- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Transient Gene Expression for the Characteristic Signal Sequences and the Estimation of the Localization of Target Protein in Plant Cell
Published: Vol 8, Iss 4, Feb 20, 2018 DOI: 10.21769/BioProtoc.2738 Views: 9754
Reviewed by: Arsalan DaudiJian ChenAlexandros Alexandratos

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

Agrobacterium-mediated Genetic Transformation of Cotton and Regeneration via Somatic Embryogenesis
Alka Srivastava [...] Praveen C. Verma
May 20, 2023 4438 Views
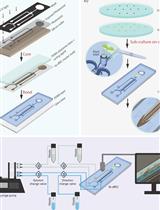
Bi-directional Dual-flow-RootChip for Physiological Analysis of Plant Primary Roots Under Asymmetric Perfusion of Stress Treatments
Claudia Allan [...] Claudia-Nicole Meisrimler
Aug 5, 2023 1985 Views

Enzymatic Starch Quantification in Developing Flower Primordia of Sweet Cherry
Nestor Santolaria [...] Afif Hedhly
Apr 5, 2025 1932 Views
Abstract
We have proposed and tested a method for characterization of the signal sequences and determinations of target protein localization in a plant cell. This method, called the AgI-PrI, implies extraction of protoplasts from plant tissues after agroinfiltration. The suggested approach combines the advantages of two widely used methods for transient gene expression in plants–agroinfiltration and transfection of isolated protoplasts. The AgI-PrI technic can be applied to other plant species.
Keywords: AgroinfiltrationBackground
To date, the following techniques for transient expression of genes in plants have been developed and widely used: agroinfiltration, biolistics of plant explants and transfection of protoplasts using polyethylene glycol or electroporation. The effectiveness of these approaches has been clearly demonstrated. Each strategy for transient expression of genes in plants, along with benefits, has its limitations and disadvantages, such as the difficulties in the fine imaging of recombinant reporter proteins in plant cell compartments owing to intricate shapes of plant epidermal cells (agroinfiltration), a low efficiency of transformation and the necessity of specialized equipment and auxiliary material (for biolistics), as well as complex preparatory procedures required for a high yield of viable protoplasts and their effective transfection. This is the reason for developing and testing new methods for transient expression of genes in a plant cell, preferably by improving the experimental protocols and preserving the physiological significance of the results of the studies. Since the cellular localization of proteins in living organisms, including plants, is closely interrelated with their functions, a fine visualization of proteins in living cells becomes an important tool for assessing the functions of the proteins.
Materials and Reagents
- Pipettes (Corning, Costar®, catalog number: 4101 )
- Inoculation loop
- Petri dishes (Thermo Fisher Scientific, Thermo ScientificTM, catalog number: 101IRR )
- Syringe without a needle (B. Braun Melsungen, catalog number: 4645103C )
- Scalpel
- Nylon mesh, pore size, 40 µm (Sterile Cell Strainers, Corning, catalog number: 431750 )
- 10-ml tubes (Corning, Axygen®, catalog number: SCT-10ML )
- Agrobacterium tumefaciens strain GV3101 (Mohamed et al., 2004; strain is available in the collection of the Institute of Plant Physiology and can be provided to researchers for experiments)
- Nicotiana benthamiana (Sheludko et al., 2007; seeds are available in the collection of the Institute of Plant Physiology and can be provided to researchers for experiments)
- LB medium (MP Biomedicals, catalog number: 113002042 )
- Rifampicin (Fisher Scientific, catalog number: BP26791 )
- Gentamicin (Thermo Fisher Scientific, GibcoTM, catalog number: 15750060 )
- Kanamycin (Thermo Fisher Scientific, GibcoTM, catalog number: 11815024 )
- Sodium chloride (NaCl) (Sigma-Aldrich, catalog number: S9888 )
- Tris-HCl, pH 7.0 (Roche Diagnostics, catalog number: 10812846001 )
- Calcium chloride (CaCl2) (Sigma-Aldrich, catalog number: C1016 )
- Acetosyringone (Abcam, catalog number: ab146533 )
- MES (Sigma-Aldrich, catalog number: 69892 )
- Magnesium sulfate heptahydrate (MgSO4·7H2O) (Merck, catalog number: 1058860500 )
- Calcium chloride dihydrate (CaCl2·2H2O) (AMRESCO, catalog number: 0556-500G )
- Ammonium phosphate monobasic (NH4H2PO4) (Sigma-Aldrich, catalog number: A3048 )
- Sorbitol (Sigma-Aldrich, catalog number: S1876 )
- Potassium hydroxide (KOH) (AppliChem, catalog number: 211514 )
- Cellulase Onozuka R10 (Kinki Yakult)
- Pectinase Macerozyme R10 (Kinki Yakult)
- Driselase (Sigma-Aldrich, catalog number: D9515 )
- Calcium nitrate tetrahydrate (Са(NО3)2·4H2O) (Sigma-Aldrich, catalog number: C2786 )
- Potassium phosphate monobasic (KH2PO4) (Fisher Scientific, catalog number: P285 )
- Magnesium sulfate (MgSO4) (Acros Organics, catalog number: AC413485000 )
- Potassium chloride (KCl) (Sigma-Aldrich, catalog number: P9333 )
- Ferric chloride (FеСl3) (Sigma-Aldrich, catalog number: 157740 )
- MS medium (Sigma-Aldrich, catalog number: M5519 )
- Sucrose (MP Biomedicals, catalog number: 04802536 )
- Solution 1 (see Recipes)
- Solution 2 (see Recipes)
- Solution 3 (see Recipes)
- Solution 4 (see Recipes)
- Solution 5 (see Recipes)
- Knop’s solution (see Recipes)
- Agroinfiltration buffer (see Recipes)
Equipment
- Incubator Shaker (Biosan, model: ES-20 , catalog number: BS-010111-AAA)
- Centrifuge (Eppendorf)
- Transilluminator (Vilber, model: ETX-F26.M , catalog number: Vilber Lourmat 2131 2600 1)
- Microscope Axio Imager Z2 (ZEISS, model: Axio Imager Z2 ) equipped with digital camera (ZEISS, model: AxioCam MRc5 ), filter set No. 38 (38 Endow GFP shift free (EX BP 470 nm/40 nm, BS FT 495 nm, EM BP 525 nm/50 nm), ZEISS, catalog number: 000000-1031-346 ) and module ApoTome (ZEISS, model: ApoTome )
Software
- ZEN, AxioVision 4.8 (ZEISS)
Procedure
For this protocol, we used plant expression vectors, with leader signal sequences providing their localization in various compartments of plant cells. The vectors were obtained according to the procedure described earlier (Tyurin et al., 2017).
- Cultivation, transformation and selection of A. tumefaciens
- Seed cells of the A. tumefaciens strain GV3101 strain using an inoculation loop in 5 ml оf LB medium containing rifampicin (50 μg/ml), gentamicin (25 μg/ml), and carbenicillin (50 μg/ml), (antibiotics do not affect the efficiency of plant transformation) and grow overnight at 28 °C in a shaker incubator (~140 pm).
- Transfer the bacterial culture to 95 ml of LB medium without antibiotics and grow to an optical density of OD600 = 0.5-0.6, at 28 °C in a shaker incubator (~140 pm).
- Centrifuge cells at 3,500 x g for 5 min, at room temperature. Resuspend the pellet in 20 ml of cold (0 °C) solution 1 (see Recipes).
- Centrifuge cells at 3,500 x g for 5 min, at room temperature. Resuspend the pellet in 2 ml of solution 2 (see Recipes).
- Competent cells should be used for transformation immediately.
- Add 1 µg of an expression vector (with the kanamycin resistance gene) to the competent cells. Incubate cells on ice for 20 min. Heat the solution for 5 min at 37 °C in the shaker incubator. Place cells for 5 min on ice, add 2 ml of LB medium without antibiotics and incubate at 28 °C in the shaker incubator (~140 rpm) for 2 h.
- Plate transformed cells on Petri dishes in the medium containing 50 µg/ml of the selective antibiotic kanamycin (use the appropriate selective agent at concentrations recommended by the manufacturer). Isolate colonies and resuspend the cells in 100-150 μl of LB medium without antibiotics, and then freeze them.
- Seed cells of the A. tumefaciens strain GV3101 strain using an inoculation loop in 5 ml оf LB medium containing rifampicin (50 μg/ml), gentamicin (25 μg/ml), and carbenicillin (50 μg/ml), (antibiotics do not affect the efficiency of plant transformation) and grow overnight at 28 °C in a shaker incubator (~140 pm).
- Plant material
Grow Nicotiana benthamiana at 20 ± 2 °C, a photoperiod of 8 h, and an illumination of 100 μmol quanta/(m2 sec) in soil or hydroponics using Knop’s solution (see Recipes) as a nutrient medium for 6 weeks. Frequency of use: 2 times per week. - Agroinfiltration of the N. benthamiana plants
- Thaw frozen A. tumefaciens cells and grow them for 48 h at 27 °C in LB medium containing rifampicin (50 μg/ml), gentamicin (25 μg/ml), and carbenicillin (50 μg/ml) in the shaker incubator (~140 pm). Replace the medium with LB medium containing the same antibiotics and solution 3 (see Recipes). Grow the cells to an optical density of OD600 = 0.8.
- Centrifuge the cells at 3,000 x g for 5 min, at room temperature. Resuspend the pellet in agroinfiltration buffer (see Recipes) to an optical density of OD600 = 2.4.
- Infiltrate the cells into abaxial epiderm of 6-week-old N. benthamiana leaves using a syringe without a needle (Figure 1).
- Estimate the quality of transformation on the 4th day after agroinfiltration by imaging of the zones with an expression of the desC-egfp hybrid gene in tobacco leaves at 312 nm, using an ETX transilluminator (Vilber Lourmat). Cut the high fluorescence regions of tobacco leaves out for protoplast extraction (Figure 2).
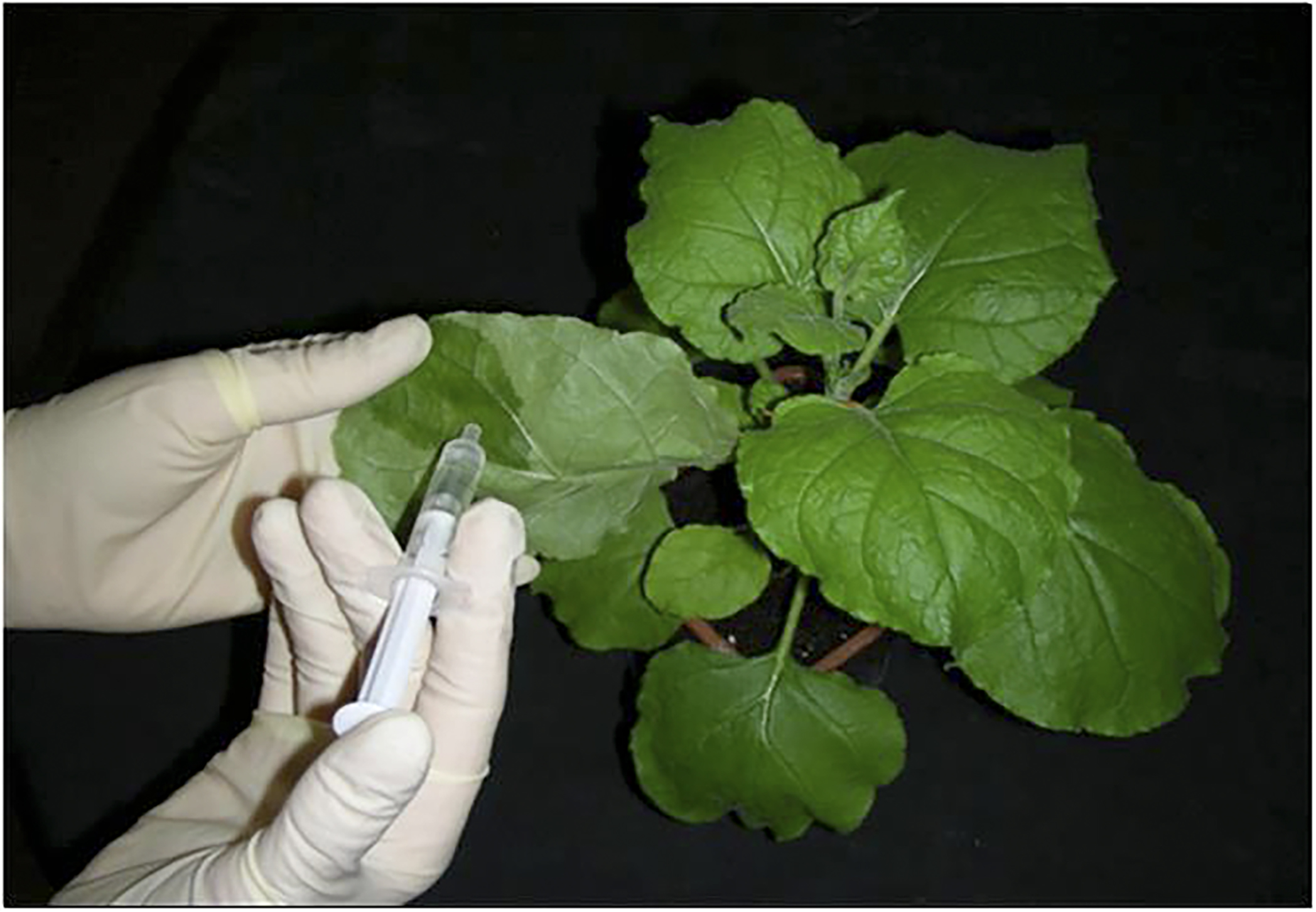

Figure 1. Procedure of agroinfiltration into abaxial epiderm of N. benthamiana leaves using a syringe without a needle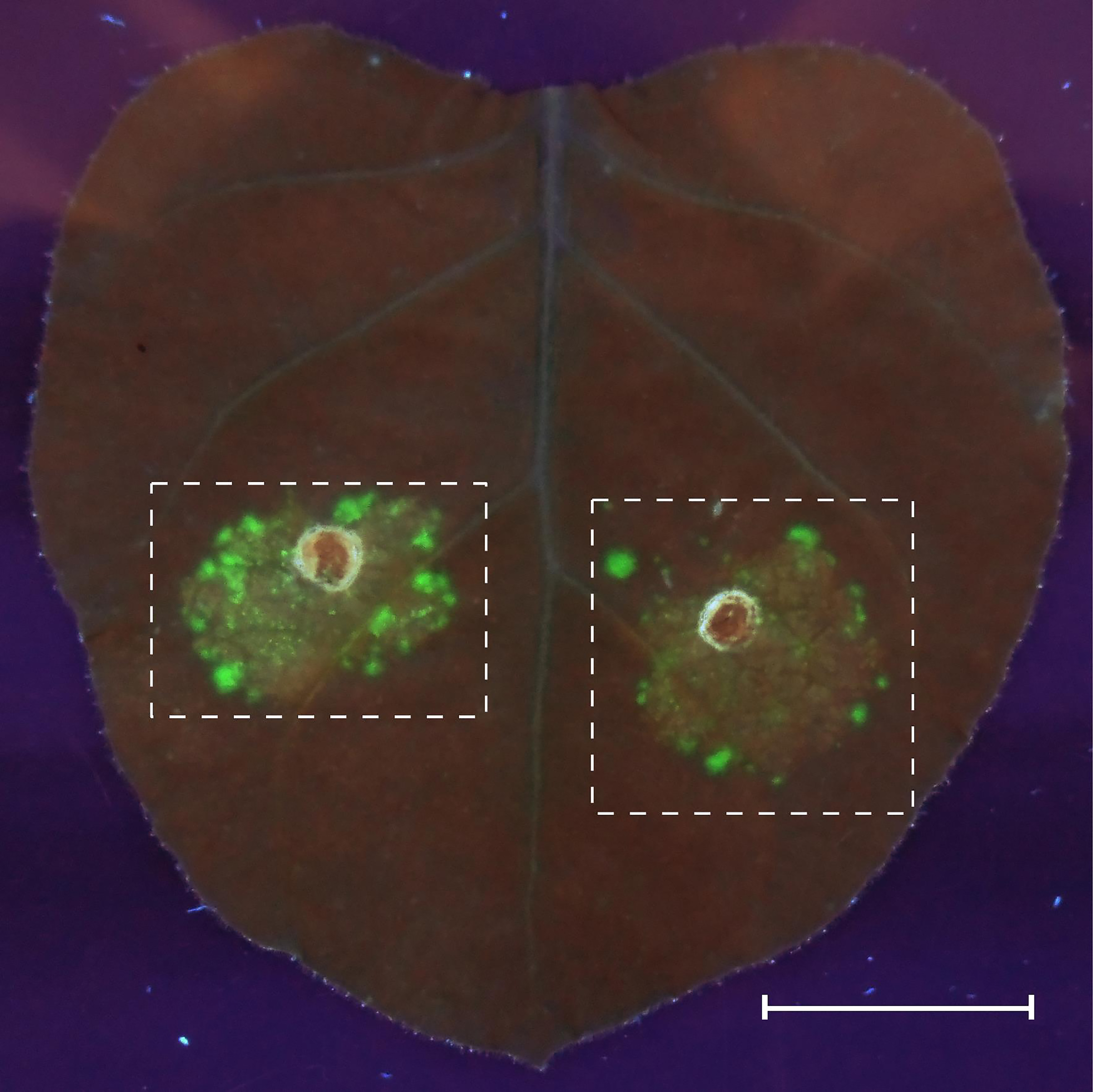
Figure 2. GFP fluorescence in agroinfiltrated Nicotiana benthamiana leaves. The agrobacteria carrying the expression cassettes of GFP fusion protein were syringe-infiltrated into an N. benthamiana leaf, and GFP fluorescence was observed 4 days after infiltration (dpi) in UV light. Fluorescence fields are marked with dashed lines. Scale bar = 20 mm.
- Thaw frozen A. tumefaciens cells and grow them for 48 h at 27 °C in LB medium containing rifampicin (50 μg/ml), gentamicin (25 μg/ml), and carbenicillin (50 μg/ml) in the shaker incubator (~140 pm). Replace the medium with LB medium containing the same antibiotics and solution 3 (see Recipes). Grow the cells to an optical density of OD600 = 0.8.
- Protoplast isolation
- Prepare solution 4 (see Recipes) for the cell wall maceration immediately before use and clarify it by centrifugation.
- Use the leaf fragments with fluorescence for protoplast isolation according to Nosov et al. (2014). Mince the leaves (about 500 mg) with a scalpel. Add 5 ml of solution 4.
- Isolate the protoplasts at 15 °C in a shaker (50 rpm) for 12 h.
- Filtrate the protoplast suspension through nylon mesh (pore size, 40 µm). Transfer the protoplast suspension into 10-ml centrifuge tubes and centrifuge at 100 x g for 5 min at room temperature.
- Resuspend the pellet of protoplasts in 10 ml of solution 5 and incubate for 5 min at room temperature, and then centrifuge at 100 x g for 5 min, repeat the procedure two times. Resuspend the pellet of protoplasts in 1.5 ml of solution 5 (see Recipes).
- Prepare solution 4 (see Recipes) for the cell wall maceration immediately before use and clarify it by centrifugation.
- Protoplast imaging
Perform the imaging of zones with target protein in freshly isolated protoplasts with an Axio Imager Z2 microscope (ZEISS) equipped with an AxioCam MR digital camera and filter units (Figure 3).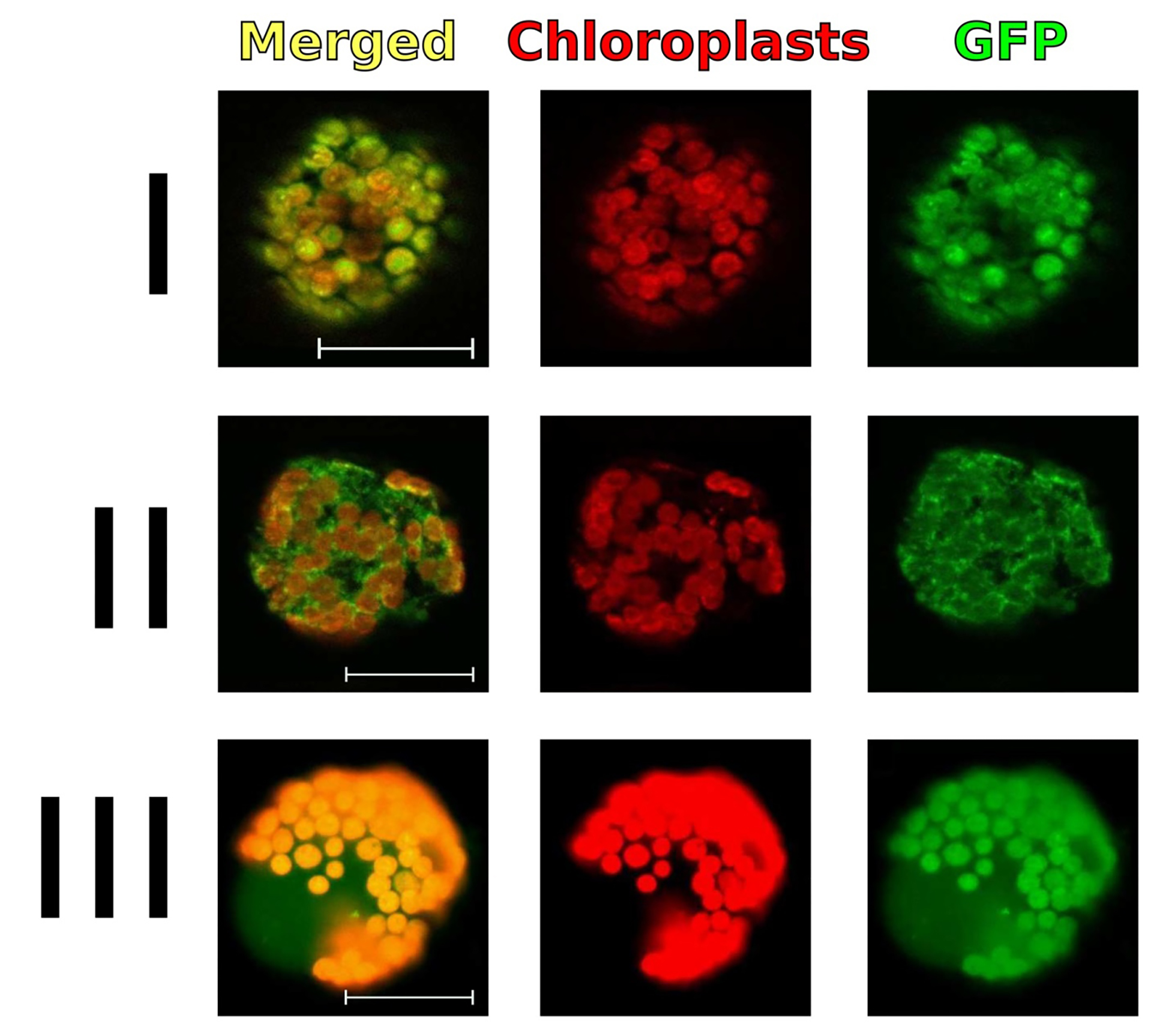
Figure 3. Results of analysis of subcellular localization of GFP fusion proteins in tobacco protoplasts isolated from the cells of an agroinfiltrated leaf area. Transient gene expression of plant expression vectors, with leader signal sequences providing their localization in various compartments of plant cells, demonstrates that GFP fusion protein is localized to the chloroplasts (I), endoplasmic reticulum (II), and cytoplasm (III). The merged images (first column) include the green channel (last column) and the chloroplast autofluorescence channel (middle column). Scale bars = 10 μm.
Data analysis
All experiments were performed in at least three independent replicates, each replicate being analyzed at least three times. To determine how well transformation was done we checked the level of GFP fluorescence in agroinfiltrated leaves of N. benthamiana in UV light. For estimation of subcellular localization of GFP fusion proteins in the tobacco protoplasts, we analyzed about 100 protoplasts and visually determined positive transient gene expression in at least 80% of protoplasts.
Notes
This protocol can be applied to other plant species (Figure 4).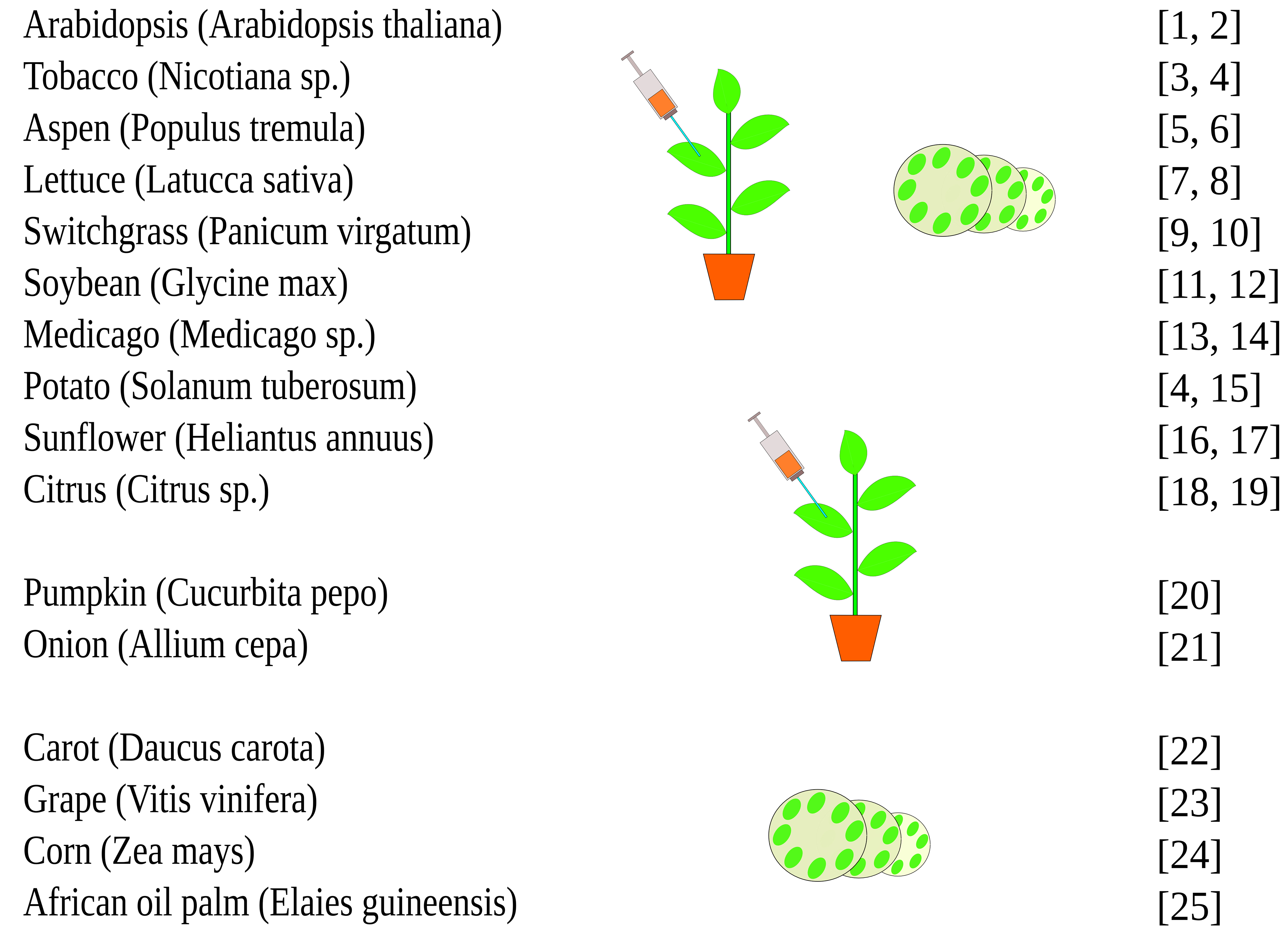
Figure 4. Potential application of the AgI–PrI technique to different plant species (Tyurin et al., 2017, supplementary data)
Recipes
- Solution 1
150 mМ NaCl
10 mМ Tris-HCl (pH 7.0) - Solution 2
20 mМ CaCl2
10 mМ Tris-HCl (pH 7.0) - Solution 3
20 μM acetosyringone
10 mМ MES - Solution 4
200 mg/L MgSO4·7H2O
100 mg/L CaCl2·2H2O
150 mg/L NH4H2PO4
0.4 M sorbitol
4 mМ CaCl2
12.5 mМ MES-KOH (pH 5.7)
1% Cellulase Onozuka R10
0.15% Pectinase Macerozyme R10
0.4% Driselase - Solution 5
0.5 М sorbitol
2.5 mМ CaCl2 - Knop’s solution 1 L
1 g Са(NО3)2
0.25 g KH2РO4
0.25 g MgSO4
0.125 g KСl
0.0125 g FеСl3 - Agroinfiltration buffer
1x MS medium
10 mM MES-KOH (pH 5.6)
2% sucrose
200 μM acetosyringone
Acknowledgments
This protocol has been adapted from an earlier study (Tyurin et al., 2017). The authors confirm that there are no known conflicts of interest associated with this publication.
References
- Mohamed, Sh., Boehm, R., Binsfeld, P. C. and Schnabl, H. (2004). Agrobacterium-mediated transformation of two high oleic sunflower (Helianthus annuus L.) genotypes assessment and optimization of important parameters. Helia 27(40): 25-40.
- Nosov, А. V., Fomenkov, A. A., Mamaeva, A. S., Solovchenko, A. E., Novikova, G. V. (2014). Extra perspectives of 5-ethynyl-2’-deoxyuridine click reaction with fluorochrome azides to study cell cycle and deoxyribonucleoside metabolism. Russ J Plant Physiol 61(6): 899-909.
- Sheludko, Y. V, Sindarovska, Y. R., Gerasymenko, I. M., Bannikova, M. A. and Kuchuk, N. V. (2007). Comparison of several Nicotiana species as hosts for high-scale Agrobacterium-mediated transient expression. Biotechnol Bioeng 6(3): 608-14.
- Tyurin, A. A., Kabardaeva, K. V., Berestovoy, M. A., Sidorchuk, Yu. V., A. Fomenkov, A. A., Nosov, A. V., Goldenkova-Pavlova, I. V. (2017). Simple and reliable system for transient gene expression for the characteristic signal sequences and the estimation of the localization of target protein in plant cell. Russ J Plant Physiol 64(5): 672-679.
Article Information
Copyright
© 2018 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Berestovoy, M. A., Tyurin, A. A., Kabardaeva, K. V., Sidorchuk, Y. V., Fomenkov, A. A., Nosov, A. V. and Goldenkova-Pavlova, I. V. (2018). Transient Gene Expression for the Characteristic Signal Sequences and the Estimation of the Localization of Target Protein in Plant Cell. Bio-protocol 8(4): e2738. DOI: 10.21769/BioProtoc.2738.
Category
Plant Science > Plant physiology > Plant growth
Molecular Biology > DNA > Transformation
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link









