. and Gasch, A. P. (2012). Cellular memory of acquired stress resistance in Saccharomyces cerevisiae. Genetics 192(2): 495-505.
- Ho, Y. H. and Gasch, A. P. (2015). Exploiting the yeast stress-activated signaling network to inform on stress biology and disease signaling. Curr Genet 61(4): 503-511.
- Kwolek-Mirek, M. and Zadrag-Tecza, R. (2014). Comparison of methods used for assessing the viability and vitality of yeast cells. FEMS Yeast Res 14(7): 1068-1079.
- McIsaac, R. S., Oakes, B. L., Wang, X., Dummit, K. A., Botstein, D. and Noyes, M. B. (2013). Synthetic gene expression perturbation systems with rapid, tunable, single-gene specificity in yeast. Nucleic Acids Res 41(4): e57.
- Morano, K. A., Grant, C. M. and Moye-Rowley, W. S. (2012). The response to heat shock and oxidative stress in Saccharomyces cerevisiae. Genetics 190(4): 1157-1195
- Schieber, M. and Chandel, N. S. (2014). ROS function in redox signaling and oxidative stress. Curr Biol 24(10): R453-462.
- Tran, K., Jethmalani, Y., Jaiswal, D. and Green, E. M. (2018). SET4 is a chromatin-associated protein, promotes survival during oxidative stress, and regulates stress response genes in yeast. J Biol Chem 293(37): 14429-14443.
- Weiner, A., Chen, H. V., Liu, C. L., Rahat, A., Klien, A., Soares, L., Gudipati, M., Pfeffner, J., Regev, A., Buratowski, S., Pleiss, J. A., Friedman, N. and Rando, O. J. (2012). Systematic dissection of roles for chromatin regulators in a yeast stress response. PLoS Biol 10(7): e1001369.
Article Information
Copyright
© 2019 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Readers should cite both the Bio-protocol article and the original research article where this protocol was used:
- Tran, K. and Green, E. M. (2019). Assessing Yeast Cell Survival Following Hydrogen Peroxide Exposure. Bio-protocol 9(2): e3149. DOI: 10.21769/BioProtoc.3149.
- Tran, K., Jethmalani, Y., Jaiswal, D. and Green, E. M. (2018). SET4 is a chromatin-associated protein, promotes survival during oxidative stress, and regulates stress response genes in yeast. J Biol Chem 293(37): 14429-14443.
Category
Microbiology > Microbial cell biology > Cell viability
Microbiology > Microbial genetics > Gene expression
Cell Biology > Cell viability > Cell survival
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link









