- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
KMnO4 Footprinting
Published: Vol 2, Iss 21, Nov 5, 2012 DOI: 10.21769/BioProtoc.280 Views: 27803

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
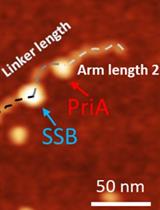
Characterize the Interaction of the DNA Helicase PriA with the Stalled DNA Replication Fork Using Atomic Force Microscopy
Yaqing Wang [...] Yuri L. Lyubchenko
Mar 5, 2021 4888 Views
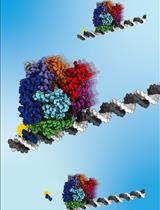
A Gel-Based Assay for Probing Protein Translocation on dsDNA
Christiane Brugger and Alexandra M. Deaconescu
Jul 20, 2021 4373 Views
Abstract
The KMnO4 footprinting method offers a rapid and easy way to detect and localize single-stranded regions within a duplex DNA molecule, such as it occurs for instance within an actively transcribing RNA polymerase-DNA complex or during R-loop formation in DNA-RNA hybrid structures. The method is based on the selective oxidation of single-stranded thymines in DNA. The modified nucleotides react with strong bases by ring opening and subsequent phosphodiester cleavage. Because the modified nucleotides will not be recognized by DNA polymerase sites of modification can also be analyzed by primer extension with Klenow DNA polymerase, which stops elongation one residue before the modification. Hence, localization of the modified base positions can be performed on denaturing polyacrylamide gels either after piperidine catalyzed phosphodiester cleavage of 3'- or 5'-32P-end-labeled DNA or by primer extension with non-labeled DNA employing 32P-labeled oligonucleotide primers. Due to the fact that KMnO4 can penetrate through membranes the footprinting method can also be used for footprint analyses within living cells.
Keywords: Single-stranded DNA localizationMaterials and Reagents
- Radiolabeled DNA fragment of interest
- 14.3 M β-mercaptoethanol
- 500 mM EDTA
- Phenol
- Bromophenol blue (Sigma-Aldrich, catalog number: BO126 )
- Xylene cyanol (Sigma-Aldrich, catalog number: X4126 )
- Formamide deionized (Panreac Applichem, catalog number: A2156 )
- Cholorophorm
- Piperidine, purity grade: pro analysis (p.a.) (e.g. Sigma-Aldrich, catalog number: 411027 )
- Ethanol (purity grade: pro analysis) (p.a.)
- X-ray films
- Glycogen (1 μg/μl) (e.g. Roche, catalog number: 10901393001 )
- NaOAc (300 mM pH = 5.5)
Optional for Procedure B (primer extension analysis) - Non-radio labeled DNA fragment of interest
- 5'-32P-labeled desoxyoligonucleotide primer
- NaOH (10 mM)
- Tris
- MgSO4
- DTT
- Klenow fragment of DNA polymerase I (e.g. Biolabs, catalog number: MO210S )
- NH4OAc
- 370 mM KMnO4 stock solution (a 1:1 dilution with H2O is used for the reaction) (see Recipes)
- Phenol/Chloroform (see Recipes)
- Neutralization solution (see Recipes)
- dNTP mix (see Recipes)
- Stop mix (see Recipes)
- Electrophoresis loading buffer (see Recipes)
Equipment
- Table-top centrifuge
- Vortex shaker
- Incubator
- Polyacrylamide gel electrophoresis system
- Vacuum concentrator
Procedure
- Footprinting with 3'- or 5'-end-labeled DNA analyzed by piperidine-catalyzed strand scission.
- Add 4 μl of 160 mM KMnO4 solution to radiolabeled DNA fragments of interest (40 ng; 5,000 to 10,000 cpm) (in the presence or absence of a binding partner) in a total volume of 40 μl.
- Incubate samples for 2 min at 30 °C; mix gently.
- Stop reaction by the addition of 4.8 μl β-mercaptoethanol; mix again.
- Put samples on ice.
- Add 5.3 μl 500 mM EDTA.
- Extract samples three times with 100 μl phenol/chloroform by vigorous shaking on a vortex for 1 min each time.
- Centrifuge samples (5 min, 3,000 rpm) and take aqueous layer.
- Precipitate samples with 2.5 volumes absolute ethanol (-20 °C).
- Centrifuge tubes (10 min, 10,000 rpm, 4 °C).
- Dissolve wet ethanol pellets in 70 μl of 10% (v/v) piperidine.
- Incubate samples at 90 °C for 30 min.
- Lyophilize samples in a vacuum concentrator until dry.
- Add 30 μl distilled water and lyophilize again.
- Repeat step 9.
- Dissolve final pellets in 50 μl distilled water.
- Add 4 μl 300 mM NaOAc (pH 5.5).
- Add 1 μl glycogen (1 μg/μl) to aid precipitation.
- Precipitate samples with 2.5 volumes absolute ethanol (-20 °C).
- Wash pellets with 150 μl 70% ethanol.
- Dry samples briefly in a vacuum concentrator to get rid of excess ethanol and dissolve in 5 μl electrophoresis loading buffer.
- Separate on a 10% or 15 % denaturing polyacrylamide gel, respectively, depending on the size of the DNA fragment.
- Visualize radioactive bands by autoradiography (over-night X-ray exposure).
- Add 4 μl of 160 mM KMnO4 solution to radiolabeled DNA fragments of interest (40 ng; 5,000 to 10,000 cpm) (in the presence or absence of a binding partner) in a total volume of 40 μl.
- Footprinting with non-labeled DNA analyzed by primer extension
- Add 4 μl of 160 mM KMnO4 solution to 40 μl non-labeled DNA fragments (~100 ng) in presence or absence of a binding partner.
- Follow steps 2 to 8 of procedure A.
- Dissolve pellet in 35 μl distilled water.
- Add 1 μl of an appropriate 15 to 20mer 5'-32P-labeled desoxyoligonucleotide primer complementary to the downstream sequence of interest (~ 5 x 105 cpm). For convenient resolution the primer should not bind farther than 50 to 80 nucleotides downstream to the region of interest.
- Add 4 μl 10 mM NaOH.
- Incubate for 2 min at 80 °C.
- Put samples on ice for 5 min.
- Add 4.5 μl of neutralization solution and mix samples.
- Incubate for 3 min at 68 °C for hybridization.
- Put samples on ice and add 5 μl of all four dNTPs (5 mM each), 1 unit Klenow fragment of DNA polymerase I and mix gently.
- Extension reaction is carried out for 10 min at 50 °C.
- Put samples on ice and terminate the reaction by the addition of 17 μl stop mix.
- Samples are precipitated with 2.5 volumes ethanol (-20 °C).
- Wash pellets with 150 μl 70% ethanol.
- Dry samples briefly in a vacuum concentrator to get rid of excess ethanol and dissolve in 5 μl electrophoresis loading buffer.
- Separate on a 10% or 15% denaturing polyacrylamide gel, respectively, depending on the size of the cDNA fragments expected.
- Visualize radioactive bands by autoradiography (over-night X-ray exposure).
- Add 4 μl of 160 mM KMnO4 solution to 40 μl non-labeled DNA fragments (~100 ng) in presence or absence of a binding partner.
Recipes
- KMnO4 stock solution (370 mM)
KMnO4 (MW = 158.04) has a solubility limit of ~60 g/L (corresponding to 0.37 M).
A 200 ml stock solution is prepared by adding 12 g KMnO4 to 220 ml distilled water. The solution is boiled until it reaches a final volume of 200 ml. This stock solution can be kept in a dark bottle for several months. - Phenol/Chloroform
A 1:1 mixture (v/v) of phenol and chloroform, purity grade: pro analysis (p.a.) is used for extraction. The phenol has been saturated with 1 M Tris-HCl (pH 7.9) before and 0.1% (final concentration) 8-Hydroxychinolin should be added to the phenol to avoid oxidation. - Neutralization solution
0.5 M Tris-HCl (pH 7.2)
0.1 M MgSO4
2 mM DTT (optional for procedure B) - dNTP mix
5 mM each, dATP, dCTP, dGTP, dTTP (optional for procedure B) - Stop mix
4 M NH4OAc
20 mM EDTA (optional for procedure B) - Electrophoresis loading buffer
0.1% (w/v) bromophenol blue
0.1% xylene cyanol
95% formamide deionized
25 mM EDTA (pH 8.0)
Acknowledgments
We like to thank people from the laboratory for helpful discussions. Work from this laboratory was funded by the Deutsche Forschungsgemeinschaft (DFG) SPP 1258 [Wa455/13-2] and [PU 435/1-1].
References
- Hayatsu, H. and Ukita, T. (1967). The selective degradation of pyrimidines in nucleic acids by permanganate oxidation. Biochem Biophys Res Commun 29(4): 556-561.
- Stratmann, T., Pul, U., Wurm, R., Wagner, R. and Schnetz, K. (2012). RcsB-BglJ activates the Escherichia coli leuO gene, encoding an H-NS antagonist and pleiotropic regulator of virulence determinants. Mol Microbiol 83(6): 1109-1123.
- Sasse-Dwight, S. and Gralla, J. D. (1989). KMnO4 as a probe for lac promoter DNA melting and mechanism in vivo. J Biol Chem 264(14): 8074-8081.
Article Information
Copyright
© 2012 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Pul, Ü., Wurm, R. and Wagner, R. (2012). KMnO4 Footprinting. Bio-protocol 2(21): e280. DOI: 10.21769/BioProtoc.280.
Category
Molecular Biology > DNA > DNA-protein interaction
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Thank you very much for your very good protocol.What is the...
Share
Bluesky
X
Copy link