- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Measurement of Cytokines
Published: Vol 2, Iss 22, Nov 20, 2012 DOI: 10.21769/BioProtoc.290 Views: 16569

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
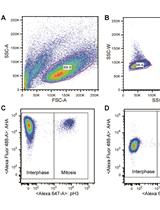
Measuring Protein Synthesis during Cell Cycle by Azidohomoalanine (AHA) Labeling and Flow Cytometric Analysis
Koshi Imami and Tomoharu Yasuda
Apr 20, 2019 9799 Views
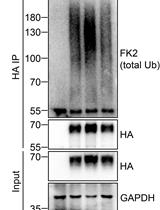
Analysis of the Ubiquitination and Phosphorylation of Vangl Proteins
Di Feng [...] Bo Gao
Oct 20, 2022 3472 Views
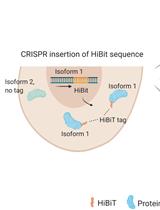
Isoform-specific, Semi-quantitative Determination of Highly Homologous Protein Levels via CRISPR-Cas9-mediated HiBiT Tagging
Kristina Seiler [...] Mario P. Tschan
Jul 20, 2023 2555 Views
Abstract
This protocol allows to measure the levels cytokines - such as VEGFs, CXCLs cytokines, PDGF or FGF - from fresh samples but also frozen tumors. The advantage of this method is to use very few micrograms of biological material and the protocol is carried out quickly.
Keywords: CytokinesMaterials and Reagents
- Fresh or frozen tumor tissues stored at - 80 °C
- Extraction buffer (Life Technologies, catalog number: FNN0011 )
- Antifoam Y-30 Emulsion (Sigma-Aldrich, catalog number: A5758 )
- BCA protein quantification kit (Interchim, catalog number: MP2920 )
- ELISA kits (Peprotech or R&D System)
- Tween-20 (Sigma-Aldrich, catalog number: P-7949 )
- BSA (Sigma-Aldrich, catalog number: A-7030 )
- Avidin-HRP conjugate solution (Sigma-Aldrich, catalog number: A-7419 )
- 10x Dulbecco’s PBS (Life Technologies, Gibco®, catalog number: 14200-075 )
- ABTS Liquid substrate solution (Sigma-Aldrich, catalog number: A3219 )
Equipment
- Homogenizer such as Precellys (Ozyme BER1011S, France) or ultraturax
- Centrifuge
- 96 wells plates (DUTSCHER SCIENTIFIC, catalog number: 047632 )
- ELISA microplates (Nunc MaxiSorp, catalog number: 439454 )
- Luminoskan (Thermo Fisher Scientific, catalog number: 5210470 )
Procedure
- 20 mg of fresh or frozen tumor tissues were resuspended in 200 μl of extraction buffer at 4 °C.
- Tissues were mechanically ground using a homogenizer such as an ultraturax or a Precellys. With ultraturax an antifoam solution as described by the manufacturer (Sigma-Aldrich – antifoam Y-30 Emulsion) is used.
- The homogenate was centrifuged for 10 min at 6,000 rpm, at 4 °C.
- The supernatant was recovered. 10 μl supernatant is used to determinate the protein concentration using a protein assay such as BCA. The sample can be stored at -80 °C.
- Measurement of cytokines must be performed as described by ELISA kit manufacturer.
- Dilute capture antibody with PBS to a concentration of 1 μg/ml and add 100 μl to each ELISA pate well. Seal the plate and incubate overnight at room temperature.
- Invert plate to remove capture antibody and blot on paper towel and wash plate 3 times by adding 300 μl of wash solution (1x PBS – 0.05% tween-20). Invert plate to remove wash solution and blot on paper towel.
- Block plate by adding 300 μl per well of 1% BSA in 1x PBS. Incubate 1 h at room temperature.
- Invert plate to remove blocking buffer and wash plate 3 times as described in step 7.
- Achieve an eight point standard curve using 2-fold serial dilutions of standard, in extraction diluent solution (1x PBS – 0.05% Tween-20 – 0.1% BSA), and a high standard of 2,000 μg/μl is recommended.
- Add 100 μl of standard or sample to each well in triplicate. Incubate at room temperature for at least 2 h.
- Invert plate to remove samples and standard and wash plate 3 times as described in step 7.
- Dilute biotinylated detection antibody in diluent (1x PBS - 0.05% Tween-20 - 0.1% BSA) to a concentration of 500 ng/ml and add 100 μl per well. Incubate at room temperature for 2 h.
- Invert plate to remove detection antibody and wash plate 3 times as described in step 7.
- Add 100 μl per well of Avidin-HRP conjugate diluted 1: 2,000 in diluent (1x PBS – 0.05% tween-20 - 0.1% BSA). Incubate 30 min at room temperature.
- Invert plate to remove Avidin-HRP conjugate and wash plate 3 times as described in step 7.
- Add 100 μl of ABTS liquid subtrate to each well. Incubate at room temperature, 10 min, for color development.
- Determine the optical density of each well, using a microplate reader set to 405 nm.
- Average the triplicate readings for each standard, control, and sample and subtract the average zero standard optical density.
- Create a standard curve by plotting the mean absorbance for each standard on the y-axis against the concentration on the x-axis and draw a curve through the points on the graph.
Acknowledgments
Our protocol was adapted from the following article: Grepin et al. (2012). We thank Dr Brahimi-Horn for editorial assistance. Financial support: Contract VEGFIL from the National Institute of Cancer (INCA), the French Association for Cancer Research (ARC), the Foundation de France, the “Conseil Général des Alpes Maritimes”, Roche France and The “Association pour la Recherche sur les Tumeurs du Rein (ARTuR)”.
References
- Grepin, R., Guyot, M., Jacquin, M., Durivault, J., Chamorey, E., Sudaka, A., Serdjebi, C., Lacarelle, B., Scoazec, J. Y., Negrier, S., Simonnet, H. and Pages, G. (2012). Acceleration of clear cell renal cell carcinoma growth in mice following bevacizumab/Avastin treatment: the role of CXCL cytokines. Oncogene 31(13): 1683-1694.
Article Information
Copyright
© 2012 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Grépin, R. and Pagès, G. (2012). Measurement of Cytokines. Bio-protocol 2(22): e290. DOI: 10.21769/BioProtoc.290.
Category
Cancer Biology > General technique > Biochemical assays > Protein analysis
Cancer Biology > Inflammation > Biochemical assays > Protein analysis
Biochemistry > Protein > Quantification
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link









