- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Intravenous Labeling and Analysis of the Content of Thymic Perivascular Spaces
Published: Vol 8, Iss 5, Mar 5, 2018 DOI: 10.21769/BioProtoc.2757 Views: 7651
Reviewed by: Ivan ZanoniYang FuMeenal Sinha

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
An Automated Imaging Method for Quantification of Changes to the Endomembrane System in Mammalian Spheroid Models
Margaritha M. Mysior and Jeremy C. Simpson
Jun 5, 2025 1693 Views

Quantifying Intracellular Distributions of HaloTag-Labeled Proteins With SDS-PAGE and Epifluorescence Microscopy
Julia Shangguan and Ronald S. Rock
Jul 20, 2025 2536 Views
Fluorescence Lifetime-Based Separation of FAST-Labeled Cellular Compartment
Aidar R. Gilvanov [...] Yulia A. Bogdanova
Oct 5, 2025 1352 Views
Abstract
Following development in the thymus, T cells are thought to exit into the periphery predominantly through perivascular spaces (PVS). This exit route is used by conventional T cells, and likely also applies to unconventional T cell subsets, such as precursors of CD8αα and TCRγδ intraepithelial lymphocytes, regulatory T cells and natural killer T cells. Additional cell types might also be found in the PVS and initiate interactions with exiting T cells. The exact content of the PVS, and the processes within, are not well studied. To distinguish vascular from resident cells within various tissues by flow cytometry, intravenous (i.v.) labeling is becoming a commonly employed method. We recently used anti-CD45.2 antibodies and magnetic enrichment to further evaluate this technique, and compared labeled and unlabeled cells in the thymus and blood. This assay can be used to specifically investigate hematopoietic cell subsets within the PVS of the thymus.
Keywords: Perivascular spacesBackground
Immature thymocytes undergo a series of maturation steps, including positive and negative selection, which eliminate the majority of developing T cells. The resulting mature T cell pool is thereby shaped towards a higher proportion of beneficial clones and a reduced proportion of dangerous self-reactive clones. The thymus also produces less abundant T cell subsets that generally act to maintain immune system, tissue, and metabolic homeostasis, including: TCRγδ cells, regulatory T (Treg) cells, natural killer T (NKT) cells, intraepithelial lymphocyte (IEL) precursors, and mucosal associated invariant T (MAIT) cells. Mature thymocytes poised to emigrate into the periphery upregulate expression of the receptor (S1PR1) that recognizes sphingosine-1 phosphate (S1P), a lipid molecule present at high concentrations in the blood. S1PR1+ T cells migrate along an S1P gradient and wind up in vascular circulation.
The thymic perivascular spaces (PVS) are basement membrane-separated compartments between the parenchyma and the vasculature. They are thought to facilitate trafficking of cells, especially mature T cells emigrating from the thymus (Mori et al., 2007; Weinreich and Hogquist, 2008; Zachariah and Cyster, 2010). The exact content of the PVS is not well characterized yet, and could include antigen presenting cells such as dendritic cells and macrophages, that carry antigens not normally expressed in the thymus, into this tissue to contribute to thymocyte selection processes.
Intravenous (i.v.) labeling is a technique commonly used to distinguish vasculature-associated circulating cells from those residing within tissues at the time of analysis (Anderson et al., 2014). Cyster and colleagues have used this approach to identify CD4+ emigrating T cells within the PVS, and showed that upon tail-vein injection of CD4-labeling antibody, CD4 T cells in the PVS are positively labeled within 3 min (Zachariah and Cyster, 2010). We sought to establish whether thymic precursors of CD8αα IEL, an agonist selected T cell subset that downregulates both CD4 and CD8 expression during thymic maturation, can also be found in the PVS. In order to do so, we adapted the i.v. labeling approach, using phycoerythrin (PE)-conjugated anti-CD45.2 (for C57BL/6 mice). As CD45 is not T cell specific, but is expressed by hematopoietic cells in general, various cells can be identified within the i.v.-labeled (IV+) fraction. Furthermore, we combined this with magnetic enrichment for the PE-conjugated antibodies. This allowed us to more closely evaluate the perivascular contents.
Materials and Reagents
- 1.5 ml microcentrifuge tubes (DOT Scientific, catalog number: 509-FTG )
- Aluminum foil (Spring Grove)
- 6-well plates (Corning, Costar®, catalog number: 3506 )
- 70 μm cell strainers (Corning, Falcon®, catalog number: 352350 )
- 1 ml insulin syringes (BD, catalog number: 329420 )
- 3 ml syringes (Covidien, catalog number: 8881513918 )
- 5 ml polystyrene round-bottom tubes (flow cytometry tubes; Corning, Falcon®, catalog number: 352008 )
- 15 ml conical centrifuge tubes (Corning, Falcon®, catalog number: 352097 )
- 96 round-bottom well plates (SARSTEDT, catalog number: 82.1582.001 )
- MACS LS columns (Miltenyi Biotec, catalog number: 130-042-401 )
- Mice, C57BL/6J, 5-6 weeks old (THE JACKSON LABORATORY, catalog number: 000664 )
- Anti-CD45.2 PE clone 104 (Tonbo Biosciences, catalog number: 50-0454-U100 ; 0.2 mg/ml)
- Heparin sodium injection (Sagent Pharmaceuticals, NDC 25021-400-10; 10,000 USP units per 10 ml)
- Phosphate buffered saline (PBS) (Corning, Mediatech, catalog number: 21-040-CV )
- Isoflurane (Piramal Healthcare, 001725CS)
- Anti-PE MicroBeads (Miltenyi Biotec, catalog number: 130-048-801 )
- Live/Dead Fixable Aqua kit (Thermo Fisher Scientific, InvitrogenTM, catalog number: L34957 )
- CD1d-tetramer (PBS57-loaded CD1d-monomers and tetramers available from NIH tetramer core facility; http://tetramer.yerkes.emory.edu)
- Anti-CD25 clone PC61 (e.g., BioLegend, catalog number: 102024 )
- Anti-TCRβ clone H57-597 (e.g., BD, BD Biosciences, catalog number: 562841 )
- Anti-CD4 clone RM4-5 (e.g., BioLegend, catalog number: 100548 )
- Anti-CD8α clone 53-6.7 (e.g., BD, BD Biosciences, catalog number: 563332 )
- Anti-CD5 clone 53-7.3 (e.g., Thermo Fisher Scientific, eBioscience, catalog number: 47-0051-82 )
- Anti-CD122 clone TM-b1 (e.g., Thermo Fisher Scientific, eBioscience, catalog number: 46-1222-82 )
- Anti-H-2Kb clone AF6-88.5 (e.g., BD, BD Biosciences, catalog number: 562942 )
- Anti-PD-1 clone J43 (e.g., Thermo Fisher Scientific, eBioscience, catalog number: 17-9985-82 )
- Anti-NK1.1 clone PK136 (e.g., BioLegend, catalog number: 108705 )
- Anti-CD11c clone N418 (e.g., Thermo Fisher Scientific, eBioscience, catalog number: 25-0114-82 )
- Anti-I-Ab clone AF6-120.1 (e.g., BioLegend, catalog number: 116421 )
- Anti-CD19 clone 1D3 (e.g., Thermo Fisher Scientific, eBioscience, catalog number: 56-0193-82 )
- Anti-CD11b clone M1/70 (e.g., Thermo Fisher Scientific, eBioscience, catalog number: 11-0112-41 )
- Anti-GR1 clone Gr-1 (e.g., BioLegend, catalog number: 108411 )
- Fetal bovine serum (FBS) (Atlanta Biologicals, catalog number: S11150 ), heat inactivated at 65 °C
- Ethylenediamine tetraacetate acid (EDTA) (Fisher Scientific, catalog number: BP120-500 )
- Sodium azide (Fisher Scientific, catalog number: BP922I-500 )
- Ammonium chloride (NH4Cl) (Sigma-Aldrich, catalog number: A4514-500G )
- Potassium bicarbonate (KHCO3) (Fisher Scientific, catalog number: P235-500 )
- Bovine serum albumin (BSA) (Sigma-Aldrich, catalog number: A7906 )
- FACS buffer (see Recipes)
- ACK lysis buffer (see Recipes)
- MACS buffer (see Recipes)
Equipment
- Class-II biosafety cabinet/laminar flow hood
- MACS Multistand (Miltenyi Biotec, catalog number: 130-042-303 )
- QuadroMACS Separator (Miltenyi Biotec, catalog number: 130-090-976 )
- Heat lamp
- Timer (Fisher Scientific, catalog number: 14-649-17 )
- Benchtop centrifuge (Beckman Coulter, model: Allegra X-12-R )
- 2,000 ml drop glass jar
- Refrigerator (4 °C) (Fisher Scientific, model: IsotempTM General-Purpose Series Lab Refrigerator )
- Hemacytometer (Sigma-Aldrich, catalog number: Z359629-1EA )
- FACS flow cytometer (BD, model: LSR-II , H10.10)
Software
- FlowJo version 10.4.0
Procedure
Note: All animal procedures must be approved by your institution’s ethics committee.
- Intravenous labeling and sample preparation
Note: Perform Steps A6-A8 in a class-II biosafety cabinet/laminar flow hood with exhaust. Figure 1 shows an outline of the procedure up to the tissue collection.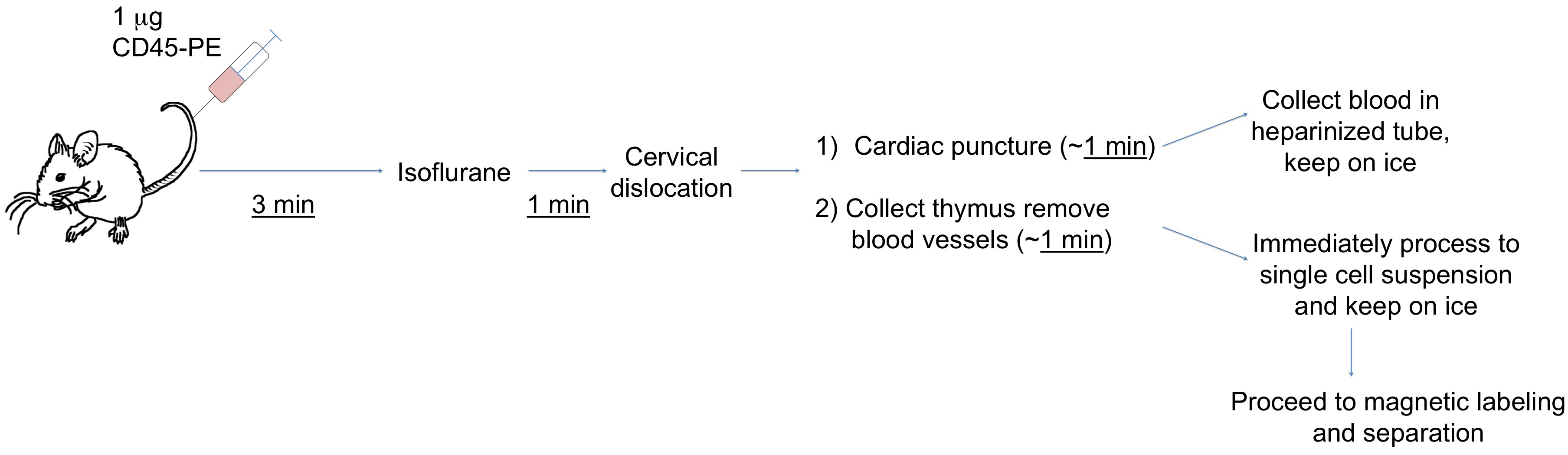
Figure 1. Outline of IV labeling and tissue collection. Experimental procedures prior to magnetic cell sorting are depicted as explained in the text. It is important to work quickly and do one animal at a time, such that the time from injection to single cell suspension does not vary greatly between samples.- Prepare anti-CD45-PE/PBS solution (1 μg/200 μl): Dilute 5 μl anti-CD45.2-PE in 195 μl PBS in a 1.5 ml microcentrifuge tube, vortex, place on ice, covered with aluminum foil.
Note: It is advisable to prepare 5-10% more solution than needed to account for solution loss within the syringe dead space. - Prepare a 6-well plate with 5 ml FACS buffer (see Recipes) and a 70 μm cell strainer per well for collection of thymi, place on ice.
- Prepare 1.5 ml microcentrifuge tubes with 20 μl heparin for blood collection, place on ice.
- Working one animal at a time, warm up mice with a heat lamp until tail vein is dilated (usually ~5 min), constantly monitoring mice to avoid overheating or burning.
- Set timer 3 min. When tail vein is dilated, inject 200 μl of the anti-CD45-PE/PBS solution into the tail-vein, using a 1 ml insulin syringe. Immediately after injection, start the timer.
Note: Include mice injected with 200 μl PBS as negative controls. - While the timer is running, prepare a glass jar inside a class-II biosafety cabinet with exhaust for the isoflurane drop method: Place a folded tissue soaked with 2 ml isoflurane into the bottom part of a 2,000 ml drop glass jar, and place insert on top, ensuring physical separation of isoflurane-soaked tissue and the mouse.
- When the timer goes off 3 min after the injection, place the mouse inside the prepared glass jar and close the lid.
- After 1 min (mouse should be anesthetized), euthanize mouse by cervical dislocation.
- Open the chest of the animal and collect blood by cardiac puncture, using a 1 ml insulin syringe: Insert the needle about 5 mm into the heart, and gently pull the plunger (Figures 2A and 2B). Empty the syringe content (usually around 0.2-0.5 ml) into a heparinized 1.5 ml microcentrifuge tube, and place the collection tube with blood on ice. Cover samples with aluminum foil to protect from light until further processing.
- Collect the thymus and remove blood vessels attached to the outside of the thymus (Figures 2C-2E). Suspend in 5 ml cold FACS buffer (6-well plate with 70 μm cell strainer) on ice, and immediately dissociate the tissue by mashing it with the plunger of a 3 ml syringe and pressing it through the cell strainer into the well. Discard the strainer with the remaining connective tissue. Repeat for each thymus. Cover samples with aluminum foil to protect from light until further processing.
- Blood samples: Transfer blood into 5 ml flow cytometry tubes, add 2 ml ACK lysis buffer (see Recipes), keep at room temperature (RT) for 2 min, spin for 3 min at 524 x g (1,500 RPM), 4 °C, discard supernatant. Repeat 2 times (resuspending pellets in ACK lysis buffer) and finally resuspend cells in 1 ml cold FACS buffer and place on ice.
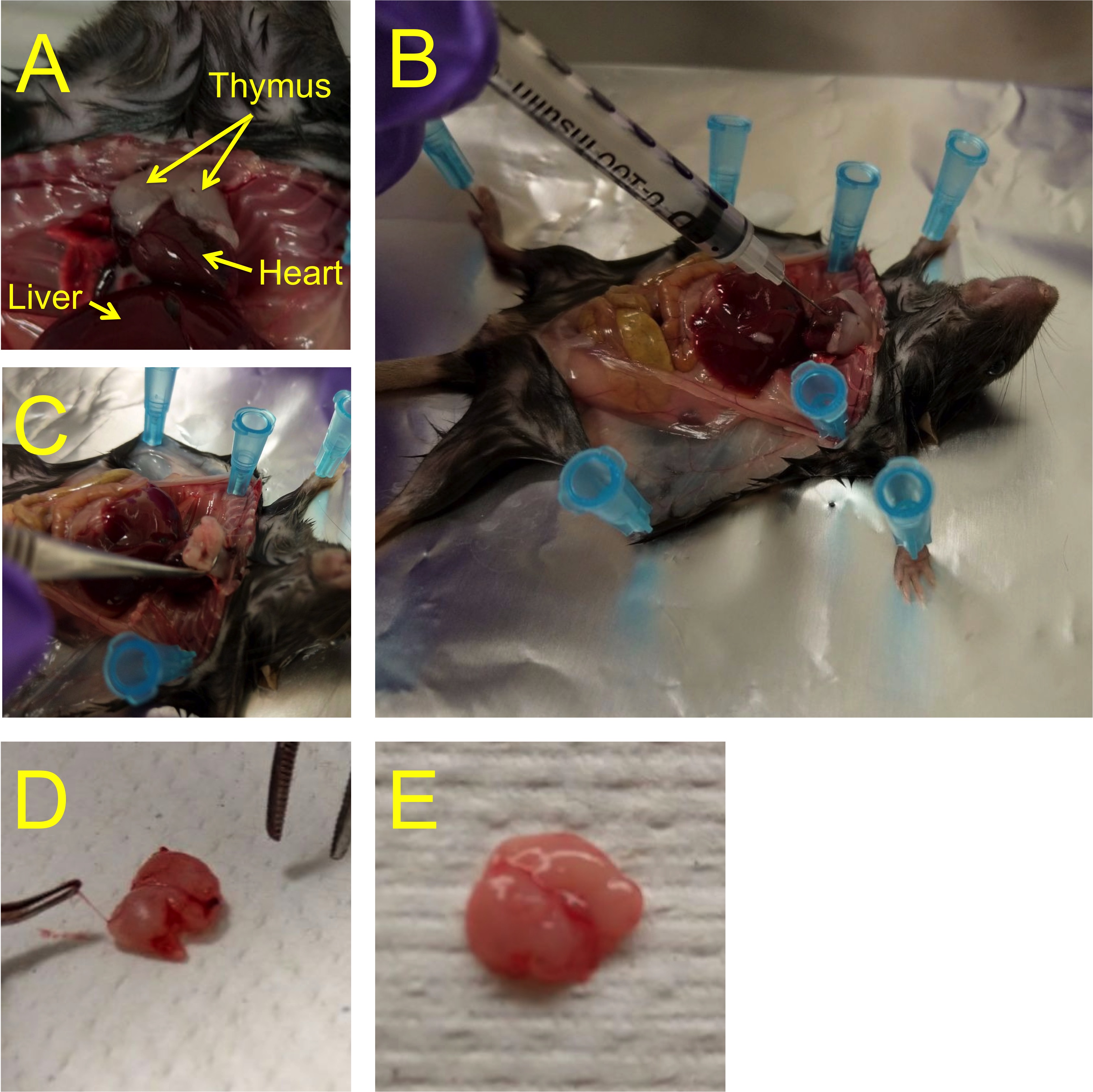
Figure 2. Outline of cardiac blood and thymus collection. A. The thymus is localized right above the heart. B. Blood is drawn from the heart with an insulin syringe. C. To remove the thymus, tweak it at the base with a curved forceps and gently pull. D. The thymus is covered with blood vessels, which should be removed with a curved forceps as best as possible. E. Thymus after removal of lining blood vessels.
- Prepare anti-CD45-PE/PBS solution (1 μg/200 μl): Dilute 5 μl anti-CD45.2-PE in 195 μl PBS in a 1.5 ml microcentrifuge tube, vortex, place on ice, covered with aluminum foil.
- MACS enrichment of i.v.-labeled (IV+) cells from thymic samples
- Determine cell count of each sample by hemacytometer (usually 150-250 x 106).
- Resuspend cells to 1 x 107/80 μl MACS buffer (see Recipes).
- Set 40 μl (5 x 106) of each sample aside as an untouched control, and fill up to 1 ml with FACS buffer, keep on ice. Cover with aluminum foil.
- In accordance with the Miltenyi protocol (included with the anti-PE MicroBeads), add 20 μl anti-PE beads per 80 μl to the remaining suspensions.
Note: In our experience, this protocol also works with lower amounts of beads. E.g., resuspend cells to 1 x 107/40 μl MACS buffer and add 10 μl beads. Further reduction of beads might be possible. If looking at abundant cells, a fraction of the thymic sample could be sufficient for enrichment. - Mix, and incubate in the refrigerator (4 °C) for 15 min, protected from light.
- Wash samples 2 x with 1 ml MACS buffer (spin for 5 min at 524 x g (1,500 RPM), 4 °C, discard supernatant, resuspend in MACS buffer).
- After the second wash step, resuspend the samples at up to 1 x 108 in 500 μl MACS buffer.
- Perform magnetic separation using LS columns and according to the Miltenyi protocol.
- Collect the magnetically-labeled (IV+) fractions in 15 ml centrifuge tubes.
Note: We use untouched fractions of samples as controls. The flowthrough from the magnetic separation step can also be collected as a negative control. We generally observed no differences between the untouched and flowthrough IV- fractions when analyzed.
- Determine cell count of each sample by hemacytometer (usually 150-250 x 106).
- Antibody- and tetramer-labeling for flow cytometry
- Determine cell count of each IV+ eluted sample (and IV- flowthrough sample, if collected)
- Spin samples (including untouched fractions) for 5 min at 524 x g RPM (1,500), 4 °C, discard supernatant, resuspend in 200 μl FACS buffer and transfer each sample into a well of a 96 round-bottom well plate, place on ice.
Note: We stain samples at up to 5 x 106 cells per 50 μl master-mix (see below). The IV+ fraction per thymus, and the untouched fractions are generally within this range. If collecting flowthrough fractions, we recommend to determine the cell number and stain 5 x 106 cells. - Determine cell numbers of the blood samples and transfer 5 x 106 cells per sample into individual wells.
- Live/Dead discriminator: In our lab we frequently use Live/Dead Fixable Aqua kit. We use this kit prior to labeling with markers of interest, and stain cells for 20 min on ice in PBS (without added protein) with 1:1,000 Live/Dead discriminator. This amine-binding discriminator stains cells that have lost their membrane integrity, and positively stained (dead) cells should be excluded when analyzing the FACS data.
- Wash samples 3 x with 200 μl FACS buffer (spin for 5 min at 524 x g (1,500 RPM), 4 °C, discard supernatant, resuspend in FACS buffer).
- Prepare master mix for labeling for FACS. We use most antibodies at 1:200 μl and CD1d-tetramer at 1:400 in FACS buffer, 50 μl per 5 x 106 cells.
We typically include the following set of antibodies to analyze CD4+CD8α- (CD4 single-positive, SP), CD4-CD8α+ (CD8 SP), CD4+CD8α+ (double-positive, DP) and IEL-precursor (CD1d-tetramer-CD25-CD4-CD8α-CD5+TCRα+CD122+H-2Kb+ PD-1+ or PD-1-NK1.1+) T cells (panel 1), or B cells (CD19+I-Ab+), dendritic cells (CD11c+I-Ab+) and CD11b+GR-1+ myeloid cells (panel 2), and these panels can be extended/substituted according to the cells of interest: Panel 1: CD1d-tetramer, CD25 TCRβ, CD4, CD8α, CD5, CD122, H-2Kb, PD-1, NK1.1; Panel 2: CD11c, I-Ab, CD19, CD11b, GR1.
Note: If B cells or myeloid cells are the specific focus of investigation, it is advisable to include an Fc receptor blocker (anti-CD16/32). - Spin 96-well plate with samples for 5 min at 524 x g (1,500 RPM), 4 °C, discard supernatant, resuspend in 50 μl master mix per 5 x 106 cells. Keep on ice, covered with aluminum foil, for 20 min.
- Wash samples 3 x with 200 μl FACS buffer (spin for 5 min at 524 x g (1,500 RPM), 4 °C, discard supernatant, resuspend in FACS buffer).
- After the final wash step, resuspend each sample in 200 μl FACS buffer and transfer into polystyrene FACS tubes. Keep on ice and cover with aluminum foil until acquisition.
- Determine cell count of each IV+ eluted sample (and IV- flowthrough sample, if collected)
- Flow cytometry
- Set up compensation on a flow cytometer.
- Acquire samples.
- Set up compensation on a flow cytometer.
Data analysis
- Samples are analyzed with FlowJo. Within singlet Live/Dead-negative thymocytes, gates are set on the IV- and IV+ fractions (see Figure 3A).
- The IV- fraction of the untouched or flowthrough samples should be compared to the IV+ fraction of the enriched and blood samples (Figure 3).
- Identify and analyze cells of interest.
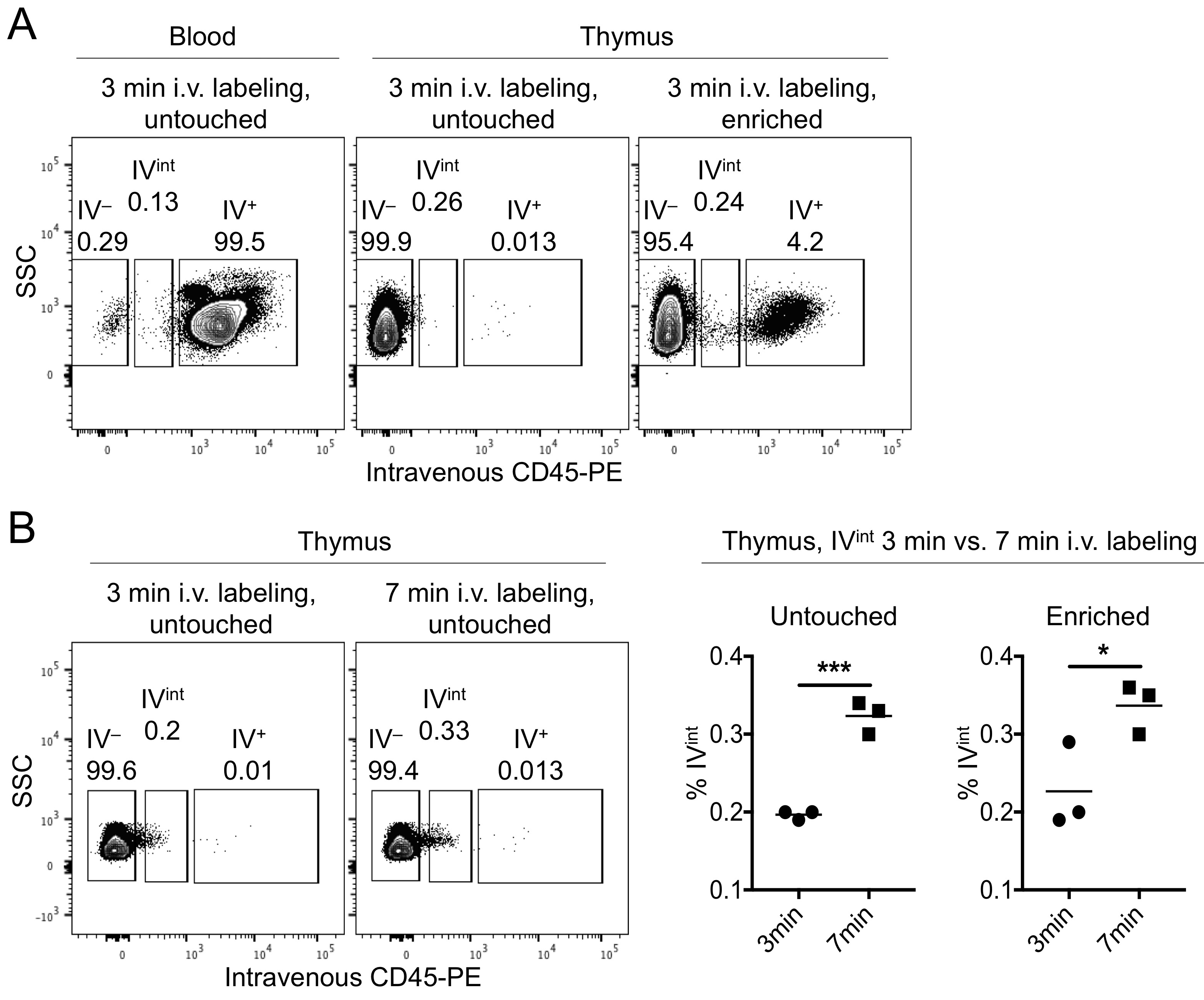
Figure 3. Gating strategy for IV+ and IV- cells. A. The IV-labeled (IV+) or–unlabeled (IV-) cells were identified within the thymic untouched (left) or MACS-enriched samples (middle), or in the blood (right). Further analysis of cells of interest can be undertaken with an appropriate staining panel. B. 3 min versus 7 min i.v. labeling times were compared in one experiment (dot plots). Graphs (right) show the percent i.v.-intermediate (IVint) cells in the untouched (left graph) and enriched (right graph) fractions. *P < 0.05, *** P < 0.001, Student’s t-test. Each symbol represents an individual mouse.
Notes
- For reference: From 6-9 week old mice (predominantly females), we normally obtain between 200-300 x 106 total cells and recover between 1.5-4.5 x 103 total enriched IV+ cells. Within the enriched IV+ cells, we found about 48.15 (± 2.52)% TCRβ+, 2.97 (± 0.31)% CD19+I-Ab+ B cells and 2.21 (± 0.54)% CD11b+GR1+ cells. Also see reference Ruscher et al. (2017), Figure 5 and Suppl. Figure 4.
- For first time users of this protocol, we recommend using up to three mice.
- We tested 3 min versus 7 min IV labeling time in one experiment. In our hands there was an increased ‘leakage’ of labeling antibody into the thymic parenchyma, as observed by a PE+ shoulder of the IV- fraction (see IVint in Figure 3B). We recommend adhering to the suggested 3 min labeling time.
- We also tested the effects of perfusion with 50 ml PBS immediately after the 3 min i.v. labeling and 1 min isoflurane steps, and did not observe differences in the proportions of cell types within the PVS with or without perfusion. Furthermore, some results suggest that perfusion can damage tissue architecture in a way that affects i.v. labeling (Anderson et al., 2014).
- Zachariah et al. confirmed accuracy of i.v. labeling of CD4+ T cells in the thymic PVS with FTY720 (an analog of S1P that works as an antagonist) treatment, and in RAG-GFP models (Zachariah and Cyster, 2010). We compared the percentages of T cell subsets, B cells and CD11b+GR-1+ cells within the blood IV+, untouched IV- (parenchyma) and enriched IV+ fractions (Ruscher et al., 2017). We further have unpublished data using CD24 and additional markers, and investigated the %GFP+ and GFP median fluorescent intensity after i.v. labeling of Rag2GFP. [Rag2GFP indicates the age of T cells after T cell receptor recombination events, with GFP fluorescence being high at the DP stage and after positive selection decreasing over time (Boursalian et al., 2004; McCaughtry et al., 2007).] The differences indicated that the enriched IV+ fractions were distinct from the blood IV+ fractions, suggesting that this protocol predominantly identifies cells that are in the thymic PVS.
- In addition to the anti-CD45 antibodies used in our PVS labeling studies, we and others (Zachariah and Cyster, 2010; Anderson et al., 2014) have employed anti-CD4 (clone RM4-4, eBioscience; clone RPA-T4, BD Biosciences) and anti-CD8α (clone 53-6.7, eBioscience) for intravenous labeling of vascular content in various peripheral tissues.
- Similarly, while not tested specifically for PVS content labeling, we have used fluorochromes such as allophycocyanin (APC) or Brilliant Violet 421 (BV421) for i.v. labeling of peripheral tissue vasculature-associated T cells. These fluorochromes might also work for the purpose of identifying the thymic PVS content by i.v. labeling.
Recipes
- FACS buffer
Supplement 1 L PBS with:
1% FBS
2 mM EDTA
0.02% sodium azide - ACK lysis buffer
Supplement 1 L ddH2O with:
150 mM NH4Cl
10 mM KHCO3
0.1 mM EDTA
Adjust pH to 7.2-7.4 - MACS buffer
Supplement 1 L PBS with:
0.5% BSA
2 mM EDTA
Acknowledgments
We would like to thank Rebecca Kummer for assistance with some i.v. labeling experiments, and Oscar Salgado Barrero for assistance with taking photographs. This work was supported by NIH grants R37 AI39560 and PO1 AI35296 to K.A.H. The authors have no competing financial interests.
References
- Anderson, K. G., Mayer-Barber, K., Sung, H., Beura, L., James, B. R., Taylor, J. J., Qunaj, L., Griffith, T. S., Vezys, V., Barber, D. L. and Masopust, D. (2014). Intravascular staining for discrimination of vascular and tissue leukocytes. Nat Protoc 9(1): 209-222.
- Boursalian, T. E., Golob, J., Soper, D. M., Cooper, C. J. and Fink, P. J. (2004). Continued maturation of thymic emigrants in the periphery. Nat Immunol 5(4): 418-425.
- McCaughtry, T. M., Wilken, M. S. and Hogquist, K. A. (2007). Thymic emigration revisited. J Exp Med 204(11): 2513-2520.
- Mori, K., Itoi, M., Tsukamoto, N., Kubo, H. and Amagai, T. (2007). The perivascular space as a path of hematopoietic progenitor cells and mature T cells between the blood circulation and the thymic parenchyma. Int Immunol 19(6): 745-753.
- Ruscher, R., Kummer, R. L., Lee, Y. J., Jameson, S. C. and Hogquist, K. A. (2017). CD8αα intraepithelial lymphocytes arise from two main thymic precursors. Nat Immunol 18(7): 771-779.
- Weinreich, M. A. and Hogquist, K. A. (2008). Thymic emigration: when and how T cells leave home. J Immunol 181(4): 2265-2270.
- Zachariah, M. A. and Cyster, J. G. (2010). Neural crest-derived pericytes promote egress of mature thymocytes at the corticomedullary junction. Science 328(5982): 1129-1135.
Article Information
Copyright
© 2018 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Ruscher, R. and Hogquist, K. A. (2018). Intravenous Labeling and Analysis of the Content of Thymic Perivascular Spaces. Bio-protocol 8(5): e2757. DOI: 10.21769/BioProtoc.2757.
Category
Immunology > Animal model > Mouse
Cell Biology > Cell imaging > Fluorescence
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link