- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Laminarin Quantification in Microalgae with Enzymes from Marine Microbes
Published: Vol 8, Iss 8, Apr 20, 2018 DOI: 10.21769/BioProtoc.2666 Views: 8050
Reviewed by: Anonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
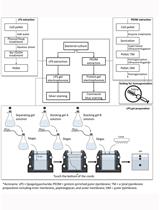
Extraction and Electrophoretic Analysis of Bacterial Lipopolysaccharides and Outer Membrane Proteins
Yue-Jia Lee and Thomas J. Inzana
Dec 20, 2021 5791 Views
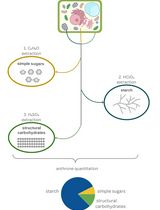
A Simple, Rapid, and Cost-Effective Method for Assessing Carbohydrate Partitioning in Microalgae and Arabidopsis thaliana
Araceli N. Bader [...] Leonardo Curatti
Dec 5, 2024 2373 Views
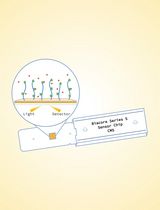
Surface Plasmon Resonance for the Interaction of Capsular Polysaccharide (CPS) With KpACE
Zhe Wang [...] Chao Cai
Jun 20, 2025 3577 Views
Abstract
The marine beta-glucan laminarin is an abundant storage polysaccharide in microalgae. High production rates and rapid digestion by heterotrophic bacteria turn laminarin into an ideal carbon and energy source, and it is therefore a key player in the marine carbon cycle. As a main storage glucan laminarin also plays a central role in the energy metabolism of the microalgae (Percival and Ross, 1951; Myklestad, 1974; Painter, 1983). We take advantage of enzymes that digest laminarin selectively and can thereby quantify only this polysaccharide in environmental samples. These enzymes hydrolyze laminarin into glucose and oligosaccharides, which are measured with a standard reducing sugar assay to obtain the laminarin concentration. Prior to this assay, the three enzymes need to be produced via heterologous expression and purification. The assay can be used to monitor laminarin concentrations in environmental microalgae, which were concentrated from seawater by filtering, or in samples derived from algal lab cultures.
Keywords: AlgaeBackground
Marine polysaccharides play an important role in the marine carbon cycle and are a major part of the physiology of phytoplankton, but are severely understudied. For decades, the agro-food industry has been using ready-to-use kits based on enzymatic assays to analyse a wide range of different polysaccharides in their processes (Whitaker, 1974). These fast, robust and specific enzyme based methods assess polysaccharides originating from land-based plants, i.e., starch, as they are widely used in food, feed and other industrial applications (Brunt et al., 1998). However, similar assays for marine polysaccharides are still lacking. Inspired by the idea of using enzymes for polysaccharide quantification in algae, we developed an enzyme-based method to quantify the ecologically relevant beta-glucan laminarin, also known as chrysolaminarin, in diatoms and other microalgae.
The three glycoside hydrolases (GH) for this application are from Formosa spp. and they were characterized as follows: FbGH30 is an exo-acting β-1,6-glucanase of the GH30 family, specifically hydrolysing the β-1,6-linked glucose monomer branches attached to the laminarin backbone; and FaGH17A and FbGH17A are two endo-acting β-1,3-glucanases of the GH family 17, which acts specifically on the β-1,3-linked laminarin backbone (Becker et al., 2017)
This method enables the quantification of laminarin in crude substrate mixtures, without the need for purification of the laminarin. This enzymatic method is fast, does not require sophisticated instruments, the enzymes are stereospecific and they selectively cleave laminarin into glucose and oligosaccharides, which can be quantified with a common reducing sugar assay. The method can be easily applied in fieldwork. The assay itself comprises only the three steps of extraction, hydrolysis and the reducing sugar assay (Figure 1). It can be done within only a few hours. The limit of detection (LOD) of the assay is at 1.5 µg/ml. The three enzymes need to be produced only once and can be stored for years. After their production and purification, one has enough material to analyse thousands of samples. We decided to include the plasmid transformation and recombinant enzyme production part into the protocol, since we consider these steps feasible to be done by marine labs with less experience in biotechnology.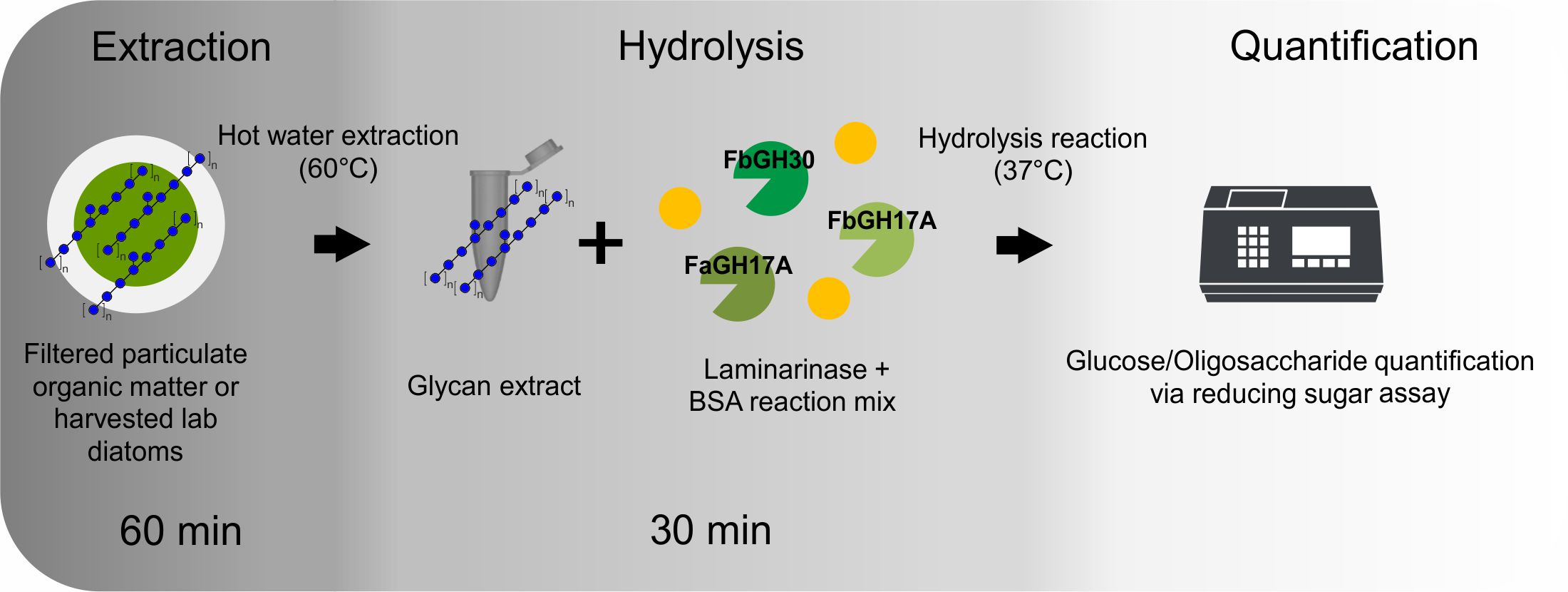
Figure 1. Schematic protocol after the production of the enzymes. A brief outline of the three main steps and their approximate duration.
Materials and Reagents
- 2.0 ml reaction tubes (SARSTEDT, catalog number: 72.691 )
- Sterile toothpick
- Aluminium foil
- 5 ml HiTrap IMAC HP column (GE Healthcare, catalog number: 17-0920-05 )
- Polyethersulfone ultrafiltration membrane with 10-kDa cutoff value (Merck, catalog number: PBGC04310 )
- 5 ml HiTrap Desalting column (GE Healthcare, catalog number: 29-0486-84 )
- Whatman GF/F filters (GE Healthcare, catalog number: 1825-047 ), size is dependent on used filtration setup
- 50 ml centrifuge tubes (SARSTEDT, catalog number: 62.547.254 )
- Half-micro-cuvettes (BRAND, catalog number: 759015 )
- Petri dishes (SARSTEDT, catalog number: 82.1473 )
Note: Alternatively a Pierce Protein Concentrator PES, 10 K MWCO, 5-20 ml (Thermo Fisher Scientific, Thermo ScientificTM, catalog number: 88527 ) (see Note 2). - BL21 (DE3) competent E. coli (New England Biolabs, catalog number: C2527I )
- SOC-medium (is usually enclosed in BL21 (DE3) competent E. coli product)
- FaGH17A plasmid (Addgene, catalog number: 86462 ) (see Note 1)
- FbGH30 plasmid (Addgene, catalog number: 86463 ) (see Note 1)
- FbGH17A plasmid (Addgene, catalog number: 100911 ) (see Note 1)
- IPTG (Isopropyl β-D-thiogalactopyranoside from AppliChem, catalog number: A4773 )
- Lysozyme (from chicken egg white) (Sigma-Aldrich, catalog number: L6876 )
- Sodium chloride (NaCl) (Carl Roth, catalog number: 3957.2 )
- Tryptone (Sigma-Aldrich, catalog number: 95039 )
- Yeast extract (Sigma-Aldrich, catalog number: 09182 )
- Agar (Sigma-Aldrich, catalog number: A7002 )
- Kanamycin sulfate (Sigma-Aldrich, catalog number: 60615 )
- Ammonium sulfate ((NH4)2SO4) (Sigma-Aldrich, catalog number: A4418 )
- Monopotassium phosphate (KH2PO4) (Sigma-Aldrich, catalog number: 9791 )
- Disodium phosphate (Na2HPO4) (Sigma-Aldrich, catalog number: 71642 )
- Glucose (Sigma-Aldrich, catalog number: G8270 )
- Lactose (Sigma-Aldrich, catalog number: 17814 )
- Glycerol (Sigma-Aldrich, catalog number: G6279 )
- Magnesium sulfate (MgSO4) (Sigma-Aldrich, catalog number: 208094 )
- Sucrose (Sigma-Aldrich, catalog number: 84100 )
- Tris(hydroxymethyl)aminomethane (Tris) (Sigma-Aldrich, catalog number: RDD008 )
- Sodium deoxycholate (Sigma-Aldrich, catalog number: D6750 )
- Triton X-100 (Sigma-Aldrich, catalog number: X100 )
- DNase I (from bovine pancreas) (Sigma-Aldrich, catalog number: 69182 )
- Nickel(II) sulfate hexahydrate (NiSO4·6H2O) (Sigma-Aldrich, catalog number: 227676 )
- Imidazole (Sigma-Aldrich, catalog number: I5513 )
- Dithiothreitol (DTT) (Sigma-Aldrich, catalog number: D0632 )
- 3-(N-Morpholino)propanesulfonic acid (MOPS) (Sigma-Aldrich, catalog number: 69947 )
- Laminarin from Laminaria digitata (Sigma-Aldrich, catalog number: L9634 )
- Bovine serum albumin (BSA), lyophilized (Sigma-Aldrich, catalog number: A2153 )
- 4-Hydroxybenzhydrazide (Sigma-Aldrich, catalog number: H9882 )
- HCl (37%) (Sigma-Aldrich, catalog number: 320331 )
- Trisodium citrate dihydrate (Sigma-Aldrich, catalog number: W302600 )
- Sodium hydroxide (NaOH) (VWR, catalog number: 28244.295 )
- Calcium chloride (CaCl2)
- LB-Kana-Agar-Plates (see Recipes)
- LB-Kana-Medium (see Recipes)
- 20x NPS stock (see Recipes)
- 50x 5052 stock (see Recipes)
- MgSO4 stock (see Recipes)
- ZY-Medium (see Recipes)
- ZYP-5052-Rich-Autoinduction-Medium (see Recipes)
- Sucrose stock (see Recipes)
- Deoxycholate stock (see Recipes)
- DNase I stock (see Recipes)
- NiSO4 stock (see Recipes)
- IMAC buffer A (see Recipes)
- IMAC buffer B (see Recipes)
- SEC buffer (see Recipes)
- SEC/DTT buffer (see Recipes)
- MOPS buffer (see Recipes)
- BSA stock solution (see Recipes)
- PAHBAH reagent A (see Recipes)
- PAHBAH reagent B (see Recipes)
- PAHBAH working reagent (see Recipes)
Equipment
- Bunsen burner or sterile hood
- 2 L Erlenmeyer flasks (or normal 2 L glass bottles)
- Rotating shaker for Erlenmeyer flasks (temperature control is recommended)
- Vortex mixer
- Microcentrifuge for 1.5 or 2.0 ml reaction tubes with cooling function, i.e., Centrifuge 5418 R (Eppendorf, model: 5418 R , catalog number: 5401000013)
- Centrifuge for 15 and 50 ml tubes with cooling function, i.e., Centrifuge 5804 R (Eppendorf, model: 5804 R , catalog number: 5805000017)
- Standard membrane or peristaltic pump and respective filtration setup, i.e., ME 1 (Vacuubrand, model: ME 1 ) and Nalgene filter holder receiver (Thermo Fisher Scientific, Thermo ScientificTM, catalog number: 300-4100 )
- Standard fast protein liquid chromatography system (FPLC), i.e., ÄKTA start (GE Healthcare, model: ÄKTA start, catalog number: 29-0220-94 )
Note: Alternatively a peristaltic pump with a flow rate of ≤ 5 ml/min. - Spectrometer, i.e., BioSpectrometer basic (Eppendorf, model: BioSpectrometer® basic , catalog number: 6135000009)
- Heated water bath, i.e., Thermolab (GFL, catalog number: 1070 )
- Two heat blocks, i.e., BioShake iQ (Analytik Jena, catalog number: 848-1808-0506 )
- Autoclave
Note: Autoclave is strongly recommended but not essential, since one always uses the antibiotic kanamycin for the bacterial growth. - Ultrafiltration stirred cell, i.e., Amicon (Merck, catalog number: UFSC05001 )
Note: Alternatively a Pierce Protein Concentrator PES, 10 K MWCO, 5-20 ml (Thermo Fisher Scientific, Thermo ScientificTM, catalog number: 88527 ) (see Note 2).
Procedure
- Transformation
- Transform each Addgene plasmid containing one enzyme into separate BL21 (DE3) competent E. coli according to the manufacturer’s instructions and by using LB-Kana-Agar-Plates (see Recipes). A common transformation protocol uses the following steps, which have to be carried out next to a Bunsen burner or in a sterile hood.
- Add 1 µl containing ca. 100 ng of plasmid DNA to the cell mixture and carefully invert the tube once. Do not vortex.
- Keep on ice for 5 min and do not mix.
- Heat shock at exactly 42 °C for exactly 10 sec and do not mix.
- Keep on ice for 5 min and do not mix.
- Add 500 µl SOC-medium, which is included in the E. coli BL21 (DE3) shipment.
- Incubate at 37 °C for 60 min and at 500 rpm.
- Spread 100 µl on an LB-Kana-Agar-Plate and incubate for at least 15 h at 37 °C.
- After picking a colony with a sterile toothpick and growing it for at least 8 h in a 3 ml LB-Kana-Medium (see Recipes) preculture at 37 °C and under constant stirring at 150 rpm, the cells can be stored at -80 °C after adding 25% (v/v) sterile glycerol (i.e., 200 µl of glycerol in 600 µl of preculture).
- Transform each Addgene plasmid containing one enzyme into separate BL21 (DE3) competent E. coli according to the manufacturer’s instructions and by using LB-Kana-Agar-Plates (see Recipes). A common transformation protocol uses the following steps, which have to be carried out next to a Bunsen burner or in a sterile hood.
- Enzyme overexpression and purification
- Use 1 ml of the preculture from Step A9 to inoculate 1 L of ZYP-5052-Rich-Autoinduction-Medium (see Recipes) (Studier, 2005) in a 2 L Erlenmeyer flask to have enough headspace. Close the flask only with aluminum foil to allow gas exchange into the flask.
Notes:- Overexpression and purification are performed in almost the exact same way for all three enzymes (FbGH30, FaGH17A and FbGH17A). However, each purification has to be performed separately. It is not possible to transform all three enzymes into one expression system and purify all together. Nonetheless, after the overexpression, it is possible to purify all three enzymes within one working day.
- Overexpression can also be performed in IPTG-induced LB-Kana-Medium. The inoculated cultures need to grow for 8 h at 37 °C before 1 ml 1 M IPTG is added. After induction, the cultures need to be incubated for 8 h more at 16 °C. Subsequent harvesting and purification are performed as already described. This method is faster but yields lower protein amounts.
- Overexpression and purification are performed in almost the exact same way for all three enzymes (FbGH30, FaGH17A and FbGH17A). However, each purification has to be performed separately. It is not possible to transform all three enzymes into one expression system and purify all together. Nonetheless, after the overexpression, it is possible to purify all three enzymes within one working day.
- Grow cultures for about 72 h at 20 °C [or room temperature (RT)] and with rotation at 150 rpm.
- Harvest cells by centrifugation at 4,500 x g for 25 min at 4 °C. Discard supernatant. The resulting pellet can be stored at -20 °C (for years) until further use.
- Resuspend the cell pellet in 15 ml sucrose stock solution (see Recipes).
- Add 5 mg of lysozyme and incubate for 10 min at RT under constant magnetic stirring at 500 rpm.
- Add 30 ml deoxycholate stock solution (see Recipes) and 0.2 ml MgSO4 stock solution (see Recipes).
- Add 1 ml DNase I stock solution (see Recipes) to liquefy the viscous solution.
- Centrifuge the lysate at 16,000 x g for 45 min at 4 °C. Transfer the supernatant containing the proteins of interest into a new tube.
- All steps during the following immobilized metal affinity chromatography (IMAC) are carried out with an FPLC at RT and a flow rate of 5 ml/min. Alternatively, the procedure can also be performed by using a peristaltic pump at the same or lower flow rates.
Note: Steps B10-B14 can be performed manually using a peristaltic pump at 5 ml/min flow rate instead of an FPLC system. The column is then placed after the pump. The linear buffer gradient in Step B14 can be achieved similarly by using a stepwise increase of the imidazole concentration over the course of the total elution volume of 5 CV (~5 min). Starting with IMAC buffer A containing 50 mM imidazole, the buffer needs to be changed every 30 sec. At each step the imidazole concentrations increases by 50 mM (30 sec each): 50, 100, 150, 200, 250, 300, 350, 400, 450, 500 mM imidazole (the usual concentration for IMAC buffer A and B are 25 and 500 mM imidazole, respectively). To prevent air bubbles on the column, the flow needs to be paused every time the inlet is switching from one buffer to the other. - Charge a 5 ml HiTrap IMAC HP column with 1 column volume (1 CV = 1 ml) of NiSO4 stock solution (see Recipes). Discard the flow that comes off the column.
- Equilibrate with 5 CVs of IMAC buffer A (see Recipes). Discard the flow that comes off the column.
- Inject the entire supernatant of the lysis onto the column. If the solution is too viscous, it needs to be diluted up to a volume of 50 ml using IMAC buffer A. Discard the flow that comes off the column.
- Wash the column with 15 CVs IMAC buffer A. Discard the flow that comes off the column.
- Elute the protein in a total number of 5 CVs of a continuous linear gradient from IMAC buffer A to IMAC buffer B (see Recipes). Collect the flow that comes off the column in thirty 1 ml fractions.
- For FbGH17A only: Add 50 µl of the reducing agent DTT (0.5 mg/ml) to each fraction immediately after the elution to prevent oxidation of the protein.
- Pool fraction 6 to 30 (if there is a UV detector, take only the samples corresponding to the major peak). Keep the protein on ice.
- The IMAC column has to be cleaned after each purification according to the manufacturer’s instruction.
- Assemble an ultrafiltration stirred cell together with a respective 10 kDa membrane. (alternatively, a centrifugation concentrator can be used, see Note 2).
- Concentrate the protein solution in the ultrafiltration chamber at 4 °C or on ice down to a volume of ≤ 1.5 ml.
- Centrifuge the protein solution for 20 min at 13,000 x g and 4 °C, to remove the precipitated, inactive protein.
- Transfer supernatant into a fresh tube and discard the pellet. Keep the protein on ice.
- The following use of a desalting column removes the imidazole. The procedure followed the manufacturer’s instructions and can be performed by using an FPLC system or a syringe at RT and a flow rate of 5 ml/min.
- For FbGH17A only: Use SEC/DTT buffer (see Recipes) instead of SEC buffer in Steps B24-26.
- Equilibrate a HiTrap desalting column with 5 CVs of SEC buffer. Discard the flow that comes off the column.
- Apply exactly 1.5 ml of the protein solution. Dilute the sample with SEC buffer if necessary. Do not exceed the sample volume of 1.5 ml. Discard the flow that comes off the column.
- Elute and collect the protein with 2 ml of SEC buffer and keep the protein on ice afterwards.
- The desalting column has to be cleaned after the purification according to the manufacturer’s instruction.
- Determine protein concentration by measuring the absorbance at 280 nm using a spectrometer and under consideration of the molecular weight and the extinction factor of each protein (FbGH30: MW = 54,700 Da, ε = 128,480 M-1 cm-1; FaGH17A: MW = 44,800 Da, ε = 87,905 M-1 cm-1; FbGH17A: MW = 46,600 Da, ε = 88,935 M-1 cm-1):
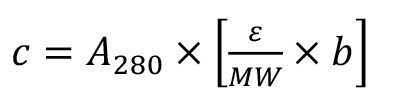
where, c: protein concentration in mg/ml; A: absorbance at 280 nm; ε: extinction factor in M-1 cm-1; MW: molecular weight in Da; b: light path length in cm.
Note: The success of overexpression (Steps B1-B3), lysis (Steps B4-B8), IMAC (Steps B9-B17), concentration (Steps B18-B21) and desalting (Steps B22-B27) can be verified by SDS-Polyacrylamide gel electrophoresis (He, 2011). - Dilute the proteins in MOPS buffer (see Recipes) to a stock concentration of 10 µM. Keep protein on ice.
- For FbGH17A only: After dilution in MOPS buffer the enzyme must be used immediately, either for preparing aliquots (Step B31) or for hydrolysis reactions (Steps D1-D7).
- Prepare ready-to-use 50 µl aliquots and freeze them at -20 °C.
Note: So far, there is no very long-term data on the shelf life of the enzymes at -20 °C, but they can be used for at least two years without any decrease in activity.
- Use 1 ml of the preculture from Step A9 to inoculate 1 L of ZYP-5052-Rich-Autoinduction-Medium (see Recipes) (Studier, 2005) in a 2 L Erlenmeyer flask to have enough headspace. Close the flask only with aluminum foil to allow gas exchange into the flask.
- Sampling and extraction
- Environmental samples, i.e., seawater, or lab cultures of microalgae are filtered onto Whatman GF/F filters at 0.1-0.5 bar by using a standard membrane or peristaltic pump and a respective filtration setup. The required volume needs to be determined by experiments. In case of sea surface water, one needs at least 5 L, for highly concentrated lab cultures the demand is much smaller (~50 ml, depending on the cell numbers). The volume must be logged for each sample. The resulting filter can be stored at -20 °C until further use.
Notes:- Polycarbonate filters can be used as well. The diameter of the filters depends on the filtration setup.
- For concentrated lab cultures, centrifugation can be used instead of filtration as well. Diatom cultures need to be centrifuged at 4,500 x g for 20 min.
- The method can be applied to macroalgal samples as well (L. Scheschonk et al. in preparation).
- Polycarbonate filters can be used as well. The diameter of the filters depends on the filtration setup.
- Place filter into a 15 or 50 ml centrifuge tube.
- Add 5-10 ml of MOPS buffer, just enough so that the entire filter is covered with liquid.
- Vortex every sample for 10 sec.
- Extract for 60 min at 60 °C in a heated water bath.
- Squeeze and take out the filter.
- Centrifuge for 15 min at 4,500 x g.
- Transfer the supernatant to a new tube. This extract can be stored for several weeks at -20 °C until further use.
Note: There is no long-term data on the stability of laminarin samples.
- Environmental samples, i.e., seawater, or lab cultures of microalgae are filtered onto Whatman GF/F filters at 0.1-0.5 bar by using a standard membrane or peristaltic pump and a respective filtration setup. The required volume needs to be determined by experiments. In case of sea surface water, one needs at least 5 L, for highly concentrated lab cultures the demand is much smaller (~50 ml, depending on the cell numbers). The volume must be logged for each sample. The resulting filter can be stored at -20 °C until further use.
- Hydrolysis
- Each sample is split up into six subsamples: three are hydrolyzed by the enzyme mixture and three are not hydrolyzed.
- Additionally, one has to prepare and treat (Steps C3 and C5-C7) samples for creating a calibration curve based on commercial laminarin from Laminarin digitata (Sigma-Aldrich). For microalgal samples, the following concentrations are suitable: 0, 7.8 µg/ml, 15.6 µg/ml, 31.3 µg/ml, 62.5 µg/ml, 125 µg/ml, 250 µg/ml, 0.5 mg/ml, 1.0 mg/ml, and 2.0 mg/ml (serial dilution in MOPS buffer).
- Thaw fresh aliquots of each enzyme stock solution and keep them on ice.
- Each hydrolyzation reaction mix consists of: 100 µl sample extract (or calibration standard), 1 µl FbGH30, 1 µl FaGH17A, 1 µl FbGH17A and 1 µl BSA stock solution (see Recipes) (see Note 3).
- Each non-hydrolyzation reaction mix consists of: 100 µl sample extract, 1 µl BSA stock solution and 3 µl of MOPS buffer (see Note 11).
- Mix all samples by shaking the reaction tubes.
- Incubate for 25 min at 37 °C in a heat block.
- Stop enzyme reaction by incubating the samples for 5 min at 99 °C in a second heat block. This extract can be stored for several weeks at -20 °C until further use.
Note: There is no long-term data on the stability of laminarin samples.
- Each sample is split up into six subsamples: three are hydrolyzed by the enzyme mixture and three are not hydrolyzed.
- Reducing sugar assay
- Add 1 ml of PAHBAH working reagent (see Recipes) (Lever, 1972) to 0.1 ml of the sample.
- Heat each sample for exactly 5 min at 100 °C in a heat block.
Note: The duration of boiling directly influences the absorbance and therefore the standard deviation of the measured triplicates. One should try to take the samples out of the heat block at the exact same speed that is used for putting them in. By doing this one can make sure that every sample is getting heated for the exact same duration. - Determine absorbance at 410 nm using half-micro-cuvettes and a spectrometer.
- Use PAHBAH working reagent as blank and for the dilution of samples that are too concentrated.
Note: The range of absorbance in which one can make reliable measurements depends on the spectrometer and must be checked in the manual in advance. The samples can be diluted in PAHBAH working reagent.
- Add 1 ml of PAHBAH working reagent (see Recipes) (Lever, 1972) to 0.1 ml of the sample.
Data analysis
- Each sample is measured in six subsamples. The value of the non-hydrolyzed triplicates needs to be subtracted from the value of the hydrolyzed samples.
- The laminarin concentration in the respective sample can be determined by comparing this difference to the calibration curve.
- The concentration, which was measured in the extract, needs to be converted into the concentration of the original sample, by taking the exact filtration volume or the cell number into account.
- Based on the calibration below, the equation and the data for laminarin concentrations in samples of 20 L are as follows (Figure 2, Table 1):
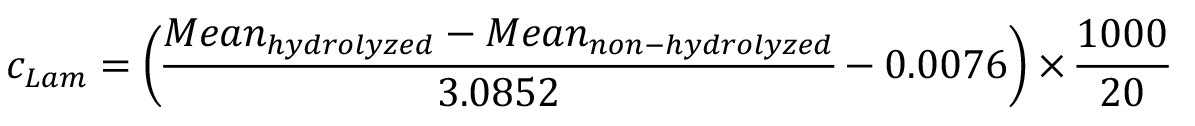
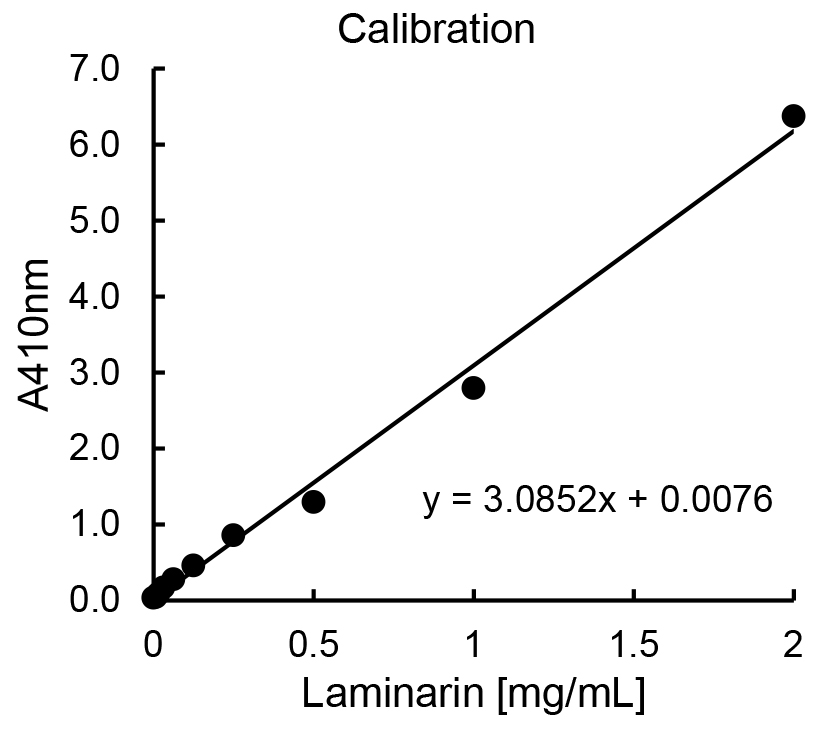
Figure 2. Exemplary calibration
Table 1. Exemplary data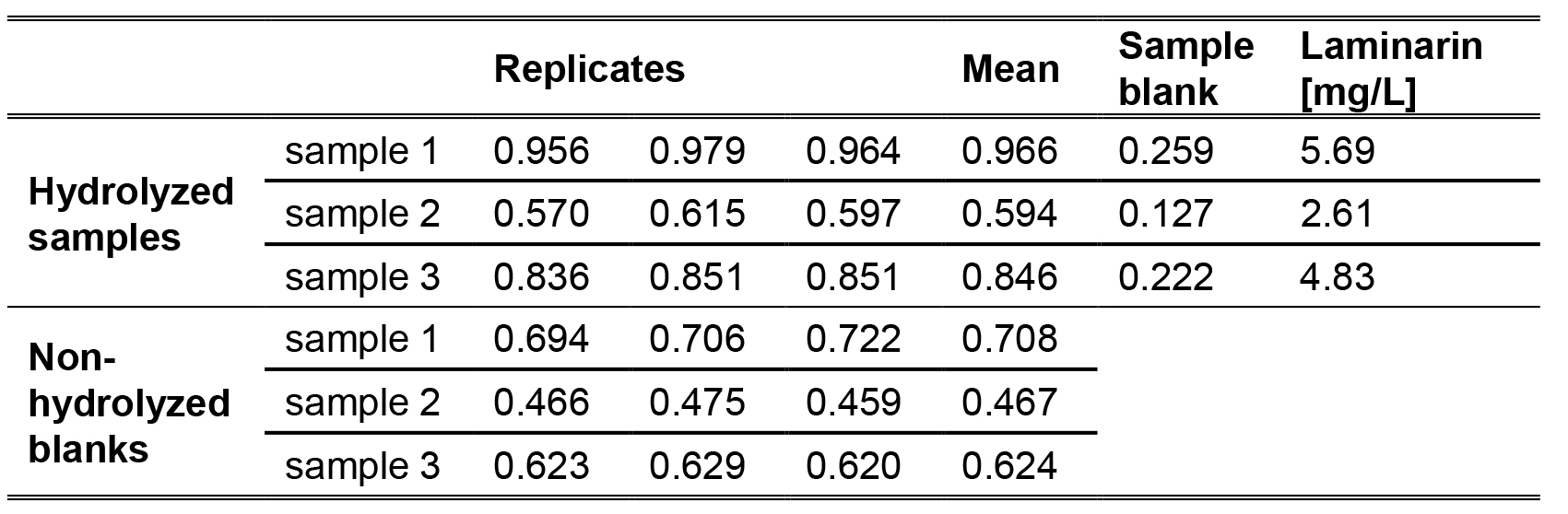
Notes
- All three plasmid can be ordered by academics and non-profits via the Addgene plasmid repository (www.addgene.org). On the website, one can find protocols for the purification of the plasmid DNA if Addgene provides only bacterial stabs. The plasmids must get transformed into E. coli BL21 (DE3).
- For protein concentration, a centrifugation-based system can be used alternatively. These tubes are not reusable and the recovery rate is quite low. However, for a single usage, it is cheaper compared to the ultrafiltration system. The concentration has to be conducted according to the manufacturer’s instruction and at 4 °C. The volume of the protein solution needs to be decreased to a volume of ≤ 1.5 ml.
- It is possible and advisable to prepare master mixes containing all components that have to be added to each sample extract.
Recipes
Note: Milli-Q water was used to make up the following solutions unless otherwise indicated.
- LB-Kana-Agar-Plates
- Dissolve:
10 g NaCl
10 g tryptone
5 g yeast extract
12.5 g agar
Complete the volume to 1 L with non-sterile water - Autoclave or boil the solution
- After letting the solution cool down to a temperature below 50 °C, add 1 ml of prepared kanamycin sulfate (50 mg/ml)
- After stirring the solution, the plates need to be poured next to a Bunsen burner or in a sterile hood
- When the plates are solid, they can be stored at 4 °C for several weeks
- Dissolve:
- LB-Kana-Medium
- Dissolve:
10 g NaCl
10 g tryptone
5 g yeast extract
Complete the volume to 1 L with non-sterile water in a 2 L Erlenmeyer flask or bottle - Close the flask with aluminum foil but keep it air-penetrable
- It is recommended to autoclave the solution
- After letting the solution cool down to a temperature below 50 °C, add 1 ml of prepared kanamycin sulfate (50 mg/ml)
- Dissolve:
- 20x NPS stock
Dissolve:
66 g (NH4)2SO4
136 g KH2PO4
142 g Na2HPO4
Complete the volume to 1 L with non-sterile water
It is recommended to autoclave the solution before use - 50x 5052 stock
Dissolve:
25 g glucose
100 g lactose
250 g glycerol (weigh in a beaker) and complete the volume to 1 L with non-sterile water
It is recommended to autoclave the solution before use - MgSO4 stock
Dissolve 12 g MgSO4 and complete the volume to 100 ml with non-sterile water
It is recommended to autoclave the solution before use - ZY-Medium
Dissolve 10 g tryptone and 5 g yeast extract in 928 ml H2O
It is recommended to autoclave the solution before use - ZYP-5052-Rich-Autoinduction-Medium
Mix 928 ml ZY-Medium
1 ml 1 M MgSO4
20 ml 50x 5052
50 ml 20x NPS and 2 ml kanamycin sulfate (50 mg/ml) next to a Bunsen burner - Sucrose stock
- Dissolve 250 g sucrose and 6 g Tris
- Adjust to pH 7.5 and complete the volume to 1 L with non-sterile water
- It is recommended to autoclave the solution before use
- Dissolve 250 g sucrose and 6 g Tris
- Deoxycholate stock
- Dissolve:
10 g sodium deoxycholate
10 ml Triton X-100
5.8 g NaCl
2.4 g Tris - Adjust to pH 7.5 and complete the volume to 1 L with non-sterile water
- It is recommended to autoclave the solution before use
- Dissolve:
- DNase I stock
Dissolve 50 mg DNase I in 35 ml IMAC buffer A and 15 ml glycerol
The solution can be stored at -20 °C - NiSO4 stock
Dissolve 3.9 g NiSO4 and complete the volume to 50 ml
Note: NiSO4 is harmful. Protect yourself by using gloves, eye and mouth protection. Discard contaminated solutions in an appropriate heavy metal waste. - IMAC buffer A
- Dissolve 29.2 g NaCl, 2.4 g Tris and 1.7 g imidazole
- Adjust to pH 7.5 and complete the volume to 1 L with non-sterile water
- It is recommended to sterile filter the solution before use
- Dissolve 29.2 g NaCl, 2.4 g Tris and 1.7 g imidazole
- IMAC buffer B
- Dissolve 29.2 g NaCl, 2.4 g Tris and 34 g imidazole
- Adjust to pH 7.5 and complete the volume to 1 L with non-sterile water
- It is recommended to sterile filter the solution before use
- Dissolve 29.2 g NaCl, 2.4 g Tris and 34 g imidazole
- SEC buffer
- Dissolve 29.2 g NaCl and 2.4 g Tris
- Adjust to pH 7.5 and complete the volume to 1 L with non-sterile water
- It is recommended to sterile filter the solution before use
- Dissolve 29.2 g NaCl and 2.4 g Tris
- SEC/DTT buffer
Dissolve 23 mg dithiothreitol in 50 ml SEC buffer - MOPS buffer
- Dissolve 5.2 g 3-(N-Morpholino)propanesulfonic acid
- Adjust to pH 7.0 and complete the volume to 500 ml
- It is recommended to sterile filter the solution before use
- Use a brown glass bottle and protect from light
- Dissolve 5.2 g 3-(N-Morpholino)propanesulfonic acid
- BSA stock solution
Dissolve 100 mg BSA in 1 ml of H2O
It is recommended to sterile filter the solution before use - PAHBAH reagent A
- Dissolve 10 g 4-hydroxybenzhydrazide in 60 ml H2O
- Add 10 ml concentrated HCl (37%) and complete the volume to 200 ml by adding more H2O
- Dissolve 10 g 4-hydroxybenzhydrazide in 60 ml H2O
- PAHBAH reagent B
- Dissolve 24.9 g trisodium citrate dehydrate in 500 ml H2O
- Add 2.2 g CaCl2 and mix
- Dissolve 40 g NaOH in a separate 1 L bottle
- Mix both solutions slowly and under constant stirring and complete the volume to 2 L with non-sterile water
- The solution must become clear within several minutes
- Dissolve 24.9 g trisodium citrate dehydrate in 500 ml H2O
- PAHBAH working reagent
Prepare a fresh 9:1 mixture of PAHBAH reagent B and PAHBAH reagent A
Can be used for several hours, but needs to be kept on ice
Acknowledgments
This protocol was adapted and modified from Becker et al., 2017. The research was supported by the Deutsche Forschungsgemeinschaft (grant HE 7217/1-1 to Jan-Hendrik Hehemann) and by the Max Planck Society.
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
References
- Becker, S., Scheffel, A., Polz, M. F. and Hehemann, J. H. (2017). Accurate quantification of laminarin in marine organic matter with enzymes from marine microbes. Appl Environ Microbiol 83(9).
- Brunt, K., Sanders, P. and Rozema, T. (1998). The enzymatic determination of starch in food, feed and raw materials of the starch industry. Starch - Stärke 50(10): 413-419.
- He, F. (2011). Laemmli-SDS-PAGE. Bio-protocol Bio101: e80.
- Lever, M. (1972). A new reaction for colorimetric determination of carbohydrates. Anal Biochem 47(1): 273-279.
- Myklestad, S. (1974). Production of carbohydrates by marine planktonic diatoms. I. Comparison of nine different species in culture. J Exp Mar Biol Ecol 15(3): 261-74.
- Painter, T. J. (1983). Algal polysaccharides. In: The polysaccharides. Elsevier 195-285.
- Percival, E. and Ross, A. (1951). 156. The constitution of laminarin. Part II. The soluble laminarin of laminaria digitata. J Chem Soc (Resumed) 0: 720-6.
- Studier, F. W. (2005). Protein production by auto-induction in high density shaking cultures. Protein Expr Purif 41(1): 207-234.
- Whitaker, J. R. (1974). Analytical applications of enzymes. In: Whitaker, J. R. (Ed.). Adv Chem Ser 136: 31-78.
Article Information
Copyright
© 2018 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Becker, S. and Hehemann, J. (2018). Laminarin Quantification in Microalgae with Enzymes from Marine Microbes. Bio-protocol 8(8): e2666. DOI: 10.21769/BioProtoc.2666.
Category
Biochemistry > Carbohydrate > Laminarin
Microbiology > Microbial biochemistry > Carbohydrate
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










