- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Use of Gas Chromatography to Quantify Short Chain Fatty Acids in the Serum, Colonic Luminal Content and Feces of Mice
Published: Vol 8, Iss 22, Nov 20, 2018 DOI: 10.21769/BioProtoc.3089 Views: 14635
Reviewed by: David CisnerosManish Kumar PatelSesha Lakshmi Arathi Paluri

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

Visualizing Hypoxia in a Murine Model of Candida albicans Infection Using in vivo Biofluorencence
José Pedro Lopes and Constantin F. Urban
Aug 5, 2019 5143 Views
RNA Extraction from Ears and Draining Lymph Nodes of Mice Infected with Leishmania amazonensis
Emilie Giraud and Evie Melanitou
Jun 5, 2020 5546 Views

TetR Regulated in vivo Repression Technology to Identify Conditional Gene Silencing in Genetically Engineerable Bacteria Using Vibrio cholerae Murine Infections as Model System
Franz G. Zingl [...] Stefan Schild
Oct 5, 2020 3779 Views
Abstract
Short-Chain Fatty Acids (SCFAs) are a product of the fermentation of resistant starches and dietary fibers by the gut microbiota. The most important SCFA are acetate (C2), propionate (C3) and butyrate (C4). These metabolites are formed and absorbed in the colon and then transported through the hepatic vein to the liver. SCFAs are more concentrated in the intestinal lumen than in the serum. Butyrate is largely consumed in the gut epithelium, propionate in the liver and acetate in the periphery. SCFAs act on many cells including components of the immune system and epithelial cells by two main mechanisms: activation of G-protein coupled receptors (GPCRs) and inhibition of histone deacetylase. Considering the association between changes in SCFA concentrations and the development of diseases, methods to quantify these acids in different biological samples are important. In this study, we describe a protocol using gas chromatography to quantify SCFAs in the serum, feces and colonic luminal content. Separation of compounds was performed using a DB-23 column (60 m x 0.25 mm internal diameter [i.d.]) coated with a 0.15 µm thick layer of 80.2% 1-methylnaphatalene. This method has a good linear range (15-10,000 µg/ml). The precision (relative standard deviation [RSD]) is less than 15.0% and the accuracy (error relative [ER]) is within ± 15.0%. The extraction efficiency was higher than 97.0%. Therefore, this is cost effective and reproducible method for SCFA measurement in feces and serum.
Keywords: Short chain fatty acidBackground
The microbiota is a complex, heterogeneous and dynamic group of microorganisms (bacteria, viruses and fungi) that colonizes the tissues in direct contact with the external environment including the skin and the genitourinary, intestinal and respiratory tracts. Several studies have demonstrated recently that these microorganisms are important for maintaining host homeostasis. Indeed, changes in their composition can lead to dysbiosis, as observed in specific pathological conditions, and have been associated with their development, as shown for inflammatory bowel disease, asthma, obesity and other chronic inflammatory conditions (Ferreira et al., 2014; Nishida et al., 2018; Sokolowska et al., 2018).
Short chain fatty acids are bacterial metabolites produced by components of the microbiota. The most abundant and studied molecules of this class are acetic, propionic and butyric acids, which are normally found in their deprotonated forms (e.g., acetate, propionate and butyrate). SCFAs modify important processes including metabolism, immune system development and activation, and intestinal barrier function strategies (den Besten et al., 2013; Correa-Fiz et al., 2016; Koh et al., 2016) have been used for changing SCFAs bioavailability in the gut including administration of probiotics (Andrade-Oliveira et al., 2015; Mendes et al., 2017), antibiotics (Fellows et al., 2018), diets with different fiber contents (Vieira et al., 2017), SCFAs in the drinking water or the butyric acid pro-drug (tributyrin) (Vinolo et al., 2012; Vieira et al., 2017). Taken together, these strategies allow us to investigate the role of these metabolites in vivo in different disease models. In this context, it is essential to have a robust method for measuring the levels of these molecules in biological samples such as feces, luminal content or serum.
Gas chromatography (GC) is most commonly used for SCFA analysis due to compatibility with the chemical properties of SCFAs, such as volatility, and the suitability of the detectors that can be coupled to this equipment, such as flame ionization detector (FID). FID is the most widely used for analysis of SCFAs. Due to the large amount of compounds present in the matrix, the sample should be pretreated using extraction and derivatization procedures. Recently, several techniques of derivatization have been described to obtain more stable compounds and provide greater compatibility between the stationary phase and the analytes (Karlsson et al., 2010; Walton et al., 2012; Zhang et al., 2013). On the other hand, the use of derivation techniques has critical disadvantages, including the points that it is time-consuming, there can be losses of SCFAs, it is costly due to the use of large quantities of reagents, and the risk of occupational exposure to allergenic and toxic reagents. Some authors have described filtration and ultracentrifugation techniques to avoid the disadvantages of derivatization and obtain a fast sample preparation (Cuervo et al., 2013; Salazar et al., 2015). In spite of this, the run time is usually increased due to low purification of the samples, which increases the quantities and variety of compounds needed for separation in the column. We have chosen the ultracentrifugation technique in this protocol. Here, we describe a method that we have used to measure SCFAs in biological samples using gas chromatography equipped with FID and a liquid-liquid extraction technique. The column used in this method is composed of a high polarity stationary phase, which makes analysis of the SCFAs possible. The liquid-liquid extraction offers some advantages such as greater sample recuperation, higher sample purification, time optimization and greater separation and peak resolution in chromatograms.
Materials and Reagents
- 100-1,000 µl pipette tips (KASVI, catalog number: K8-1000B)
- Eppendorf Safe-Lock Tubes, 1.5 ml (Eppendorf, catalog number: 0030120086)
- Amber, write-on spot, certified, 2 ml, screw top vial packs (Agilent Technologies, catalog number: 5182-0554)
- Serum, colonic luminal content and feces of C57BL/6 mice
- Hydrochloric acid, 37% (v/v) (Sigma-Aldrich, catalog number: 320331)
- Anhydrous citric acid (Cinética Produtos Químicos, catalog number: 278)
- Sodium chloride (Cinética Produtos Químicos, catalog number: 415)
- Tetrahydrofuran (Merck, catalog number: 1081012500)
- Acetonitrile (Sigma-Aldrich, catalog number: 60004)
- N-butanol (Sigma-Aldrich, catalog number: 34867)
- Acetic acid, analytical standard, GC assay ≥ 9.8% (Sigma-Aldrich, catalog number: 71251)
- Propionic acid, analytical standard, GC assay ≥ 9.8% (Sigma-Aldrich, catalog number: 94425)
- Butyric acid, analytical standard, GC assay ≥ 9.8% (Sigma-Aldrich, catalog number: 19215)
- HCl
- Distilled water
- 1-methylnaphthalene
- N2
- H2
- 0.1 M HCl solution (see Recipes)
- 3,000 µg/ml acetic acid in serum stock solution (see Recipes)
- 3,000 µg/ml propionic acid stock solution (see Recipes)
- 3,000 µg/ml butyric acid stock solution (see Recipes)
- 1,000 µg/ml SCFAs standard mixture (see Recipes)
Equipment
- J&W DB-23 GC column, 60 m, 0.32 mm, 0.25 µm, 7 inch cage (Agilent Technologies, catalog number: 123-2362)
- Forceps
- Freezer FE26/127V (Electrolux Appliances, model: FE26, catalog number: 04251FBA106/ 04251FBB206)
- Stainless steel double spatula, 180 mm length x 3 mm diameter (Metalic Acessórios para Laboratório, catalog number: 063-A3)
- PIPETMAN Classic P1000 pipette (Gilson, catalog number: 20170-170)
- Gas Chromatograph Agilent 6850 series (Agilent Technologies, discontinued by the manufacturer)
- DB 23 Agilent capillary column, 60 m x 0.25 mm internal diameter (i.d.)
- Oven
- Automatic liquid sampler 7683B (Agilent Technologies, catalog number: G2880A)
- Analytical balance (SHIMADZU, model: ATX224 series, catalog number not found)
- Vortex mixer (Phenix Luferco, model: AP56)
- Centrifuge 5810 R (Eppendorf, model: 5810 R, catalog number: 5810000424)
- Gas Chromatograph Shimadzu 2010 model, equipped with FID (SHIMADZU, catalog number: C184-E019)
- AOC-20i automatic liquid sampler (SHIMADZU, model: AOC-20i)
Software
- EZChrom software, 3.3.1 version (Agilent Technologies)
- GC solution software, 3.2 version (SHIMADZU)
Procedure
- Sample preparation
- Collect blood from cardiac puncture or axillary plexus. Maintain blood at room temperature (RT) for 30 min and centrifuge (3,000 x g, 8 min). Collect the serum and freeze at -80 °C in 1.5 ml microtubes.
- Collect fecal pellets directly from each mouse in microtubes. For this, raise the animal holding the tail, position the microtube in the proximity of the anus and collect the pellets that are excreted. Two or three pellets from each animal are adequate.
- After euthanasia, collect the colons and gently remove the luminal contents of the proximal part. To do this, use the forceps to collect directly into microtubes and freeze at -80 °C.
- Keep all biological samples frozen at -80 °C until the day of analysis.
- For feces and colonic luminal content, transfer 20 mg of these into 1.5 ml microtubes and add 200 µl of distilled water. Homogenize using a metal spatula.
- In 1.5 ml microtubes, add the sample (200 µl serum or homogenate of feces with water, see Step A5), a 200 µl mixture of organic solvents composed of N-butanol, tetrahydrofuran and acetonitrile in a 50:30:20 ratio, 40 µl HCl 0.1 M, 20 mg citric acid and 40 mg sodium chloride. Shake the microtubes vigorously using the vortex stirrer for 1 min.
- Centrifuge the samples at 14,870 x g at room temperature for 10 min.
- Using an automatic pipette, transfer the supernatant to chromatographic vials equipped with 200 µl inserts and analyze by GC-FID (Figure 1).

Figure 1. Sample preparation. A. Liquid-liquid extraction procedure of samples. Serum samples (200 µl) were extracted with the same volume of a mixture of solvents. B. Liquid-liquid extraction procedure for feces and colonic luminal content samples. Fecal samples (20 mg) were previously homogenized with distilled water and subjected to the extraction procedures described in Steps A5-A8 of Sample preparation section.
- GC procedure
- Adjust the temperature of the injector to 250 °C.
- Inject contents (5 µl) in a split ratio of 25:1, using the DB 23 capillary column Agilent, 60 m x 0.25 mm of internal diameter (i.d.), and coated with a film of 0.15 µm composed of 80.2% 1-methylnaphthalene.
- The mobile phase is composed of N2 at an initial flow rate of 1 ml/min and maintaining this for 1 min, then changing to 0.8 ml/min for 1 min, changing to 0.6 ml/min for 1 min and then reverting to 1 ml/min for 9.2 min.
- Adjust and maintain the temperature of the FID detector at 260 °C.
- Adjust the flow of the H2 and the synthetic air to 30 ml/min and 350 ml/min, respectively.
- Adjust the temperature program of the oven to100 °C, maintain for 7 min and increase to 200 °C at a rate of 25 °C/min and maintain for 5 min.
Data analysis
- Chromatograms generated by the analysis of the serum (Figure 1A) and the feces (Figure 1B) are shown in Figure 2. SCFAs are identified by the times of retention in comparison with analytical standard solutions.

Figure 2. Chromatogram of serum (A) and fecal (B) samples. The retention times of each SCFA are as follows: acetate, 5.2 min; propionate, 7.1 min; butyrate, 9.08 min. The analysis is performed using a Gas Chromatograph Shimadzu 2010 model equipped with 7683B automatic liquid sampler and FID. - Use the EZChrom software to integrate the areas of each peak, corresponding to its respective SCFA.
- Prepare standard calibration curves in triplicate, with a concentration range of 15-1,000 µg/ml for SCFAs using the matrices in this study (serum and feces) as solvent to dilute the analytes. Construct the calibration curve in serum matrix, make a 1,000 µg/ml stock solution of SCFA using serum as solvent, and dilute serially in a range of 15-1,000 µg/ml (see Recipes).
- Construct the calibration curve in feces matrix, make a 1000 µl/ml SCFAs using distilled water as solvent in accordance with obtaining method of stock solution of SCFA in serum (see Recipes).
- Prepare samples according to the liquid-liquid extraction procedures described in the Procedure A, Steps 5-8 and analyze the samples by GC-FID as described in the Procedure B, GC procedure.
The standard calibration curves have presented linear regression relationships for all acids (R2 = 0.998) (Figure 3).
Figure 3. Linearity of standard calibration curves of (A) serum and (B) feces. Calibration curves obtained using serum (A) and feces (B). Linearity obtained using Gas Chromatograph Agilent 6850 series, equipped with 7683B automatic liquid sampler and FID following the parameters described in this article. - The retention time is 7.2, 8.2, and 9.2 min for acetic, propionic and butyric acid, respectively and the total run time for each analysis is 16.5 min.
- The precision, expressed as relative standard deviation (RSD) was less than 15.0%, and the accuracy, expressed as Relative Error (RE) was within ± 15.0%. The extraction efficiency (EE) was greater than 97.0%. To calculate the RSD, RE and EE, use the formulas described below:

- For standard curves, plot areas of each SCFA standard sample on the y-axis against your respective theoretical concentration values (15-1,000 µg/ml) and obtain the equation of the function y = ax + b, where a is the angular coefficient, x is the area found for the sample and b is the linear coefficient.
- Use the equation showed below to calculate the concentrations of each SCFA in the samples. The concentration of each SCFA is expressed as mg/ml.

- Convert the concentrations for mM and mmol/kg for the serum and fecal/colonic luminal content samples, respectively using the formulas described below:

Notes
- Agilent equipment is not available in our laboratory. Thus, we have used the GC Shimadzu 2010 model to obtain the chromatograms to illustrate analytes separation (Figures 2 and 4). The chromatograms were obtained following the bioanalytical method conditions described in this article.
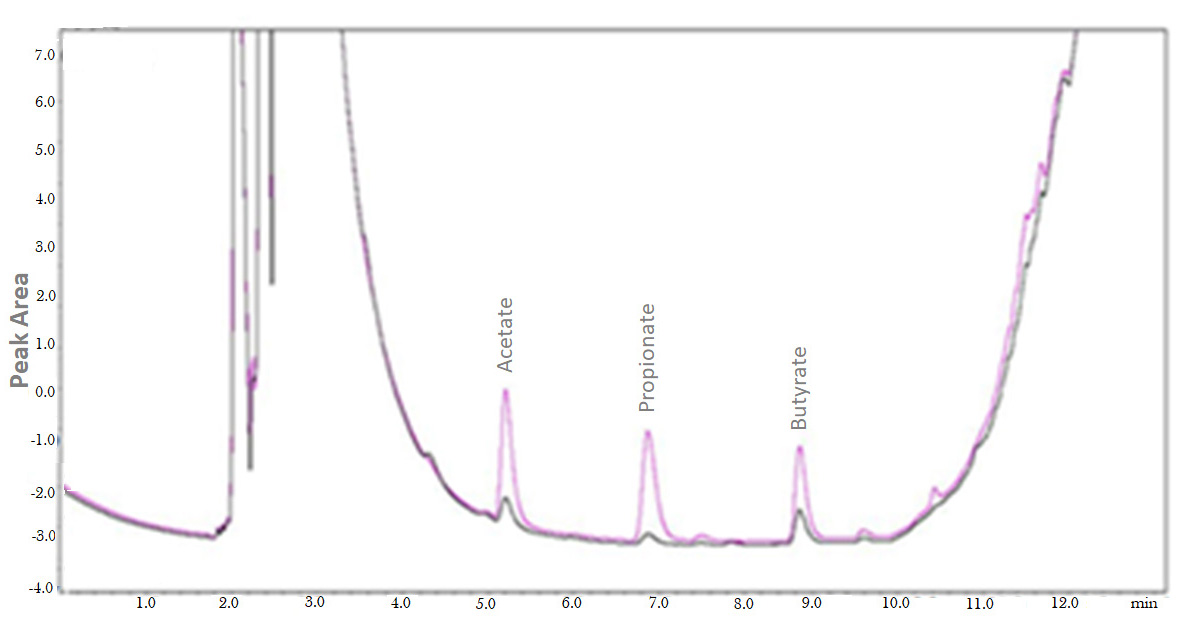
Figure 4. Overlap of chromatograms obtained of colonic luminal content from antibiotic-treated mice and control animals (not treated with antibiotics). The chromatogram shows animals treated with antibiotics (blue line) and untreated animals (black line) (Fellows et al., 2018). The analyses are performed using a Gas Chromatograph Shimadzu 2010 model equipped with AOC-20i automatic liquid sampler and FID, in accordance with the parameters described in this article. The chromatograms were integrated using the GC solution software. - The retention time and the detection range of the analytes may vary according to capillary column type and length and equipment brands. DB-23 capillary column Agilent can be used in GC Shimadzu 2010 model.
- This method was developed and validated in accordance with the RDC 27/2012 of ANVISA (National Sanitary Vigilance Agency).
- No alterations to the protocol are required if different mouse strains are used.
- Animals were housed in facility of the Institute of Biomedical Sciences, University of Sao Paulo and Institute of Biology, University of Campinas.
- Blank serum is the serum without SCFAs standards addition.
Recipes
- 0.1 M HCl solution
0.835 ml HCl 37% (v/v)
Add 99.165 ml distilled water - 3,000 µg/ml acetic acid in serum stock solution
5.7 µl analytical standard acetic acid, GC assay ≥ 99.8%
Add 2,000 µl blank serum
Vortex for 1 min - 3,000 µg/ml propionic acid stock solution
6.06 µl analytical standard propionic acid, GC assay ≥ 99.8%
Add 2,000 µl blank serum
Vortex for 1 min - 3,000 µg/ml butyric acid stock solution
6.2 µl analytical standard butyric acid, GC assay 99.8%
Add 2,000 µl blank serum
Vortex for 1 min - 1,000 µg/ml SCFAs standard mixture
Mix 1 ml each SCFA stock solution to give 1,000 µg/ml each SCFA (final volume 3 ml)
Standard SCFA solutions- Perform a serial dilution of this mixed organic acid solution using serum as solvent to generate a range of 15-1,000 µg/ml as a standard curve of SCFAs in serum matrix
- Perform the same serial dilution using distilled water as matrix for the construction of the standard curve of SCFAs in feces matrix. To carry this out, prepare seven tubes containing 20 mg of feces and add 200 µl of their respectively solutions obtained in the serial dilution in each tube
Acknowledgments
We are grateful to Claudete Justina Valduga and Patrícia Sartorelli for providing equipment for this study. This work was supported by FAPESP 2012/50410-8 and CNPq (The National Council for Scientific and Technological Development), project number 486037/2012-6 to C.M.F, and 2017/10653-9 to M.A.V.
Competing interests
The authors have declared that no conflict of interest exists.
Ethics
All experimental procedures were approved by the Ethics Committee on Animal Use of the Institute of Biology, University of Campinas (protocol number 3742-1), and Animal Care Committee of the University of Sao Paulo (protocol number. 181/114/02). Also, it was performed in accordance with the guidelines of the Brazilian Control and Experimentation Committee.
References
- Andrade-Oliveira, V., Amano, M. T., Correa-Costa, M., Castoldi, A., Felizardo, R. J., de Almeida, D. C., Bassi, E. J., Moraes-Vieira, P. M., Hiyane, M. I., Rodas, A. C., Peron, J. P., Aguiar, C. F., Reis, M. A., Ribeiro, W. R., Valduga, C. J., Curi, R., Vinolo, M. A., Ferreira, C. M. andCamara, N. O. (2015). Gut bacteria products prevent AKI induced by ischemia-reperfusion. J Am Soc Nephrol 26(8): 1877-1888.
- Correa-Fiz, F., Fraile, L. and Aragon, V. (2016). Piglet nasal microbiota at weaning may influence the development of Glasser's disease during the rearing period. BMC Genomics 17: 404.
- Cuervo, A., Salazar, N., Ruas-Madiedo, P., Gueimonde, M. and Gonzalez, S. (2013). Fiber from a regular diet is directly associated with fecal short-chain fatty acid concentrations in the elderly. Nutr Res 33(10): 811-816.
- den Besten, G., van Eunen, K., Groen, A. K., Venema, K., Reijngoud, D. J. and Bakker, B. M. (2013). The role of short-chain fatty acids in the interplay between diet, gut microbiota, and host energy metabolism. J Lipid Res 54(9): 2325-2340.
- Fellows, R., Denizot, J., Stellato, C., Cuomo, A., Jain, P., Stoyanova, E., Balazsi, S., Hajnady, Z., Liebert, A., Kazakevych, J., Blackburn, H., Correa, R. O., Fachi, J. L., Sato, F. T., Ribeiro, W. R., Ferreira, C. M., Peree, H., Spagnuolo, M., Mattiuz, R., Matolcsi, C., Guedes, J., Clark, J., Veldhoen, M., Bonaldi, T., Vinolo, M. A. R. and Varga-Weisz, P. (2018). Microbiota derived short chain fatty acids promote histone crotonylation in the colon through histone deacetylases. Nat Commun 9(1): 105.
- Ferreira, C. M., Vieira, A. T., Vinolo, M. A., Oliveira, F. A., Curi, R. and Martins Fdos, S. (2014). The central role of the gut microbiota in chronic inflammatory diseases. J Immunol Res 2014: 689492.
- Karlsson, C., Ahrne, S., Molin, G., Berggren, A., Palmquist, I., Fredrikson, G. N. and Jeppsson, B. (2010). Probiotic therapy to men with incipient arteriosclerosis initiates increased bacterial diversity in colon: a randomized controlled trial. Atherosclerosis 208(1): 228-233.
- Koh, A., De Vadder, F., Kovatcheva-Datchary, P. and Backhed, F. (2016). From dietary fiber to host physiology: Short-chain fatty acids as key bacterial metabolites. Cell 165(6): 1332-1345.
- Mendes, E., Acetturi, B. G., Thomas, A. M., Martins, F. D. S., Crisma, A. R., Murata, G., Braga, T. T., Camara, N. O. S., Franco, A., Setubal, J. C., Ribeiro, W. R., Valduga, C. J., Curi, R., Dias-Neto, E., Tavares-de-Lima, W. and Ferreira, C. M. (2017). Prophylactic supplementation of Bifidobacterium longum 51Aprotects mice from ovariectomy-induced exacerbated allergic airway inflammation and airway hyperresponsiveness. Front Microbiol 8: 1732.
- Nishida, A., Inoue, R., Inatomi, O., Bamba, S., Naito, Y. and Andoh, A. (2018). Gut microbiota in the pathogenesis of inflammatory bowel disease. Clin J Gastroenterol 11(1): 1-10.
- Salazar, N., Dewulf, E. M., Neyrinck, A. M., Bindels, L. B., Cani, P. D., Mahillon, J., de Vos, W. M., Thissen, J. P., Gueimonde, M., de Los Reyes-Gavilan, C. G. and Delzenne, N. M. (2015). Inulin-type fructans modulate intestinal Bifidobacterium species populations and decrease fecal short-chain fatty acids in obese women. Clin Nutr 34(3): 501-507.
- Sokolowska, M., Frei, R., Lunjani, N., Akdis, C. A. and O'Mahony, L. (2018). Microbiome and asthma. Asthma Res Pract 4: 1.
- Vieira, A. T., Castelo, P. M., Ribeiro, D. A. and Ferreira, C. M. (2017). Influence of oral and gut microbiota in the health of menopausal women. Front Microbiol 8: 1884.
- Vinolo, M. A., Rodrigues, H. G., Festuccia, W. T., Crisma, A. R., Alves, V. S., Martins, A. R., Amaral, C. L., Fiamoncini, J., Hirabara, S. M., Sato, F. T., Fock, R. A., Malheiros, G., dos Santos, M. F. and Curi, R. (2012). Tributyrin attenuates obesity-associated inflammation and insulin resistance in high-fat-fed mice. Am J Physiol Endocrinol Metab 303(2): E272-282.
- Walton, G. E., Lu, C., Trogh, I., Arnaut, F. and Gibson, G. R. (2012). A randomised, double-blind, placebo controlled cross-over study to determine the gastrointestinal effects of consumption of arabinoxylan-oligosaccharides enriched bread in healthy volunteers. Nutr J 11: 36.
- Zhang, H., Sun, J., Liu, X., Hong, C., Zhu, Y., Liu, A., Li, S., Guo, H. and Ren, F. (2013). Lactobacillus paracasei subsp. paracasei LC01 positively modulates intestinal microflora in healthy young adults. J Microbiol 51(6): 777-782.
Article Information
Copyright
© 2018 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Ribeiro, W. R., Vinolo, M. A. R., Calixto, L. A. and Ferreira, C. M. (2018). Use of Gas Chromatography to Quantify Short Chain Fatty Acids in the Serum, Colonic Luminal Content and Feces of Mice. Bio-protocol 8(22): e3089. DOI: 10.21769/BioProtoc.3089.
Category
Microbiology > Microbe-host interactions > In vivo model > Mammal
Immunology > Animal model > Mouse
Systems Biology > Metabolomics > Whole organism
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link











