- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
In vitro Nitrate Reductase Activity Assay from Arabidopsis Crude Extracts
Published: Vol 8, Iss 7, Apr 5, 2018 DOI: 10.21769/BioProtoc.2785 Views: 10313
Reviewed by: Renate WeizbauerVenkatasalam ShanmugabalajiAswad Khadilkar

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
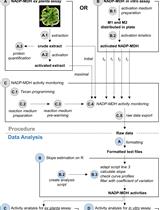
A Semi-throughput Procedure for Assaying Plant NADP-malate Dehydrogenase Activity Using a Plate Reader
Kevin Baudry and Emmanuelle Issakidis-Bourguet
Aug 20, 2023 1496 Views
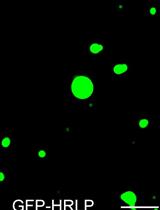
An in vitro Assay to Probe the Formation of Biomolecular Condensates
Yu Zhang and Shen Lisha
Sep 5, 2023 3237 Views
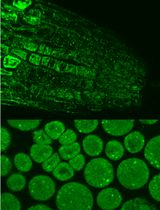
Immunofluorescence for Detection of TOR Kinase Activity In Situ in Photosynthetic Organisms
Ana P. Lando [...] Giselle M. A. Martínez-Noël
Dec 20, 2024 1846 Views
Abstract
Nitrate reductase (NR) reduces the major plant nitrogen source, NO3-, into NO2-. NR activity can be measured by its final product, nitrite through its absorbance under optimized condition. Here, we present a detailed protocol for measuring relative enzyme activity of NR from Arabidopsis crude extracts. This protocol offers simple procedure and data analysis to compare NR activity of multiple samples.
Keywords: Nitrate reductaseBackground
Nitrogen is crucial macronutrient required by plants and is mainly absorbed in the form of nitrate. Nitrate reductase is the first enzyme of the nitrogen assimilation in higher plants. Homodimers of plant nitrate reductase catalyze the NAD(P)H-dependent reduction of nitrate to nitrite as follows:
NO3- + NADH + H+ → NO2- + NAD+ + H2O
Methods to measure NR activity may be a powerful tool to investigate biological factors influencing NR activity (Park et al., 2011). Nitrogen assimilation affects the contents of amino acid in plant, thus regulating NR activity could be used for increasing quality of some crop (Croy and Hageman, 1970; Dalling and Loyn, 1977; Ruan et al., 1998).
In this protocol, increased nitrite concentrations during limited time in optimized buffer condition are acquired as comparable values. Nitrite concentration is measured by its absorbance at 540 nm through Griess assay. Briefly, nitrite forms a diazonium salt with sulfanilic acid, then N-(1-naphthyl) ethylenediamine dihydrochloride is formed colored azo compound. It is possible to compare the values to determine how samples have different NR activity. Furthermore, the values could be converted to exact increased nitrite concentration through a simple process.
Materials and Reagents
- 3MTM MicroporeTM surgical tape (3M, MicroporeTM, catalog number: 1530-1 )
- Reaction tube, 1.5 ml (Greiner Bio One International, catalog number: 616201 )
- 150 x 25 mm (d x h) plastic Petri dish (SPL Life Sciences, catalog number: 10151 )
- Cuvette (Ratiolab, catalog number: 2712120 )
- Arabidopsis seeds
- Ethanol (Merck, EMSURE®, catalog number: 1009831011 )
- Liquid nitrogen
- Potassium nitrite (Sigma-Aldrich, catalog number: P8394 )
- Murashige & Skoog medium including vitamins (Duchefa Biochemie, catalog number: M0222 )
- MES monohydrate (Duchefa Biochemie, catalog number: M1503 )
- Potassium hydroxide (Merck, catalog number: 814353 )
- Sucrose (Duchefa Biochemie, catalog number: S0809 )
- Plant agar (Duchefa Biochemie, catalog number: P1001 )
- Tris-HCl (Duchefa Biochemie, catalog number: T1513 )
- Ethylenediaminetetraacetic acid (EDTA) (Sigma-Aldrich, catalog number: EDS )
- Sodium molybdate dihydrate (Na2MoO4·2H2O) (Sigma-Aldrich, catalog number: M1003 )
- Flavin adenine dinucleotide disodium salt hydrate (FAD-Na2) (Sigma-Aldrich, catalog number: F8384 )
- Dithiothreitol (DTT) (Duchefa Biochemie, catalog number: D1309 )
- Bovine serum albumin (BSA) (Merck, Probumin®, catalog number: 821006 )
- 2-Mercaptoethanol (Sigma-Aldrich, catalog number: M6250 )
- Phenylmethylsulfonyl fluoride (PMSF) (Sigma-Aldrich, catalog number: 78830 )
- Sodium nitrate (NaNO3) (Sigma-Aldrich, catalog number: S5506 )
- Sodium phosphate dibasic (Na2HPO4) (Bio Basic, catalog number: S0404 )
- Sodium phosphate monobasic, anhydrous (NaH2PO4) (Bio Basic, catalog number: SB0878 )
- Nicotinamide adenine dinucleotide (NADH) (Sigma-Aldrich, catalog number: 43420 )
- Hydrochloric acid (HCl) (DAEJUNG CHEMICAL & METALS, catalog number: 4090-4405 )
- Sulfanilamide (Sigma-Aldrich, catalog number: S9251 )
- N-(1-naphthyl) ethylenediamine dihydrochloride (Sigma-Aldrich, catalog number: 222488 )
- MS agar media (see Recipes)
- Extraction buffer (see Recipes)
- Reaction buffer (see Recipes)
- 1% sulfanilamide solution (see Recipes)
- 0.05% N-(1-naptyl) ethylenediamine hydrochloride (see Recipes)
Equipment
- Pipette kit (Gilson, model: PIPETMAN® Classic, catalog number: F167300 )
- 2 L flask (DWK Life Sciences, Duran®, catalog number: 21 216 63 )
- Arabidopsis growth chamber (Hanbaek, model: HB-301L-3 )
- Stainless steel tweezers
- Mortar & pestle (Silico & Chemico Porcelain Works, catalog number: J-753 )
- Centrifuge (Thermo Fisher scientific, model: SorvallTM LegendTM 17 , catalog number: 75002431)
- Spectrophotometer (Biochrom, model: Libra S22 )
- Magnetic stirrer (Vision Scientific, model: VS-130SH )
- Stirring bar
- pH meter (Fisher Scientific, model: accumetTM AB15 )
- Autoclave (Hanbaek, model: HB-506-6 )
Procedure
- Plant preparation
- Sterilize Arabidopsis seeds with 70% EtOH three times.
- Plate the seeds on Murashige and Skoog (MS) medium (see Recipes) containing 0.75% agar.
- Wrap 3M micropore tape around the circumference of each MS plate and store the plates in the dark at 4 °C for 2 days.
- After 2 days, move the plates to fully automated growth chambers under long day conditions (16 h, 22 °C under light/8 h, 20 °C under dark).
- Sterilize Arabidopsis seeds with 70% EtOH three times.
- Tissue extraction
- After 15 days, take tissue of interest (0.5 g fresh weight) using stainless steel tweezer.
- Freeze the samples in liquid nitrogen.
- Grind samples in liquid nitrogen with chilled mortar and pestle.
- Add 750 μl of chilled extraction buffer (see Recipes) to ground sample to homogenize.
- Collect the homogenate into new 1.5 ml tubes through a pipette.
- Centrifuge at 17,000 x g for 5 min.
- Collect supernatant into new 1.5 ml tubes and discard pellet.
- After 15 days, take tissue of interest (0.5 g fresh weight) using stainless steel tweezer.
- Nitrate reductase enzymatic assay
- Add 150 μl of supernatant to 850 μl of reaction buffer (see Recipes) in a 1.5 ml tube.
Note: The reaction buffer contains substrate (nitrate) and NADH to initiate reaction. - Incubate at room temperature for 2 h.
Note: To make blank, skip incubation procedure and do next step. - Add 200 μl of 1% sulfanilamide and 200 μl of 0.05% N-(1-naphthyl) ethylenediamine hydrochloride by pipetting to stop the reaction.
- Incubate at room temperature for 15 min.
- Add 150 μl of supernatant to 850 μl of reaction buffer (see Recipes) in a 1.5 ml tube.
- Measure absorbance
- Transfer 1 ml of the reaction mixture to a cuvette.
- Measure the absorbance of the reaction mixture at 540 nm.
- Transfer 1 ml of the reaction mixture to a cuvette.
- Preparation of nitrite standard curve
- Prepare a nitrite dilution series using potassium nitrite (0, 5, 10, 20, and 40 μM) in 1,250 μl of 0.6x diluted extraction buffer (add 500 μl of ddH2O to 750 μl of extraction buffer).
Note: The dilution ratio of extracted buffer for standards is determined based on the assumption that 1 mg fresh weight has a volume of about 1 µl. - Add 150 μl of dilution series to 850 μl of reaction buffer (see Recipes) in a 1.5 ml tube.
- Add 200 μl of 1% sulfanilamide and 200 μl of 0.05% N-(1-naptyl) ethylenediamine hydrochloride by pipetting.
- Incubate at room temperature for 15 min.
- Transfer 1 ml of the reaction mixture to a cuvette.
- Measure the absorbance of the reaction mixture at 540 nm.
- Generate standard curve for known concentrations of nitrite.
- Prepare a nitrite dilution series using potassium nitrite (0, 5, 10, 20, and 40 μM) in 1,250 μl of 0.6x diluted extraction buffer (add 500 μl of ddH2O to 750 μl of extraction buffer).
- Waste disposal
- All waste must be disposed of in accordance with federal, state and local environmental control regulations.
- All waste must be disposed of in accordance with federal, state and local environmental control regulations.
Data analysis
Values acquired from this protocol are quantity of nitrite molecules. Thus, calculate the NR activity as follows:
ΔA = Asample - Ablank
Each sample is assayed in triplicate and ΔA value is blank subtracted from the average of triplicates.
ΔA value means the increased concentration of nitrite which is produced during 2 h incubation time (see Step C2) and it could be used as a comparable scale of NR activity. For someone who wants to acquire the exact increased nitrite concentration, calculate the concentration change using the standard curve. The ratio of increasing nitrite content of the sample (nmol NO2- g-1 FW h-1) using the formula as follows:
[change of nitrite concentration (μM)] x [extracted volume (ml)]/[fresh weight (g)]/[reaction time (h)]
Recipes
- MS agar media
4.4 g Murashige Skoog basal salt mixture and 0.5 g MES monohydrate in 1 L of deionized water
Adjust pH to 5.8 with 5 M KOH
Add 10 g sucrose
Add 7.5 g plant agar
Autoclave at 121 °C for 15 min
Store the medium at room temperature
Pour media to 150 x 25 mm (d x h) plastic Petri dish
Poured plated can be stored at 4 °C - Extraction buffer
250 mM Tris-HCl (pH 8.0)
1 mM EDTA
1 μM Na2MoO4
5 μM flavin adenine dinucleotide(FAD)
3 mM dithiothreitol
1% BSA
12 mM 2-mercaptoethanol
250 μM PMSF
Note: Extraction buffer should be prepared fresh, used immediately and stored on ice. - Reaction buffer
40 mM NaNO3
80 mM Na2HPO4
20 mM NaH2PO4 (pH 7.5)
0.2 mM NADH
Note: Reaction buffer without NADH can be stored at 4 °C for several months. Since NADH is added, the buffer should be used immediately. - 1% sulfanilamide solution
Dissolved in 3 M HCl
Note: This solution is stable for several months. - 0.05% N-(1-naphthyl) ethylenediamine hydrochloride
Dissolved in distilled water. Store the solution in a dark bottle
Note: This solution is stable for 1 month.
Acknowledgments
This work was supported by a grant from the Next-Generation BioGreen 21 Program (Plant Molecular Breeding Center No. PJ01327601), Rural Development Administration, Republic of Korea.
This protocol was adapted and modified as described previously (Samuelson and Larsson, 1993; Yu et al., 1998, Hachiya and Okamoto, 2016). The authors declare that they have no conflict of interest.
References
- Croy, L. I. and Hageman, R. H. (1970). Relationship of nitrate reductase activity to grain protein production in wheat. Crop Science 10(3): 280-285.
- Dalling, M. and Loyn, R. (1977). Level of activity of nitrate reductase at the seedling stage as a predictor of grain nitrogen yield in wheat (Triticum aestivum L.). Aust J Agr Res 28(1):1-4.
- Hachiya, T. and Okamoto, Y. (2016). Simple spectroscopic determination of nitrate, nitrite, and ammonium in Arabidopsis thaliana. Bio-protocol 7(10):e2280.
- Park, B. S., Song, J. T. and Seo, H. S. (2011). Arabidopsis nitrate reductase activity is stimulated by the E3 SUMO ligase AtSIZ1. Nat Commun 2: 400.
- Ruan, J., Wu, X., Ye, Y. and Härdter, R. (1998). Effect of potassium, magnesium and sulphur applied in different forms of fertilisers on free amino acid content in leaves of tea (Camellia sinensis L). J Sci Food Agr 76(3):389-396.
- Samuelson, M. E. and Larsson, C. M. (1993). Nitrate regulation of zeation riboside levels in barley roots: effects of inhibitors of N assimilation and comparison with ammonium. Plant Science 93(1-2):77-84.
- Yu, X., Sukumaran, S. and Mrton, L. (1998). Differential expression of the Arabidopsis Nia1 and Nia2 genes. cytokinin-induced nitrate reductase activity is correlated with increased nia1 transcription and mrna levels. Plant Physiol 116(3): 1091-1096.
Article Information
Copyright
© 2018 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Kim, J. Y. and Seo, H. S. (2018). In vitro Nitrate Reductase Activity Assay from Arabidopsis Crude Extracts. Bio-protocol 8(7): e2785. DOI: 10.21769/BioProtoc.2785.
Category
Plant Science > Plant biochemistry > Protein > Activity
Plant Science > Plant biochemistry > Other compound
Biochemistry > Protein > Activity
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link









