- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Quantitative Analysis of Cargo Density in Single-extracellular Vesicles by Imaging
Published: Vol 8, Iss 24, Dec 20, 2018 DOI: 10.21769/BioProtoc.3111 Views: 6464
Reviewed by: Yanjie LiLuis Alberto Sánchez VargasNaresh Kasoju

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
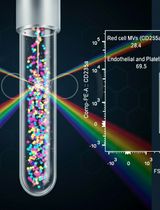
Protocol for the Isolation and Analysis of Extracellular Vesicles From Peripheral Blood: Red Cell, Endothelial, and Platelet-Derived Extracellular Vesicles
Bhawani Yasassri Alvitigala [...] Lallindra Viranjan Gooneratne
Nov 5, 2025 1503 Views

Production of Genetically Engineered Extracellular Vesicles for Targeted Protein Delivery
Leyla A. Ovchinnikova [...] Yakov A. Lomakin
Nov 5, 2025 1674 Views
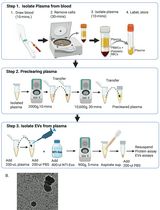
A Rapid and High-Recovery Extracellular Vesicle (EVs) Isolation Technique from Blood Samples
Aryan N. Sankpal and Narendra V. Sankpal
Mar 5, 2026 82 Views
Abstract
Function of extracellular vesicles such as exosomes and microvesicles is determined by their wide ranges of cargoes inside them. Even in the pure exosomes or microvesicles the cargo contents are very heterogeneous. To understand this heterogeneous nature of extracellular vesicles, we need information of the vesicles, which will give us some parameters including vesicle size, number and cargo content of each vesicle. Here, we describe a new method to quantify cargo density in single-extracellular vesicles. Staining of extracellular vesicles in a membrane lipid content-proportionate manner and immobilization of extracellular vesicles onto glass substrate allow us to obtain cargo density information of single-extracellular vesicles. This protocol will be useful to analyze the effects of various drugs or genetic manipulation on vesicle generation and maturation including cargo sorting into heterogeneous extracellular vesicles.
Keywords: Extracellular vesicleBackground
Extracellular vesicles are small membrane-bound vesicles released from a variety of cells, comprising exosomes and microvesicles based on the current knowledge of their biogenesis (van Niel et al., 2018). Extracellular vesicles contain various kinds of cargoes including cytosolic proteins, membrane proteins, lipids, microRNAs, and mRNAs. The cargoes are highly heterogeneous in their contents and variety depending on cell types. Extracellular vesicles play a role in erythrocyte maturation, antigen presentation, tumor metastasis, or prion disease and Alzheimer’s disease propagation, through cell-to-cell communication by carrying their cargoes from the donor to the recipient cells (Maas et al., 2017).
Exosomes are formed by fusion of multivesicular endosomes (MVEs) with the plasma membranes. On the other hand, microvesicles are formed by shedding of the plasma membranes. Endosomal sorting complexes required for transport (ESCRT) is known to be critical for the generation of lysosomal MVEs. It is reported that ceramide is important for the generation of exosomal MVEs (Trajkovic et al., 2008). The size of exosomes ranges from 30 nm to 100 nm, whereas that of microvesicles is from 50 nm to 1,000 nm (Karpman et al., 2017; van Niel et al., 2018). The mechanism of cargo sorting into extracellular vesicles, especially microvesicles, is still mysterious. For understanding the heterogeneous nature of extracellular vesicles, it is important to get more information on cargo density as well as the number and the size of single extracellular vesicles.
Conventionally, we can quantitate the amount of cargo in total extracellular vesicles obtained from culture media using western blot analysis. Since extracellular vesicles are highly heterogeneous as mentioned above, we need to get more precise figures of extracellular vesicles for the understanding of vesicle generation and maturation including the cargo sorting. We can solve this issue with detecting “cargo density of single-extracellular vesicles”. Here, we provide details on our new method for getting information of each extracellular vesicle with a cargo density. This protocol provides a useful readout of cargo content in each extracellular vesicle, and promote to unravel detailed molecular mechanism of the cargo sorting and accelerate clinical application of extracellular vesicles.
Materials and Reagents
- 10 μl, 200 μl, or 1,000 μl tips (Fukae-Kasei, Watson, catalog numbers: 123R-254CS, 123R-755CS, 110-706C)
- 200 μl gel loading tips (Quality Scientific Plastics, catalog number: 010-Q)
- 5 ml or 10 ml pipettes (Corning, catalog numbers: 4487, 4488)
- 50 ml conical centrifuge tubes (BD, Falcon, catalog number: 352070)
- 1.5 ml microcentrifuge tubes (Ina Optika, BIO-BIK, catalog number: CF-0150)
- 35 mm or 100 mm culture dish (Corning, catalog numbers: 430165, 430167)
- 6-well plate (Corning, catalog number: 3516)
- 35 mm glass bottom culture dish (MatTek, catalog number: P35G-0-14-C)
- 0.22 μm syringe filters (Millipore, catalog number: SLGVJ13SL)
- Kimwipes (Nippon Paper Crecias, catalog number: 62011)
- Aluminum foil (Mitsubishi Aluminum, Nippaku foil)
- Ultracentrifuge thick wall polycarbonate tubes (Beckman Coulter, catalog number: 349622)
- HeLa cells (RIKEN cell bank, catalog number: RCB0007)
- MDA-MB-231 cells (ATCC, catalog number: HTB-26)
- Dulbecco's modified Eagle medium (DMEM), high glucose, with L-Glutamine and Phenol Red (FUJIFILM Wako Pure Chemical, catalog number: 044-29765)
- Fetal bovine serum (FBS) (Biowest, catalog number: S1820)
- Penicillin-streptomycin Mixed Solution (P/S) (NACALAI TESQUE, catalog number: 09367-34)
- Trypsin-EDTA solution with phenol red (NACALAI TESQUE, catalog number: 32778-34)
- FuGENE HD transfection reagent (Promega, catalog number: E2311)
- Opti-MEM, reduced serum medium (Thermo Fisher Scientific, InvitrogenTM, catalog number: 31985070)
- Extracellular vesicles-free fetal bovine serum (Thermo Fisher Scientific, GibcoTM, catalog number: A2720803)
- Total exosome isolation reagent (from cell culture media) (Thermo Fisher Scientific, InvitrogenTM, catalog number: 4478359)
- Paraformaldehyde (PFA) powder (Sigma-Aldrich, catalog number: P6148)
- Normal goat serum (Jackson Immuno Research, catalog number: 005-000-121)
- Lipophilic Tracer, DiD (Ex. 644 nm; Em. 665 nm) (Thermo Fisher Scientific, InvitrogenTM, catalog number: D7757)
- Biotinyl Cap PE (PE-biotin) (Avanti Polar Lipids, catalog number: 870277)
- Biotin-conjugated bovine serum albumin (BSA-biotin) (Sigma-Aldrich, catalog number: A8549)
- Bovine serum albumin (BSA) (Sigma-Aldrich, catalog number: A8022)
- Streptavidin (Thermo Fisher Scientific, Invitrogen, catalog number: 434301)
- Antibody of cargo (e.g., anti-cMET antibody; Cell Signaling, catalog number: 3127S)
- Secondary antibody (e.g., anti-mouse IgG Alexa Fluor 594; Cell Signaling, catalog number: 8890S)
- Plasmid DNA of cargo (e.g., constructing plasmid DNA of GFP- or mCherry-fused cargo protein)
- Sodium chloride (NaCl) (NACALAI TESQUE, catalog number: 31319-45)
- Potassium chloride (KCl) (Wako, catalog number: 163-03545)
- di-Sodium hydrogenphosphate (Na2HPO4) (NACALAI TESQUE, catalog number: 31723-35)
- Potassium dihydrogenphosphate (KH2PO4) (NACALAI TESQUE, catalog number: 28721-55)
- NaOH (NACALAI TESQUE, catalog number: 31511-05)
- Phosphate-buffered saline (PBS) (see Recipes)
- 4% PFA (see Recipes)
Equipment
- Glass vial (Maruemu, catalog number: 5-115-02)
- Forceps
- Pipettes (Gilson, models: PIPETMAN P10, P20, P200, P1000)
- Pipetter (Drummond Scientific, model: Pipet-Aid XP)
- Polypropylene microtube storage rack, 80 positions
- Test tube rack (Sanwakaken, catalog number: SS30-10)
- Clean hood (NK System, model: VH-1300BH-2A)
- CO2 incubator (Forma Scientific, model: 3158)
- Confocal laser scanning microscope (Zeiss, model: LSM 510 Meta)
- Transmitted Light Microscope (Olympus, model: IMT-2)
- Ultracentrifuge (Beckman Coulter, model: Optima TL)
- Ultracentrifuge rotor (Beckman Coulter, model: TLA-100.3)
- Refrigerated microcentrifuge (Tomy Seiko, model: MRX-151)
- Vortex mixer (Scientific Industries, model: Vortex-Genie 2)
- Refrigerator, 4 °C
- Freezer, -20 °C
- Cold room, 4 °C
Software
- LSM510 v4.2 operating software (Zeiss)
- ImageJ (https://imagej.nih.gov/ij/)
- Excel for Mac 2011 (Microsoft)
Procedure
- Overall procedure
Overall procedure for analyzing cargo density of single-extracellular vesicles is shown in Figure 1.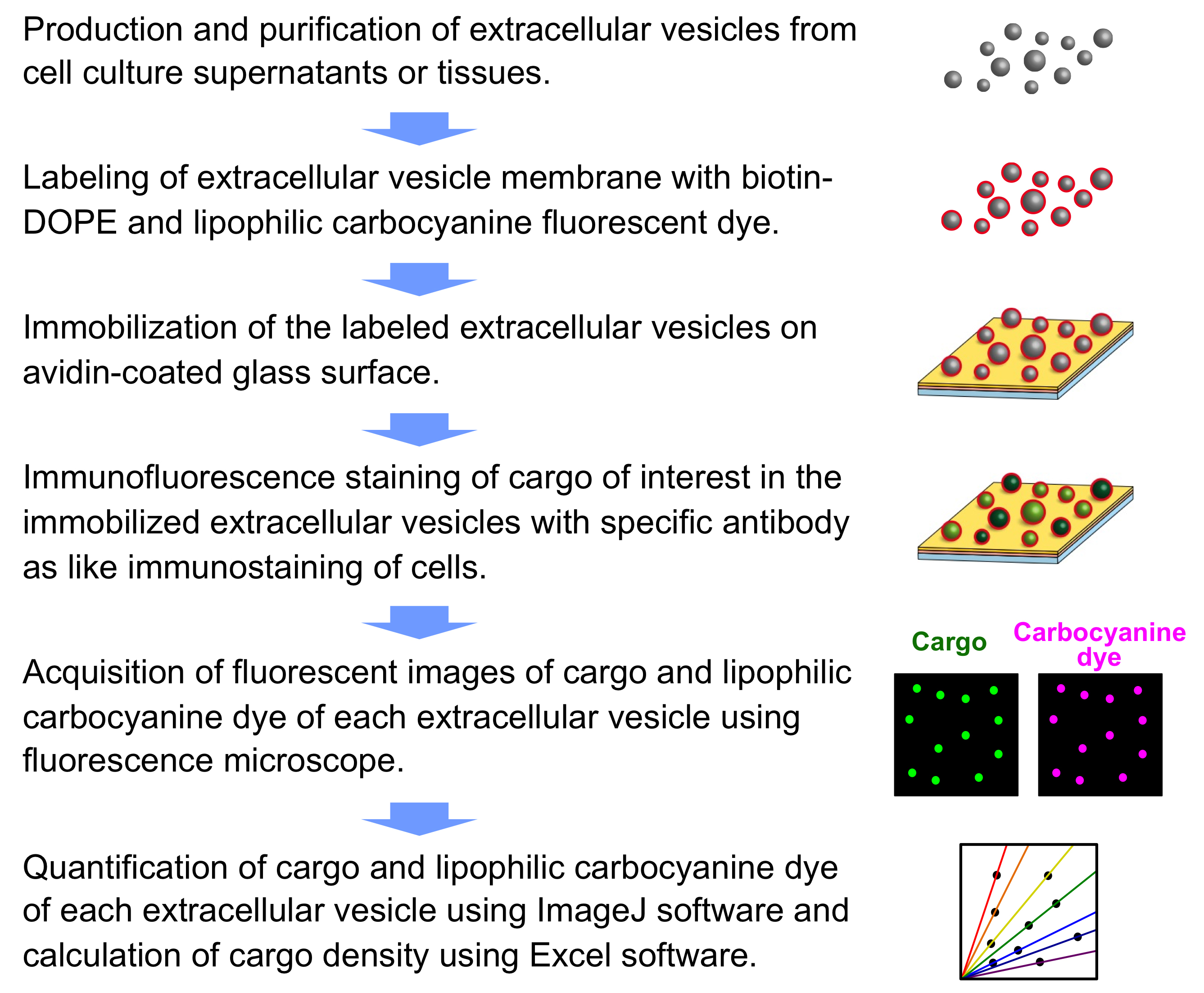
Figure 1. Overall procedure for detecting cargo density in single-extracellular vesicles - Production and purification of extracellular vesicles
- Cell culture and transfection for producing released extracellular vesicles
- HeLa cells or MDA-MB-231 cells are plated onto 35 mm culture dish or 6-well plate and maintained in Dulbecco’s modified Eagle medium (DMEM) containing 10% fetal bovine serum (FBS) and 1% penicillin/streptomycin (P/S) at 37 °C in 5% CO2.
- When cells are at 70% confluence, remove the medium and wash the cells 2 times each with 2 ml of 37 °C DMEM without FBS and P/S.
- Replace medium with 1.2 ml of 37 °C DMEM containing 10% extracellular vesicles-free fetal bovine serum and 1% P/S.
- Incubate at 37 °C in 5% CO2 for 24 to 48 h.
- Purification of extracellular vesicles
- Flowchart of this part is shown in Figure 2.
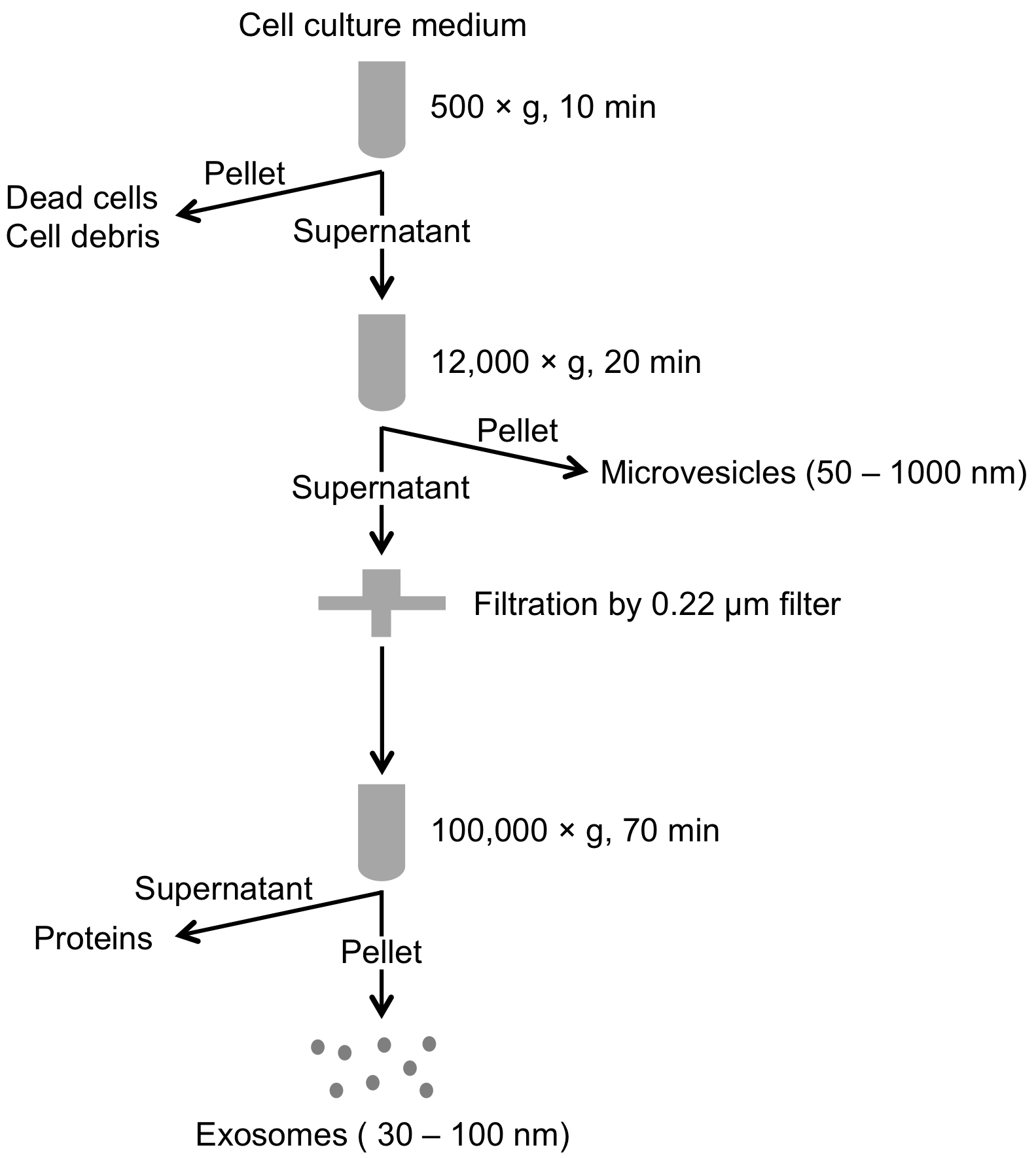
Figure 2. Flowchart of isolation of extracellular vesicles (exosomes and microvesicles) by differential centrifugation - Cell culture media (1.2 ml) from 90%-95% confluent cells is collected in 1.5 ml microcentrifuge tube.
- Centrifuge at 500 x g at 4 °C for 10 min to remove the cell debris using a refrigerated microcentrifuge.
- Collect all 1.2 ml supernatant and transfer to a new 1.5 ml microcentrifuge tube.
- Centrifuge at 12,000 x g at 4 °C for 20 min using a refrigerated microcentrifuge to separate microvesicles and others.
- Collect the upper 1 ml of supernatant is directly purified by filtration through a 0.22 μm pore filter and transfer into the ultracentrifuge thick wall polycarbonate tube. Approximately 70 µl of sample is lost after the filtration.
- The pellet from Step B2e is diluted in 200 μl of phosphate-buffered saline (PBS) (Microvesicle-enriched fraction).
- Ultracentrifuge the filtrated supernatant using ultracentrifuge (Optima TL, rotor: TLA-100.3) at 100,000 x g at 4 °C for 70 min.
- Remove supernatant (approximately 930 µl) carefully.
- Dilute the pellet in 200 μl of PBS (Exosome-enriched fraction).
- Flowchart of this part is shown in Figure 2.
- Cell culture and transfection for producing released extracellular vesicles
- Preparation of functionalized glass surface
- Flowchart of this part is shown in Figure 3.
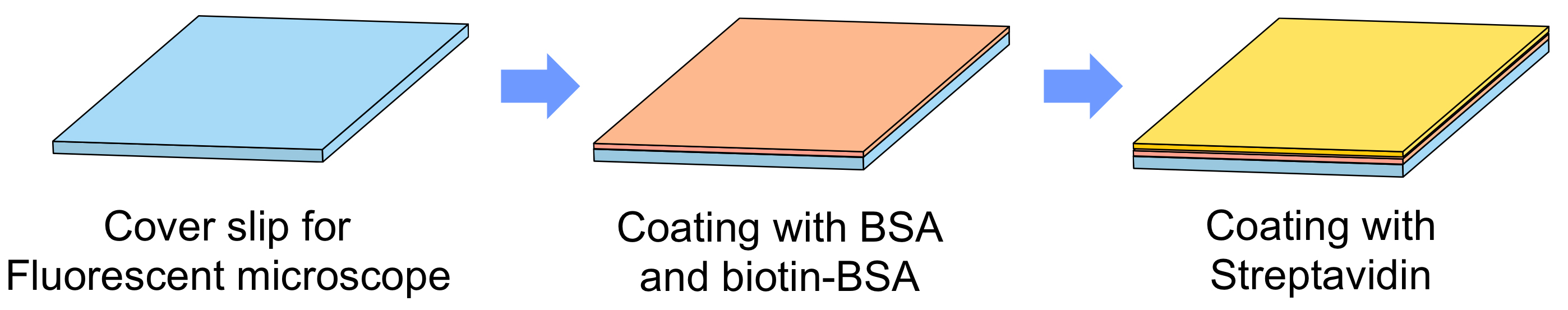
Figure 3. Flowchart of preparation of functionalized glass surface - Prepare BSA/BSA-biotin mixture with mixing 5 μl of 10 M BSA and 5 μl of 1 M BSA-biotin in 490 μl of PBS.
- Cover the glass surface of glass bottom 35 mm culture dish with 400 μl of the BSA/BSA-biotin mixture.
- Incubate at room temperature for 20-30 min.
- Prepare 25 mM streptavidin solution by diluting 8 μl of 1 M streptavidin in 312 μl of PBS.
- Remove excess BSA/BSA-biotin by gently rinsing the glass surface once with 1 ml of PBS.
- Cover the glass surface with 300 μl of the streptavidin solution.
- Incubate at room temperature for 10 min.
- Gently rinse the BSA/BSA-biotin/streptavidin-coated glass surface twice each with 1 ml of PBS.
- Flowchart of this part is shown in Figure 3.
- Labeling of extracellular vesicles
- Transfer the isolated extracellular vesicles (200 μl total) into a glass vial.
- Add 2 μl of 10 mg/ml DOPE-biotin into the glass vial and suspend to mix.
- Incubate at room temperature for 5 min.
- Add 2 μl of 1 mg/ml DiD into the glass vial and suspend to mix.
Note: Because DiD staining is the most critical step for calculating the cargo density, it is important to take exact amount of DiD. - Spin down the glass vial in a 50 ml conical tube at 200 x g at room temperature and take it out with forceps.
- Then incubate at room temperature for 15 min in the dark (Figure 4).

Figure 4. Labeling of extracellular vesicles with DOPE-biotin and DiD in glass vial
- Immobilization of extracellular vesicles on glass surface
- The DOPE-biotin- and DiD-labeled extracellular vesicles are applied onto the functionalized glass surface using gel loading long tip (Figure 5).
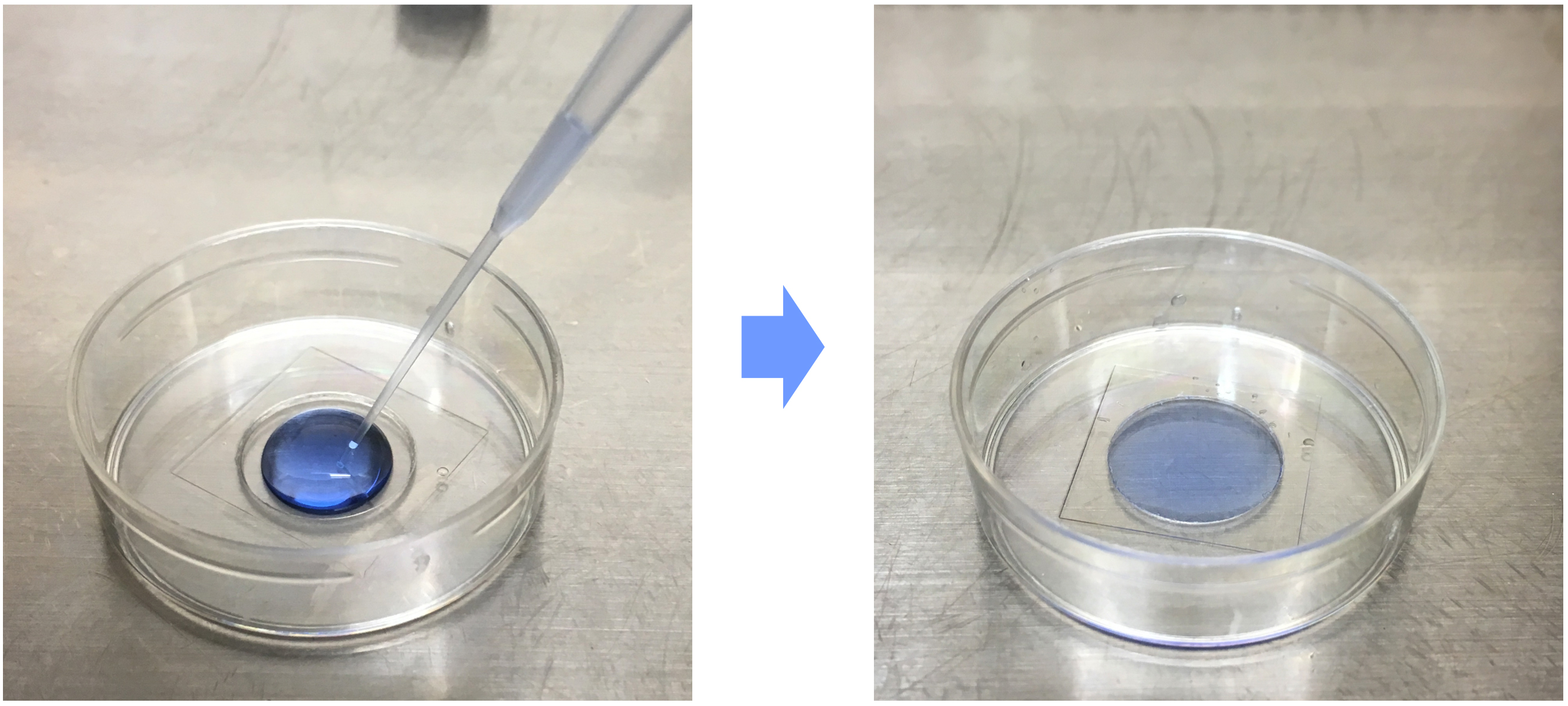

Figure 5. Immobilization of extracellular vesicles on functionalized glass surface. DOPE-biotin- and DiD-labeled extracellular vesicle solution is applied onto functionalized glass surface and made it flat against the surface tension. - Leave to stand at 4 °C for 2-3 h in dark condition.
- Rinse twice each with 2 ml of PBS.
- The DOPE-biotin- and DiD-labeled extracellular vesicles are applied onto the functionalized glass surface using gel loading long tip (Figure 5).
- Immunofluorescence staining of endogenous cargo in extracellular vesicles
- Fix immobilized extracellular vesicles with 4% PFA for 5 min in dark condition.
- Rinse extracellular vesicles twice with 2 ml of PBS and pipette out all solution.
- Add 150 μl of PBS containing 10% normal goat serum and incubate at room temperature for 30 min in dark condition.
- Rinse extracellular vesicles once with 2 ml of PBS and pipette out all solution.
- Add 150 μl of anti-cargo antibody (e.g., anti-cMet antibody) in PBS and incubate at room temperature for 60-90 min in dark condition.
- Rinse extracellular vesicles once with PBS and pipette out all solution.
- Add 150 μl of fluorescent-dye conjugated secondary antibody (e.g., Alexa 488 or Alexa 594) in PBS and incubate at room temperature for 30 min in dark condition.
- Rinse extracellular vesicles twice each with 2 ml of PBS.
- Store at 4 °C in dark condition, and observe in 2-3 days.
- Imaging of fluorescence-labeled cargo and DiD using fluorescence microscope
Note: For this imaging part we will show about the exosome sample as a representative.- Turn on the confocal laser scanning microscope (Zeiss, LSM 510 Meta).
- Open the Zeiss LSM510 v4.2 operating software.
- Turn on the appropriate laser (e.g., Argon laser for excitation of GFP or Alexa 488, 543 nm wavelength He-Ne laser for excitation of mCherry or Alexa 594, 633 nm wavelength He-Ne laser for excitation of DiD).
- Select 63x objective lens.
- Set the glass bottom 35 mm culture dish with immobilized and cargo-labeled extracellular vesicles on the specimen holder.
- Carefully find the glass surface using visible light under eyepiece viewing.
- Change the observation status to laser scanning.
- In the “Scan Control panel”, “Mode”, set the level of zoom to x1 (Default).
- In the “Scan Control panel”, “Channels”, set the pinhole to “200”.
- Adjust the focus with DiD channel using “Fast XY”.
- Collect averaged images (e.g., number is selected to 4 for the average) of DiD and fluorescent cargo (e.g., GFP- or mCherry-fused cargo or immunostained cargo with anti-mouse IgG Alexa 488 or anti-mouse IgG Alexa 594 as secondary antibody) and save them as lsm format.
- The acquired images of exosomes are shown in Figure 6 (in case of endogenous cargo) or in Kajimoto et al. (2013) or (2018) (in case of exogenous GFP- or mCherry-fused cargo).
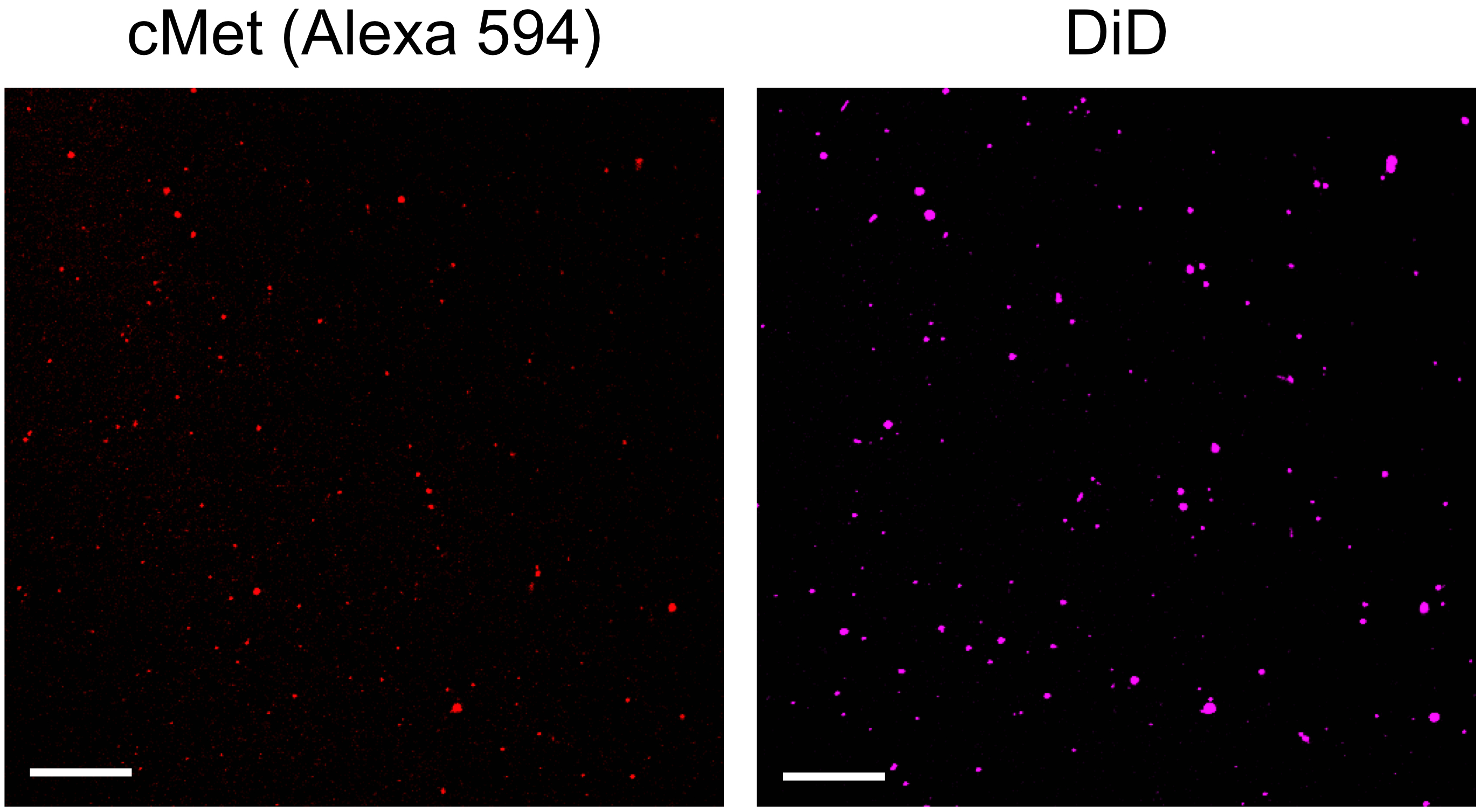
Figure 6. Fluorescent images of cargo (cMet-Alexa 594) and DiD in single-exosomes. Exosomes released from MDA-MB-231 cells are stained with anti-cMet antibody with Alexa 594 secondary antibody and DiD. Scale bars, 10 μm.
Data analysis
- Open the ImageJ software.
- Go to “Adjust” → “Set Measurements”, and check “Area”, “Mean gray value”, and “Integrated Density”, then click “Ok” button.
- Open the acquired images (lsm format file, here using the data of Figure 6 as example) with drag-and-drop to ImageJ icon (Figure 7A).
- Select DiD image (Channel 2) (Figure 7B).
- Set “Image” → “Type” → “8-bit” and confirm that the image is converted to greyscale (Figure 7C).
- Set “Edit” → “Invert” → “No (against the question “Process all 3 images?”)” for inverting the image of DiD (Figure 7D).
- Use “Image” → “Adjust” → “Threshold” to highlight the extracellular vesicles (Figure 7E). Basically using “Auto” would be working well. If necessary, adjust the threshold to the suitable level to count.
- Go to “Analyze” → “Analyze Particles”. There are some options in the “Analyze Particles” window (Figure 7F). Basically use default setting (Size: 0-Infinity, Circularity: 0.00-1.00, Show: Nothing) and just check “Display results” and “Add to Manager”. If there are too many small noises or you want to exclude particles based on size, adjust the numbers of “Size”.
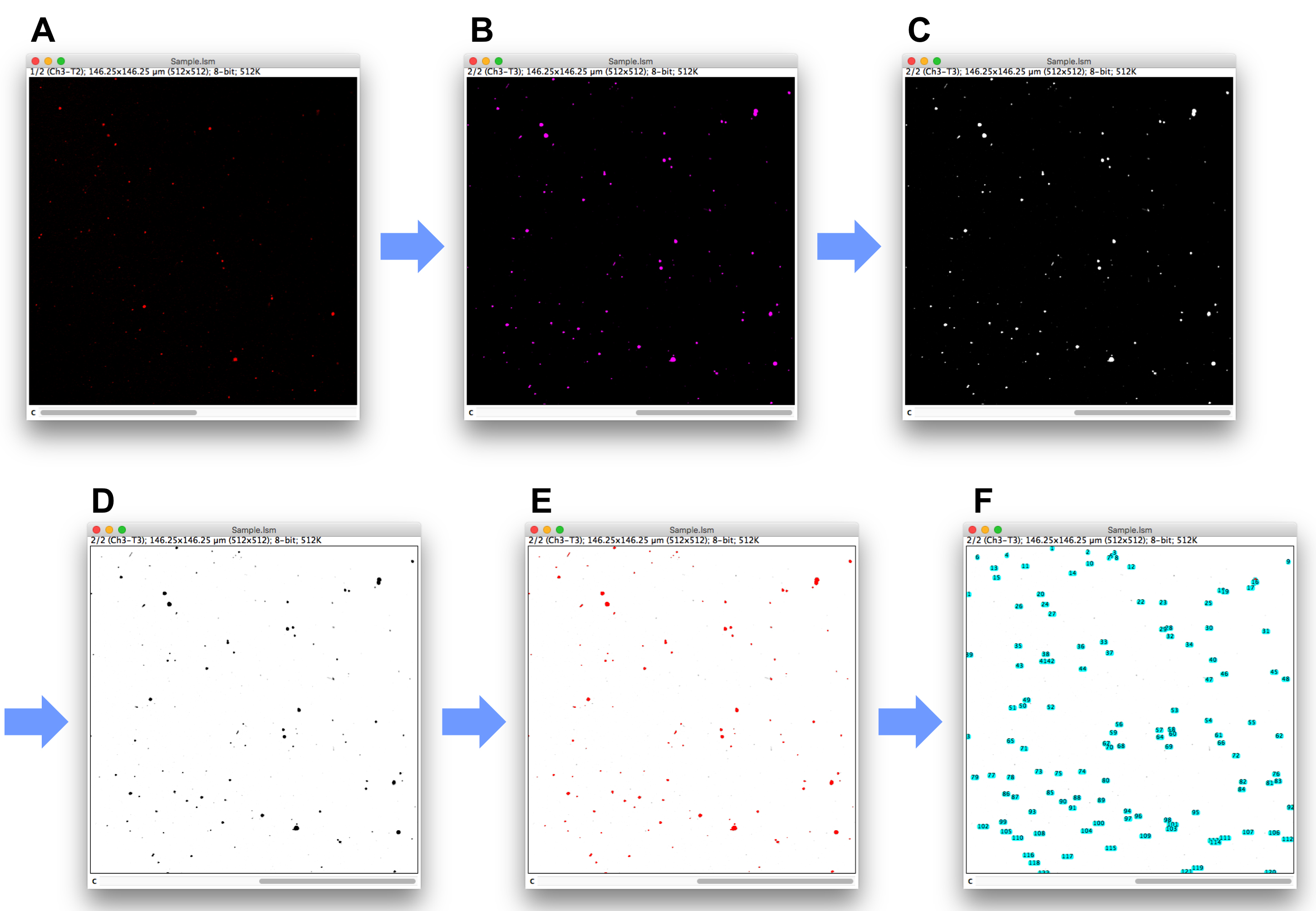
Figure 7. Conversion of acquired images of cargo (cMet-Alexa 594) and DiD to be calculated with ImageJ - In the “ROI Manager” window, go to “More” → “Multi Measure”, and just check “Measure All 2 Slices” and click “Ok” button.
- Get “Area”, “Mean gray value (Mean)”, and “Integrated Density (IntDen)” information of cargo (in channel 1) and DiD (in channel 2) of each extracellular vesicle (Figure 8).
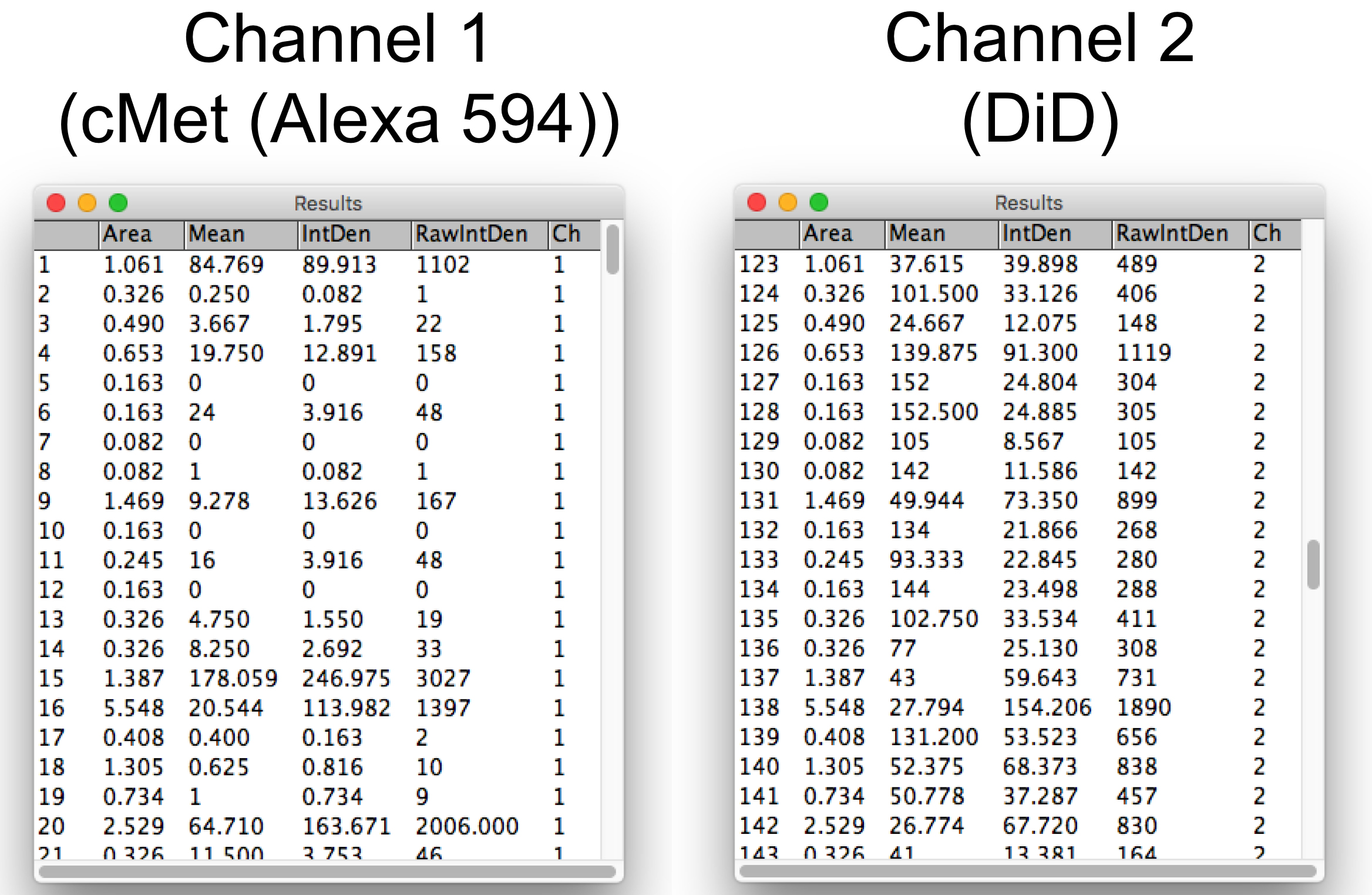
Figure 8. Results of “Area”, “Mean gray value (Mean)”, and “Integrated Density (IntDen)” of cargo (cMet-Alexa 594) and DiD in each exosome quantified by ImageJ. The total number of exosomes analyzed in this image is 122. - Transfer the data sets to Excel by copy and paste.
- Obtain “Mean gray value (Mean)” information of background of cargo and DiD channels respectively.
- Open the acquired images again.
- Select vesicle-free dark area with “Rectangular” or “Oval” tool.
- Go to “Analyze” → “Measure”, and get “Area”, “Mean gray value (Mean)”, and “Integrated Density (IntDen)” information of both cargo and DiD channels.
- Transfer the data sets to Excel with copy and paste.
- Calculate background of cargo and DiD of each extracellular vesicle respectively.
Calculating formula: Background = (“Mean gray value [Mean]” of background) * (“Area” of extracellular vesicles)
Note: If the calculating result becomes less than zero “0”, it would be fixed to zero “0”. This negative value means undetectable level of cargo or DiD. - Calculate cargo density of each extracellular vesicle.
Calculating formula:
Cargo density = [(“Integrated Density (IntDen)” of cargo) - (Background of cargo)]/[(“Integrated Density (IntDen)” of DiD) - (Background of DiD)] - Then make presentation diversely depending on the purpose of the study as in Kajimoto et al. (2013) or (2018).
Notes
- This protocol is applicable to all samples including adherent and non-adherent cells, plasma, and tissues. We used adherent cells as an example to describe the detailed protocol.
- The amount of extracellular vesicles varies in different cell types. In most cases, small-scale culture in 35 mm dishes or 6-well plates is enough, but in some cases, a large-scale culture in 60 mm or 100 mm dishes is required.
- If using the size- and density- dependent purification method for extracellular vesicles like differential centrifugation, it is important to use healthy cells for avoiding the contamination of apoptotic body or cell debris.
- Detailed conditions of Procedures C, D, and E are basically according to the previously reported fluorescence-based assay to measure single liposomes (Hatzakis et al., 2009).
Recipes
- Phosphate-buffered saline (PBS)
- Mix 1.37 M NaCl, 27 mM KCl, 100 mM Na2HPO4, and 18 mM KH2PO4 in 900 ml H2O
- Adjust to a final pH of 7.4
- Add H2O to 1,000 ml
- Autoclave to store (This is 10x PBS stock)
- Dilute 1:10 with H2O to make 1x PBS
- 4% PFA
- Prepare 30 ml of H2O in 50 ml conical tube and heat to 60 °C
- Add 1.6 g of PFA powder in the 30 ml of H2O, 60 °C in the fume hood and mix well
- Add 4 μl of 10 N NaOH and mix well until the solution gets clear
- Add 4 ml of 10x PBS
- Add H2O to 40 ml and mix
- Cool, store at 4 °C in dark condition, and use in two weeks
Acknowledgments
This protocol was adapted from procedures published in Kajimoto et al. (2013) and (2018). This work was supported by grants from the JSPS KAKENHI, the Uehara Memorial Foundation, the Osaka Medical Research Foundation, and the Nakatani Foundation.
Competing interests
There are no conflicts of interest.
References
- Hatzakis, N. S., Bhatia, V. K., Larsen, J., Madsen, K. L., Bolinger, P. Y., Kunding, A. H., Castillo, J., Gether, U., Hedegard, P. and Stamou, D. (2009). How curved membranes recruit amphipathic helices and protein anchoring motifs. Nat Chem Biol 5(11): 835-841.
- Kajimoto, T., Okada, T., Miya, S., Zhang, L. and Nakamura, S. (2013). Ongoing activation of sphingosine 1-phosphate receptors mediates maturation of exosomal multivesicular endosomes. Nat Commun 4: 2712.
- Kajimoto, T., Mohamed, N. N. I., Badawy, S. M. M., Matovelo, S. A., Hirase, M., Nakamura, S., Yoshida, D., Okada, T., Ijuin, T. and Nakamura, S. I. (2018). Involvement of Gβγ subunits of Gi protein coupled with S1P receptor on multivesicular endosomes in F-actin formation and cargo sorting into exosomes. J Biol Chem 293(1): 245-253.
- Karpman, D., Stahl, A. L. and Arvidsson, I. (2017). Extracellular vesicles in renal disease. Nat Rev Nephrol 13(9): 545-562.
- Maas, S. L. N., Breakefield, X. O. and Weaver, A. M. (2017). Extracellular vesicles: Unique intercellular delivery vehicles. Trends Cell Biol 27(3): 172-188.
- Trajkovic, K., Hsu, C., Chiantia, S., Rajendran, L., Wenzel, D., Wieland, F., Schwille, P., Brugger, B. and Simons, M. (2008). Ceramide triggers budding of exosome vesicles into multivesicular endosomes. Science 319(5867): 1244-1247.
- van Niel, G., D'Angelo, G. and Raposo, G. (2018). Shedding light on the cell biology of extracellular vesicles. Nat Rev Mol Cell Biol 19(4): 213-228.
Article Information
Copyright
© 2018 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Readers should cite both the Bio-protocol article and the original research article where this protocol was used:
- Kajimoto, T. and Nakamura, S. (2018). Quantitative Analysis of Cargo Density in Single-extracellular Vesicles by Imaging. Bio-protocol 8(24): e3111. DOI: 10.21769/BioProtoc.3111.
- Kajimoto, T., Mohamed, N. N. I., Badawy, S. M. M., Matovelo, S. A., Hirase, M., Nakamura, S., Yoshida, D., Okada, T., Ijuin, T. and Nakamura, S. I. (2018). Involvement of Gbetagamma subunits of Gi protein coupled with S1P receptor on multivesicular endosomes in F-actin formation and cargo sorting into exosomes. J Biol Chem 293(1): 245-253.
Category
Cell Biology > Cell imaging > Fluorescence
Cell Biology > Organelle isolation > Extracellular vesicle
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link








