- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
In vitro NLK Kinase Assay
(*contributed equally to this work) Published: Vol 7, Iss 21, Nov 5, 2017 DOI: 10.21769/BioProtoc.2593 Views: 11369
Reviewed by: Ralph BottcherAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

Detection of ALT Associated Promyelocytic Leukemia Nuclear Bodies (APBs) by Immunofluorescence-FISH (IF-FISH)
Siamak A. Kamranvar and Maria G. Masucci
Dec 5, 2014 14468 Views
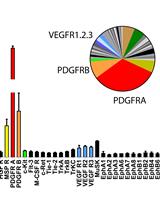
Unbiased Screening of Activated Receptor Tyrosine Kinases (RTKs) in Tumor Extracts Using a Mouse Phospho-RTK Array Kit
Julian Naipauer [...] Enrique A. Mesri
Apr 20, 2019 5697 Views
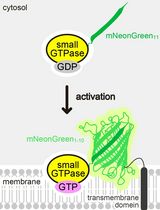
Detecting the Activation of Endogenous Small GTPases via Fluorescent Signals Utilizing a Split mNeonGreen: Small GTPase ActIvitY ANalyzing (SAIYAN) System
Miharu Maeda and Kota Saito
Jan 5, 2026 542 Views
Abstract
This protocol provides step by step instructions to perform an in vitro kinase assay for nemo-like kinase. In addition, this protocol also describes an efficient method using mild lysis buffer for expression and purification of Glutathione S-transferase (GST) fusion proteins.
Keywords: NLKBackground
Drosophila nemo has been identified as a pivotal regulator of photoreceptor clusters in the developing Drosophila eyes (Choi and Benzer, 1994). It has been reported that nemo is conserved across various species. The mammalian homolog is nemo-like kinase (NLK) (Brott et al., 1998; Rocheleau et al., 1999). NLK is a member of the CMGC group (CDK, MAPK, GSK3 and CLK) and also belongs to the atypical MAPK family. NLK regulates multiple cellular mechanisms through transcription factors including TCF/LEF1, NICD, and FOXO, which are key players in diverse signaling pathways (Ishitani et al., 1999; Ishitani et al., 2010; Kim et al., 2010). Moreover, recently we have identified NLK as a novel kinase for Yes-associated protein (YAP), a co-transcriptional activator in the Hippo pathway (Moon et al., 2017). This protocol describes an in vitro method to measure the nemo-like kinase activity.
Materials and Reagents
- Pipette tips–1,000 μl tips (OHAUS, catalog number: PCB-1000 ), 200 μl tips (Neptune Scientific, catalog number: 2100 )
- 14 ml round bottom tubes (SPL, catalog number: 40014 )
- 15 ml conical centrifuge tubes (Ihanil Scientific, catalog number: LP-00015CP-00 )
- 60 mm cell culture dish (SPL, catalog number: 20060 )
- Microcentrifuge tube (SARSTEDT, catalog number: 72.690 )
- X-ray film cassette (Advansta, catalog number: L-07019-001 )
- X-ray film (XSANATEC, AGFA, catalog number: EA75Q )
- Cell scraper (SARSTEDT, catalog number: 83.1832 )
- Centrifuge bottles (Sigma-Aldrich, catalog number: B1033 )
- pGEX-4T-3 vector (GE Healthcare, catalog number: 28-9545-52 )
- pFLAG-CMV4 vector (Sigma-Aldrich, catalog number: E7158 )
- Flag-NLK vectors (Ishitani et al., 1999)
- Competent cells-BL21 (DE3) (EMD Millipore, Novagen, catalog number: 70235 )
- Homo sapiens (Human) HEK293 cells (ATCC, catalog number: CRL-1573 )
- Zinc chloride (ZnCl2) (Sigma-Aldrich, catalog number: 229997 )
- Isopropyl β-D-1-thiogalactopyranoside (IPTG) (Bio Basic, catalog number: IB0168 )
- Glutathione Sepharose 4B (GE Healthcare, catalog number: 17075601 )
- Ethanol (Merk, catalog number: 1.00983.1011 )
- Bovine serum albumin (BSA) (Bio Basic, catalog number: AD0023 )
- Dulbecco’s modified Eagle medium (DMEM) (Lonza, catalog number: 12-604F )
- Fetal bovine serum (FBS) (Thermo Fisher Scientific, GibcoTM, catalog number: 16000044 )
- Antibiotic-Antimycotic (Thermo Fisher Scientific, GibcoTM, catalog number: 15240062 )
- TurboFect transfection reagents (Thermo Fisher Scientific, Thermo ScientificTM, catalog number: R0531 )
- Protein Assay Dye Reagent Concentrate (Bio-Rad Laboratories, catalog number: 5000006 )
- Anti-FLAG M2 antibody (Sigma-Aldrich, catalog number: F3165 )
- Protein A/G plus agarose (Santa Cruz Biotechnology, catalog number: sc-2003 )
- Adenosine 5’-triphosphate (ATP) [γ-32P] (PerkinElmer, catalog number: NEG502A )
- LB broth high salt (Duchefa Biochemie, catalog number: L1704 )
- Micro agar (Duchefa Biochemie, catalog number: M1002 )
- Ampicillin (Duchefa Biochemie, catalog number: A0104 )
- Tryptone (Duchefa Biochemie, catalog number: T1332 )
- Yeast extract (Duchefa Biochemie, catalog number: Y1333 )
- Sodium chloride (NaCl) (Duchefa Biochemie, catalog number: S0520 )
- Potassium chloride (KCl) (Duchefa Biochemie, catalog number: P0515 )
- Di-sodium hydrogen phosphate dihydrate (Na2HPO4·2H2O) (Duchefa Biochemie, catalog number: S0537 )
- Potassium phosphate dibasic (K2HPO4) (Merck, catalog number: 105108 )
- Tris (Duchefa Biochemie, catalog number: T1501 )
- NP-40 (IGEPAL CA-630) (Sigma-Aldrich, catalog number: I8896 )
- Ethylenediaminetetraacetic acid (EDTA) (Duchefa Biochemie, catalog number: E0511 )
- Ethylene glycol-bis(2-aminoethylether)-N,N,N’,N’,-tetraacetic acid (EGTA) (MP Biomedicals, catalog number: 194823 )
- β-Glycerophosphate (MP Biomedicals, catalog number: 195206 )
- Sodium fluoride (NaF) (Sigma-Aldrich, catalog number: S1504 )
- Dithiothreitol (DTT) (Bio Basic, catalog number: DB0058 )
- Phenylmethylsulfonyl fluoride (PMSF) (Sigma-Aldrich, catalog number: P7626 )
- Leupeptin (Sigma-Aldrich, catalog number: L2884 )
- Triton X-100 (Sigma-Aldrich, catalog number: T8787 )
- L-Glutathione reduced (Sigma-Aldrich, catalog number: G4251 )
- Methanol (Merck, catalog number: 106099 )
- Acetic acid (Merck, catalog number: 100056 )
- Coomassie Blue (Thermo Fisher Scientific, Thermo ScientificTM, catalog number: 20279 )
- HEPES (pH 7.5) (Duchefa Biochemie, catalog number: H1504.0100 )
- Magnesium chloride hexahydrate (MgCl2·6H2O) (Sigma-Aldrich, catalog number: M2670 )
- Sodium dodecyl sulfate (SDS) (Duchefa Biochemie, catalog number: S1377 )
- Glycerol (Duchefa Biochemie, catalog number: G1345.1000 )
- β-Mercaptoethanol (AMRESCO, catalog number: 0482 )
- Bromophenol Blue (BPB) (AMRESCO, catalog number: 0449 )
- Sodium orthovanadate (Sigma-Aldrich, catalog number: S6508 )
- Lysozyme (Agilent Technologies, catalog number: EC 3.2.1.17 )
- LB medium (see Recipes)
- LB agar plate (see Recipes)
- 2x YTA medium (see Recipes)
- 1x phosphate-buffered saline (PBS) (see Recipes)
- Cell lysis buffer A (see Recipes)
- Cell lysis buffer B (see Recipes)
- Elution buffer (see Recipes)
- Coomassie Blue staining buffer (see Recipes)
- 10x kinase buffer (see Recipes)
- 4x SDS loading dye (see Recipes)
- Destaining buffer (see Recipes)
Equipment
- Shaking incubator (VISION SCIENTIFIC, model: VS-8480SF )
- Erlenmeyer flask (Corning, PYREX®, catalog number: 4980-500 )
- UV spectrophotomter (Mecasys, Optizen, model: 2120UV )
- Centrifuge for 50/250 ml tubes
- 50 ml centrifuge tubes (Thermo Fisher Scientific, Thermo ScientificTM, model: 3119 )
- 250 ml centrifuge bottles (Thermo Fisher Scientific, Thermo ScientificTM, model: 3120 )
- Sonicator (Sonics & Materials, model: VCX 500 )
- High speed refrigerated micro centrifuge (TOMY DIGITAL BIOLOGY, model: MX-300 )
- Water-Jacketed CO2 incubator (Thermo Fisher Scientific, model: Forma Series II 3111 )
- Pipettes (Gilson, model: PIPETMAN®, catalog number: F167300 )
- Micro centrifuge rotor rack (TOMY DIGITAL BIOLOGY, model: AR015-24 )
- Micro centrifuge rotor rack (TOMY DIGITAL BIOLOGY, model: AR015-24 )
- Electrophoresis apparatus (Bio-Rad Laboratories, model: 1658033FC )
- Flat scanner (Epson, model: Epson Perfection V37 )
- Gel drying equipment (Vision Scientific, model: ME-2B-S )
- -80 °C freezer (NuAire, model: NU-6625D36 )
- Water bath (VISION SCIENTIFIC, model: VS-1205SW1 )
- Thermo shaker (BIOAND, model: MSC100 )
- Solid tapered microtip (Sonics & Materials, catalog number: 630-0418 )
Procedure
- Purification of GST-fusion protein
- Clone the gene of interest into the pGEX vector multiple cloning site.
- Transform the pGEX-YAP plasmid (Moon et al., 2017) into BL21 bacteria and then grow overnight on an LB plate with ampicillin (see Recipes).
Note: The purification protocol can be applied to other GST fusion proteins after cloning the gene of interest in frame with GST into the multiple cloning site of the pGEX vector. - Pick one colony and culture overnight at 37 °C in a shaking incubator (180 rpm) with 6 ml of 2x YTA medium in a 14 ml round bottom tube (add ampicillin; 300 μg/6 ml of 2x YTA medium, see Recipes).
- Transfer all cultured cells to Erlenmeyer flask.
- Incubate for 1 h at 37 °C in a shaking incubator (180 rpm) with 100 ml of 2x YTA (add ampicillin; 5 mg/100 ml of 2x YTA medium) and 50 μM ZnCl2.
- Measure optical density at 600 nm (OD600) by a UV spectrophotometer (OD should be between 0.5 and 0.8).
Note: If OD600 is less than 0.5, incubate until it exceeds 0.5. - To induce protein expression, add 1 ml of 100 mM IPTG to make a final concentration of 1 mM and incubate for 2 h at 37 °C in a shaking incubator.
Note: Prepare 100 mM IPTG stock in sterile water; store at -20 °C. - Centrifuge at 3,500 x g for 10 min at 4 °C and remove supernatant.
- Add 4 ml ice cold lysis buffer A (see Recipes) and resuspend the bacterial pellet by pipetting up and down followed by incubation for 10 min at 4 °C.
- Transfer the bacterial suspension to a 15 ml conical tube and add 1 mg of lysozyme followed by incubation for 20 min at 4 °C.
- Sonicate at output of 40% on ice for 10 sec (0.5 sec sonication/1 sec break) and keep on ice for 30 sec. Repeat sonication step one more time.
- Transfer lysate to four new 1.5 ml micro-centrifuge tubes (aliquot 1 ml) and then centrifuge at 14,000 x g for 10 min at 4 °C.
Note: Henceforth, perform all procedures in micro-centrifuge tubes.
- Transfer supernatant to four new 1.5 ml micro-centrifuge tubes.
- Add 30 μl PBS-washed 50% glutathione Sepharose 4B slurry (see Notes) to each tube and then incubate while rotating for 90 min at 4 °C.
Note: To remove the storage solution, wash 125 µl of the original glutathione Sepharose 4B slurry (it was stored in 20% ethanol) by adding with 1 ml ice cold PBS. Centrifuge at 500 x g for 5 min at 4 °C and discard supernatant. To obtain 50% glutathione Sepharose 4B slurry, resuspend the pellet in 100 µl 1x PBS. - Centrifuge at 500 x g for 5 min at 4 °C and discard supernatant.
- Add 1 ml lysis buffer A (see Recipes) and then incubate while rotating for 10 min at 4 °C.
- Centrifuge at 500 x g for 5 min at 4 °C and discard supernatant.
- Wash by adding 1 ml of 50 mM Tris (pH 8.0) and immediately centrifuge at 500 x g for 5 min at 4 °C.
- Discard supernatant.
- Add 350 μl elution buffer (see Recipes) to the glutathione Sepharose 4B beads and incubate while rotating for 4 h or overnight at 4 °C.
- Centrifuge at 500 x g for 5 min at 4 °C and divide the supernatant into 20 µl aliquots and store the protein aliquots at -80 °C for future use.
- To quantify the amount of purified recombinant GST-YAP, run an aliquot of the purified protein with a BSA standard on a 10% SDS-PAGE. For this add 5 µl 4x SDS loading dye (see Recipes) to 15 µl sample and boil for 5 min.
Note: Add 1x SDS loading dye to the other SDS-PAGE samples, e.g., molecular weight marker and BSA standard, to obtain a total volume of 20 μl. - Separate 20 µl SDS-PAGE samples on 10 % SDS-PAGE.
- Run the gel for Coomassie Blue staining, then incubate the gel for 20 min at RT in Coomassie Blue staining buffer on a rocker, followed by 5 washes with destaining buffer, 10 min each time.
Note: Quantify the final concentration and size of protein by Coomassie Blue staining with a BSA standard (see Figure 1 and Recipes). - Store the aliquots at -80 °C freezer for future use.
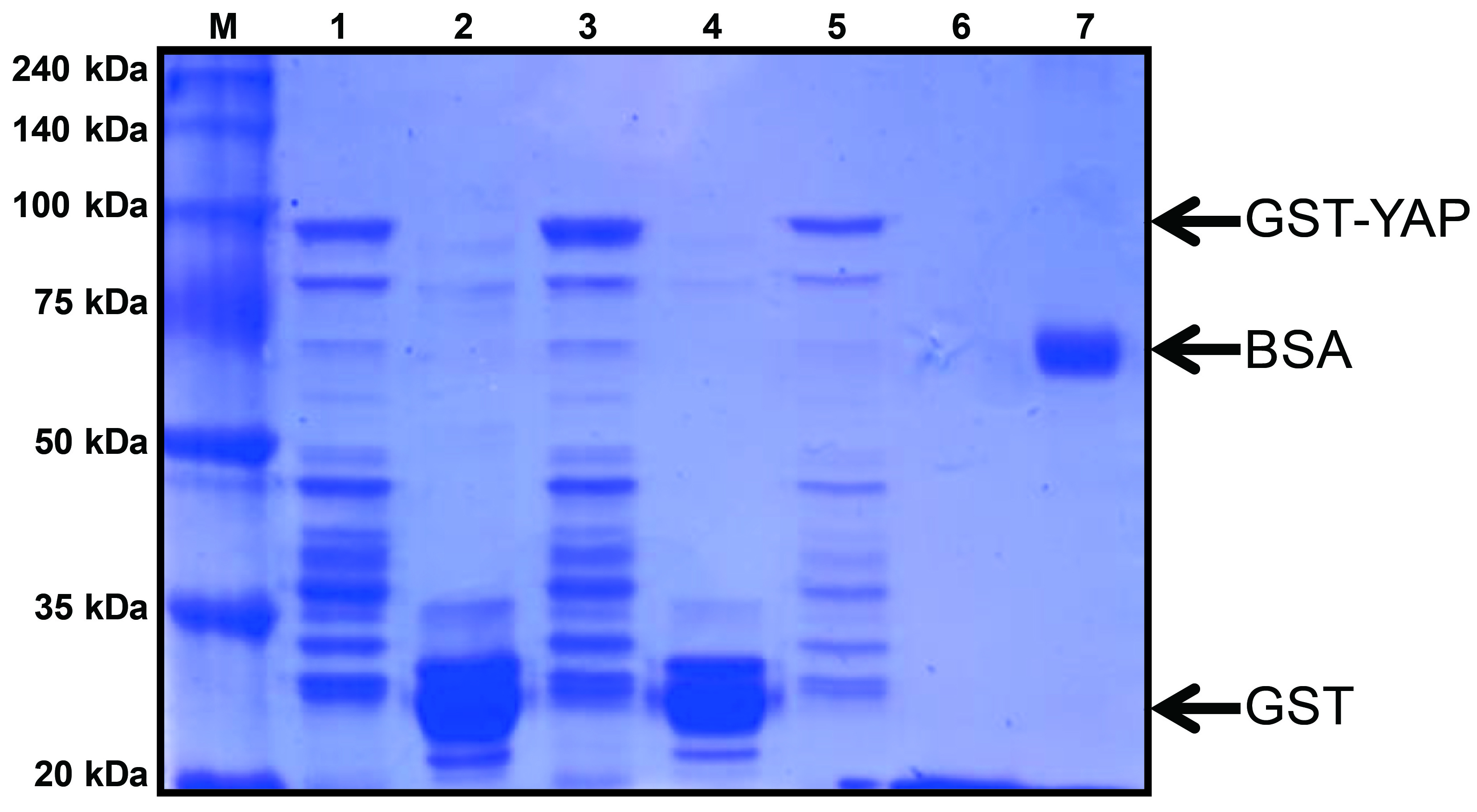
Figure 1. Coomassie Blue stained gel of purified GST-YAP. GST-YAP is evident as a 95 kDa band in the gel, GST alone runs at a molecular weight of around 25 kDa and BSA at 66 kDa. Lane M: protein size marker; Lane 1, 3, 5: GST-YAP; Lane 2, 4: GST; Lane 6: elution buffer; Lane 7: BSA (1 μg).
- Clone the gene of interest into the pGEX vector multiple cloning site.
- In vitro NLK kinase assay
- Incubate HEK293 cells at 37 °C in a humidified 5% CO2 incubator in DMEM supplemented with 10% FBS and 1x antibiotics.
- Seed 0.8 x 106 HEK293 cells on two 60 mm untreated cell culture dishes.
- After 24 h, transfect cells with 3 μg Flag-NLK-WT (wild type) or Flag-NLK-KM (kinase negative) plasmid (Ishitani et al., 1999) according to the manufacturer’s protocol (TurboFect transfection reagent).
Note: The ratio of DNA (μg) to TurboFect (μl) is 1:2 to 1:3. - Wash cells once with 1 ml ice cold 1x PBS (see Recipes), harvest cells by scraping off the cells in ice cold 1 ml 1x PBS, transfer the cells to micro-centrifuge tubes and centrifuge at 14,000 x g for 5 min at 4 °C.
- Discard supernatant and add 200 μl lysis buffer B (see Recipes) to the pellet.
- Lyse the cells by pipetting up and down around 40 times with a 200 μl pipette and incubate for 30 min at 4 °C.
- Centrifuge at 14,000 x g for 5 min at 4 °C and transfer supernatant into new micro-centrifuge tubes.
- Measure protein concentration with Protein Assay Dye Reagent kit using UV spectrophotometer.
- Incubate 400 μg of protein in 800 μl of lysis buffer B with anti-FLAG antibody (0.5 μg) on rotator overnight at 4 °C.
- Add Protein A/G plus agarose (20 μl per sample).
- Incubate on microfuge-tube rotator for 120 min at 4 °C.
- Centrifuge at 3,000 x g for 2 min at 4 °C and discard supernatant.
- Add 800 μl of lysis buffer B (see Recipes) and incubate on rotator for 5 min at 4 °C.
- Centrifuge at 3,000 x g for 2 min at 4 °C and discard supernatant (repeat steps B12-B13 three times).
- Add and gently mix 800 μl of ice cold 1x kinase buffer (see Recipes) and immediately centrifuge at 3,000 x g for 2 min at 4 °C.
- Discard supernatant and add 3 μl of 10x kinase buffer, 1 μg GST-YAP (referred to section of ‘Purification of GST-fusion protein’), 10 μCi of Adenosine 5’-triphosphate (ATP) [γ-32P] and add H2O to make the total volume up to 30 μl in two different tubes (Flag-NLK-WT/-KM).
Note: Radioactive isotopes should be used only by authorized persons and at authorized places. - Incubate for 5 min at 25 °C.
- To stop the reaction, add 10 μl of 4x SDS loading dye (see Recipes) and boil for 5 min.
- Centrifuge at 3,000 x g for 2 min at 4 °C and carefully take 30 μl of supernatant and separate proteins on 10% SDS-PAGE.
- Detach SDS-PAGE gel from electrophoresis apparatus.
- Incubate gel at a rocker with Coomassie Blue staining buffer (see Recipes) at room temperature for 30 min.
- Discard staining buffer and rinse for 20 min with destaining buffer (see Recipes).
Note: Repeat destaining until the background is nearly clear. - Capture image of SDS-PAGE gel using a flat scanner for Coomassie Blue staining.
- Dry SDS-PAGE gel for 60 min in a Gel dryer.
- Expose radioactivity with X-ray film for 30 min to 4 h in X-ray film cassette in a dark room (Figure 2).
- Develop and fix X-ray film for analyzing NLK kinase activity (see Data analysis).
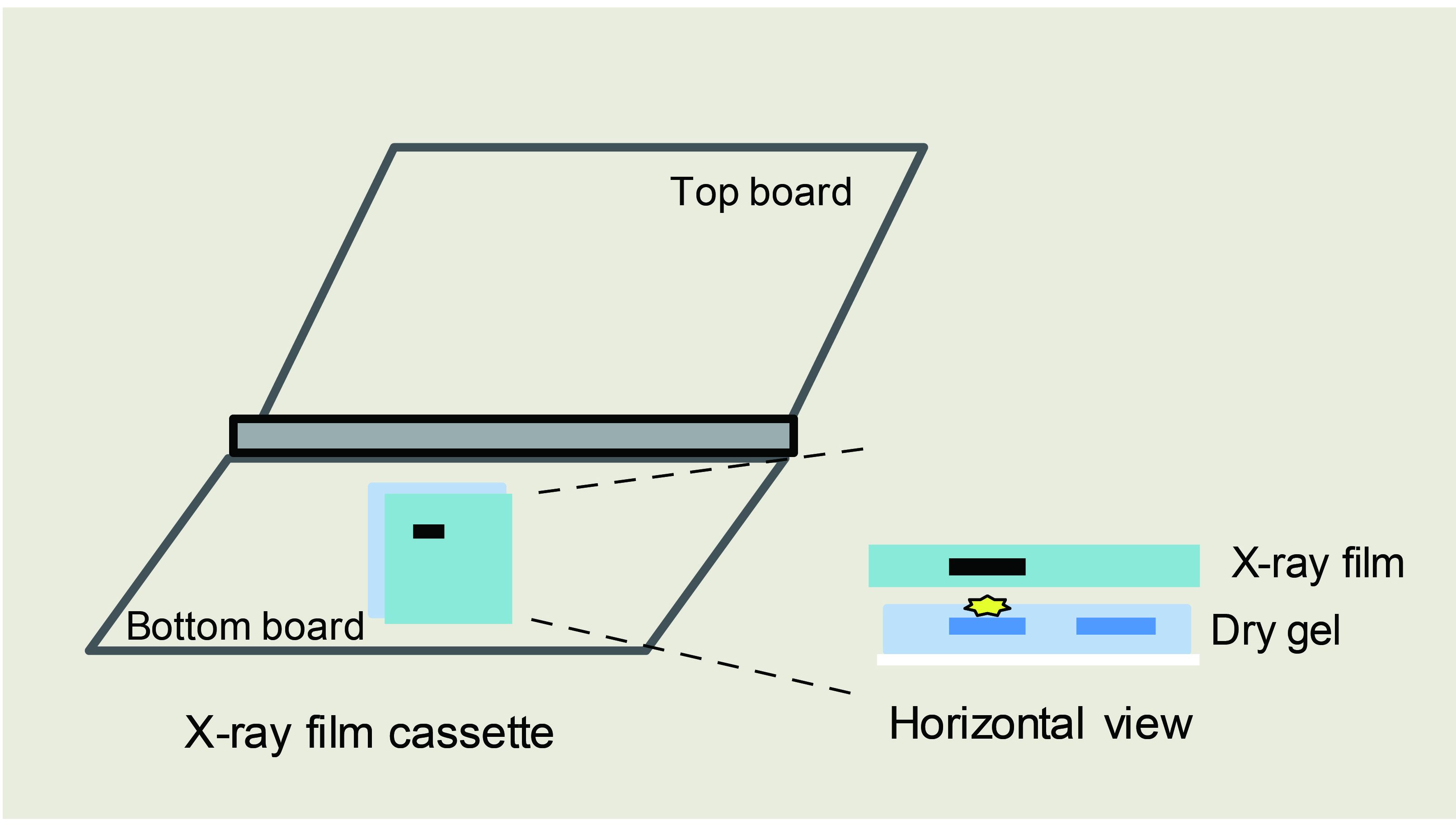
Figure 2. Schematic diagram of gel exposure
- Incubate HEK293 cells at 37 °C in a humidified 5% CO2 incubator in DMEM supplemented with 10% FBS and 1x antibiotics.
Data analysis
We have performed the in vitro kinase assay by incubating GST-YAP with immunoprecipitated Flag-NLK constructs (wild type or kinase negative) from lysates of HEK293 cells transfected with Flag-NLK. The level of autoradiography was detected by Flag-NLK-WT, but Flag-NLK-KM was not evident (Figure 3, red box). The in vitro NLK Kinase Assay suggests that NLK interacts with and directly phosphorylates YAP. Note that image analysis software (ImageJ) can be used to quantify the intensity of radiography. For this, we compared the band intensities of the phosphorylated GST-YAP (autoradiography) normalized to the total amount of GST-YAP (Coomassie Blue staining).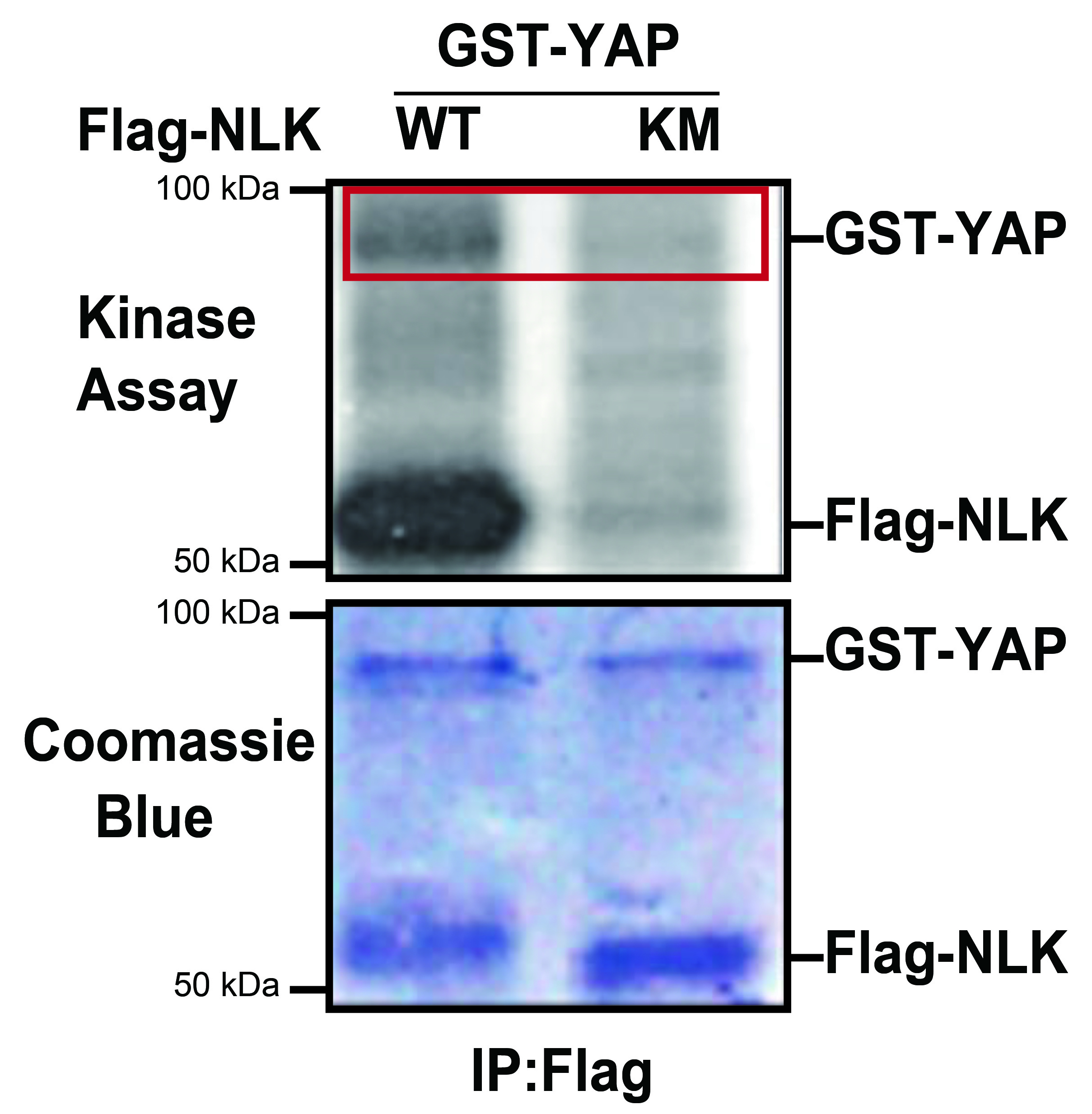
Figure 3. NLK phosphorylates YAP in vitro. HEK293 cells were transiently transfected with the indicated plasmids. Cell lysates were immunoprecipitated with anti-Flag antibodies and in vitro kinase assays were performed. The phosphorylation of GST-YAP and autophosphorylation of NLK were shown by autoradiography (top panel). Similar levels of GST-YAP and Flag-NLK were visualized by Coomassie Blue staining (bottom panel). This figure has been modified from Moon et al. (2017). EMBO Rep. doi:10.15252/embr.201642683
Recipes
- LB medium (autoclaved, store at 4 °C)
25 g/L LB broth high salt - LB agar plate (store at 4 °C)
25 g/L LB broth high salt
15 g/L Micro agar
Autoclave and pour it into plates. Cool down in sterile place - 2x YTA medium (autoclaved, store at 4 °C)
16 g/L tryptone
10 g/L yeast extract
5 g/L NaCl
Adjust pH to 7.0 - 1x phosphate-buffered saline (PBS) (autoclaved, store at 4 °C)
8 g/L NaCl
0.2 g/L KCl
1.46 g/L Na2HPO4
0.24 g/L KH2PO4
Adjust pH to 7.4 - Cell lysis buffer A (store at -20 °C)
50 mM Tris (pH 7.5)
150 mM NaCl
0.05% NP-40 (IGEPAL CA-630) - Cell lysis buffer B (prepare daily fresh, store at 4 °C)
20 mM Tris (pH 7.5)
100 mM NaCl
1 mM EDTA
2 mM EGTA
50 mM β-glycerophosphate
50 mM NaF
1 mM sodium vanadate
2 mM dithiothreitol (DTT)
1 mM phenylmethylsulfonyl fluoride (PMSF)
1 μg/ml of leupeptin
1% Triton X-100 - Elution buffer (prepare daily fresh)
50 mM Tris (pH 8.0)
6.8 mg/ml reduced glutathione - Coomassie Blue staining buffer (store at RT)
450 ml/L methanol
100 ml/L acetic acid
2.5 g/L Coomassie Blue - 10x kinase buffer (store at -20 °C)
100 mM HEPES (pH 7.5)
10 mM dithiothreitol (DTT)
50 mM MgCl2 - 4x SDS loading dye (store at -20 °C)
200 mM Tris (pH 6.8)
5% SDS
25% glycerol
200 mM β-mercaptoethanol
0.4% bromophenol blue (BPB) - Destaining buffer (store at RT)
450 ml/L methanol
100 ml/L acetic acid
Acknowledgments
This protocol was adapted from the research article by Moon et al. (2017). FLAG-NLK constructs were kindly provided by Dr. Ishitani T from Kyushu University, Japan. This work was supported by the grants from the National Research Foundation of Korea (2016R1E1A1A01943544) to E. Jho. There is no conflict of interest or competing interest that may impact the design and implementation of their protocol.
References
- Brott, B. K., Pinsky, B. A. and Erikson, R. L. (1998). Nlk is a murine protein kinase related to Erk/MAP kinases and localized in the nucleus. Proc Natl Acad Sci U S A 95(3): 963-968.
- Choi, K. W. and Benzer, S. (1994). Rotation of photoreceptor clusters in the developing Drosophila eye requires the nemo gene. Cell 78(1): 125-136.
- Ishitani, T., Hirao, T., Suzuki, M., Isoda, M., Ishitani, S., Harigaya, K., Kitagawa, M., Matsumoto, K. and Itoh, M. (2010). Nemo-like kinase suppresses Notch signalling by interfering with formation of the Notch active transcriptional complex. Nat Cell Biol 12(3): 278-285.
- Ishitani, T., Ninomiya-Tsuji, J., Nagai, S., Nishita, M., Meneghini, M., Barker, N., Waterman, M., Bowerman, B., Clevers, H., Shibuya, H. and Matsumoto, K. (1999). The TAK1-NLK-MAPK-related pathway antagonizes signalling between β-catenin and transcription factor TCF. Nature 399(6738): 798-802.
- Kim, S., Kim, Y., Lee, J. and Chung, J. (2010). Regulation of FOXO1 by TAK1-Nemo-like kinase pathway. J Biol Chem 285(11): 8122-8129.
- Moon, S., Kim, W., Kim, S., Kim, Y., Song, Y., Bilousov, O., Kim, J., Lee, T., Cha, B., Kim, M., Kim, H., Katanaev, V. L. and Jho, E. H. (2017). Phosphorylation by NLK inhibits YAP-14-3-3-interactions and induces its nuclear localization. EMBO Rep 18(1): 61-71.
- Rocheleau, C. E., Yasuda, J., Shin, T. H., Lin, R., Sawa, H., Okano, H., Priess, J. R., Davis, R. J. and Mello, C. C. (1999). WRM-1 activates the LIT-1 protein kinase to transduce anterior/posterior polarity signals in C. elegans. Cell 97(6): 717-726.
Article Information
Copyright
© 2017 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Moon, S., Kim, J. and Jho, E. (2017). In vitro NLK Kinase Assay. Bio-protocol 7(21): e2593. DOI: 10.21769/BioProtoc.2593.
Category
Cancer Biology > Proliferative signaling > Biochemical assays > Protein analysis
Biochemistry > Protein > Activity
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










