- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Co-sedimentation Assay for the Detection of Direct Binding to F-actin
Published: Vol 2, Iss 19, Oct 5, 2012 DOI: 10.21769/BioProtoc.270 Views: 21474

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
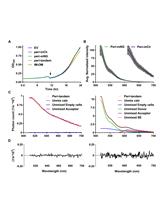
Detection of Protein Interactions in the Cytoplasm and Periplasm of Escherichia coli by Förster Resonance Energy Transfer
Nils Y. Meiresonne [...] Tanneke den Blaauwen
Jan 20, 2018 14487 Views
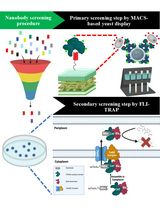
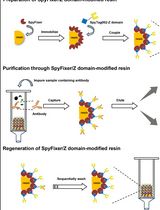
Isolation of Antigen-Specific Nanobodies From Synthetic Libraries Using a Protein Selection Strategy That Combines MACS-Based Screening of YSD and FLI-TRAP
Apisitt Thaiprayoon [...] Dujduan Waraho-Zhmayev
Jan 20, 2026 537 Views
Abstract
This protocol describes measurement of direct protein-protein interactions by actin co-sedimentation assay. The actin co-sedimentation assay is a well-established technique and has been commonly used to demonstrate binding of proteins that interact directly with actin filaments (Ahrens et al., 2012; Mehta and Sibley, 2010; Schuler et al., 2005; Singh et al., 2011). We and others have previously shown that the damaged cell-recognition molecule C-type lectin 9A (Clec9A) recognises a conserved component within nucleated and non-nucleated cells that is exposed only when the cell membrane is damaged (Srivastava and Barber, 2008; Zhang et al., 2012). Here we use Clec9A as an example and present a detailed procedure for demonstrating the direct binding of purified recombinant Clec9A ectodomain to actin filaments.
Keywords: ActinMaterials and Reagents
- Rabbit muscle G-actin (Cytoskeleton, catalog number: AKL95-C )
- Recombinant Clec9A ectodomain was expressed and purified by affinity chromatography and size-exclusion chromatography as described previously 5
- Tris
- CaCl2
- ATP
- β-mercaptoethanol
- KCl
- MgCl2
- NuPAGE Novex 4-12% Bis-Tris gel (Life Technologies, Invitrogen™, catalog number: NP0327BOX )
- Laemmli reducing SDS-PAGE sample buffer (Laemmli 2x concentrate) (Sigma-Aldrich, catalog number: S3401 )
- Precision Plus Protein molecular weight marker (Bio-Rad Laboratories, catalog number: 161-0373 )
- Coomassie Brilliant Blue
- 1x G-buffer (see Recipes)
- 10x F-buffer (see Recipes)
Equipment
- Superdex S200 10/300 GL column (GE Healthcare)
- AKTA Fast Protein Liquid Chromatography (FPLC) system (GE Healthcare)
- TLA 100 rotor and Beckman Coulter Optima TL Ultracentrifuge (Beckman Coulter)
- Polycarbonate centrifuge tubes (7 x 20 mm, 230 μl) (Beckman Coulter, catalog number: 343775 )
- GS-800 calibrated densitometer and Quantity one image analysis software (Transmission densitometer) (Bio-Rad Laboratories, catalog number: 1707980 )
Procedure
- Preparation of actin
- Reconstitute vial of lyophilised powder of rabbit muscle G-actin in water (2 mg/ml) and incubate protein for 1/2 h at room temperature (RT).
Note: If making up a large stock of actin, freeze in small, single use aliquots at -80 °C. - Pre-equilibrate a Superdex S200 10/300 GL column with a minimum of 35 ml of G-buffer.
Note: G-buffer is used as both pre-elution and elution buffer. - Load 0.5 ml of actin (2 mg/ml) onto the Superdex S200, and run column in G-buffer at a flow rate of 0.5 ml/min, and collect column fractions corresponding to elution volumes of 12-15 ml for analysis.
- Analyse fractions reflecting the peak of actin protein (G-actin molecular mass is 42-43 kDa) by SDS-PAGE on a NuPAGE Novex 4-12% Bis-Tris gel under reducing conditions, and visualise proteins by Coomassie Brilliant Blue staining to confirm purity of G-actin.
- Pool fractions reflecting the peak of G-actin.
- Polymerise G-actin (pooled monomeric G-actin from step 1e) to F-actin by adding 1/10 volume of 10x F-buffer and incubating for 1 h at RT.
- Reconstitute vial of lyophilised powder of rabbit muscle G-actin in water (2 mg/ml) and incubate protein for 1/2 h at room temperature (RT).
- Assay for actin filament binding
- Use the actin filaments generated in step A-6 to set up a series of reaction tubes in polycarbonate ultracentrifuge tubes with a range of concentrations of actin filaments (0, 1, 2, 5 μM) and add a fixed amount of potential actin-interacting protein (Clec9A; 5 μM) or a control protein that does not interact with actin (5 μM) in a total volume of 50 μl. As a control, also include 1 tube of actin alone.
Note: The concentration of actin filaments used for each reactions should be based on the initial concentration of G-actin in step A-5. - Incubate reactions for 1 h at RT.
- Sediment actin filaments and associated proteins by ultracentrifugation at 100,000 x g using a fixed angle rotor for 1 h at RT.
- Transfer supernatant to a fresh tube.
- Wash pellet by adding 50 μl of 1x F-buffer (diluted in G-buffer) and sediment actin filaments and associated proteins by ultracentrifugation at 100,000 x g for 1 h at RT.
- Remove supernatant and resuspend pellet in 1x Laemmli reducing SDS-PAGE sample buffer (equal volume to supernatant).
- Set up tubes containing an equal volume of supernatant (from step B-4) with SDS-PAGE sample buffer, or resuspended pellet in SDS-PAGE sample buffer (from step B-6) for each of the reactions of actin +/- actin binding proteins, then incubate samples at 95 °C for 5 min.
- Analyse samples by SDS-PAGE and visualise proteins by Coomassie Brilliant Blue staining.
- Quantitate the amount of Clec9A in the supernatant and pellet fractions by densitometry analysis using a GS-800 calibrated densitometer.
Note: It is important to note that as the samples are being analysed under reducing conditions, the sizes of the bands do not change on SDS-PAGE. What changes is the shift of the bands from supernatant to pellet as a result of F-actin cosedimentation. - Calculate the intensity of the Clec9A bands in the pellet, and the intensity of the total amount of Clec9A (pellet + supernatant). Then calculate the proportion of Clec9A co-sedimenting with F-actin as the ratio of Clec9A in the pellet relative to the total Clec9A (Clec9A in the Supernatant + Pellet), that is, the intensity of Clec9A in pellet divided by the intensity of Clec9A in (supernatant + pellet). The proportion of Clec9A co-sedimenting with F-actin relative to a nonspecific binding control is interpreted as the specific binding to F-actin.
- Use the actin filaments generated in step A-6 to set up a series of reaction tubes in polycarbonate ultracentrifuge tubes with a range of concentrations of actin filaments (0, 1, 2, 5 μM) and add a fixed amount of potential actin-interacting protein (Clec9A; 5 μM) or a control protein that does not interact with actin (5 μM) in a total volume of 50 μl. As a control, also include 1 tube of actin alone.
Recipes
- 1x G-buffer
20 mM Tris (pH 8)
0.1 mM CaCl2
0.1 mM ATP
5 mM β-mercaptoethanol - 10x F-buffer
1 M KCl
20 mM ATP
20 mM MgCl2
Acknowledgments
Jian-Guo Zhang was supported by Australian Government NHMRC Program Grants 461219 and 1016647. His work was also made possible through Victorian State Government Operational Infrastructure Support and Australian Government NHMRC IRIISS.
References
- Ahrens, S., Zelenay, S., Sancho, D., Hanc, P., Kjaer, S., Feest, C., Fletcher, G., Durkin, C., Postigo, A., Skehel, M., Batista, F., Thompson, B., Way, M., Reis e Sousa, C. and Schulz, O. (2012). F-actin is an evolutionarily conserved damage-associated molecular pattern recognized by DNGR-1, a receptor for dead cells. Immunity 36(4): 635-645.
- Mehta, S. and Sibley, L. D. (2010). Toxoplasma gondii actin depolymerizing factor acts primarily to sequester G-actin. J Biol Chem 285(9): 6835-6847.
- Schuler, H., Mueller, A. K. and Matuschewski, K. (2005). A Plasmodium actin-depolymerizing factor that binds exclusively to actin monomers. Mol Biol Cell 16(9): 4013-4023.
- Singh, B. K., Sattler, J. M., Chatterjee, M., Huttu, J., Schuler, H. and Kursula, I. (2011). Crystal structures explain functional differences in the two actin depolymerization factors of the malaria parasite. J Biol Chem 286(32): 28256-28264.
- Srivastava, J. and Barber, D. (2008). Actin co-sedimentation assay; for the analysis of protein binding to F-actin. J Vis Exp(13).
- Zhang, J. G., Czabotar, P. E., Policheni, A. N., Caminschi, I., Wan, S. S., Kitsoulis, S., Tullett, K. M., Robin, A. Y., Brammananth, R., van Delft, M. F., Lu, J., O'Reilly, L. A., Josefsson, E. C., Kile, B. T., Chin, W. J., Mintern, J. D., Olshina, M. A., Wong, W., Baum, J., Wright, M. D., Huang, D. C., Mohandas, N., Coppel, R. L., Colman, P. M., Nicola, N. A., Shortman, K. and Lahoud, M. H. (2012). The dendritic cell receptor Clec9A binds damaged cells via exposed actin filaments. Immunity 36(4): 646-657.
Article Information
Copyright
© 2012 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Wong, W., Zhang, J., Baum, J., Lahoud, M. H. and Shortman, K. (2012). Co-sedimentation Assay for the Detection of Direct Binding to F-actin. Bio-protocol 2(19): e270. DOI: 10.21769/BioProtoc.270.
Category
Biochemistry > Protein > Interaction > Protein-protein interaction
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link