- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Isolation of Rice Embryo Single Cell Type using Laser Capture Microdissection (LCM)
Published: Vol 2, Iss 18, Sep 20, 2012 DOI: 10.21769/BioProtoc.259 Views: 12930

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

Closed Systems to Study Plant–Filamentous Fungi Associations: Emphasis on Microscopic Analyses
Vasiliki Skiada and Kalliope K. Papadopoulou
Feb 20, 2025 2878 Views

Transgene-free Genome Editing in Grapevine
Edoardo Bertini [...] Sara Zenoni
Feb 20, 2025 2550 Views
Isolation and Transfection of Protoplasts From Maize Mesophyll Cells
Lauren A. Higa [...] Zhi-Yan Du
Feb 5, 2026 181 Views
Abstract
A lot of transcriptional profiling in plant and animals has used RNAs samples from many different cell types. The laser-capture microdissection (LCM) can identify and harvest pure cellular populations directly from heterogenous tissues based on histological identification. The molecules or protein isolated from LCM-captured cells can be suitable for single cell type analysis by using chip expression profiling or sequencing.
Materials and Reagents
- Ethanol
- Acetic acid
- Histoclear (also named CitriSolv) (Thermo Fisher Scientific, catalog number: 5989-27-5 )
- DEPC H2O
- 75% (v/v) ethanol and 25% (v/v) acetic acid (see Recipes)
- Gradient series of ethanol solutions in H2O or histoclear (see Recipes)
Equipment
- Microscope
- Microtome (Waldorf, model: HM310 )
- Pix-Cell IIe LCM system (Arcturus)
- RNase-free glass slides
- 15 μm laser beam
Procedure
- Crytosectioning, fixation and dehydration of rice embryos
- Dissect rice embryo from the seeds under the microscope.
- Fix samples immediately on ice in fixation solution containing 75% (v/v) ethanol and 25% (v/v) acetic acid. The samples were left in the vials at 4 °C over night.
- Dehydrate the tissue in a series of ethanol concentrations (v/v) in order, 70%, 85%, 95%, 100%, each for 1 h at room temperature, followed by ethanol: histoclear (3:1, v/v), followed by ethanol: histoclear (1:1), ethanol: histoclear (1:3) and 100% histoclear treatment each for 1 h.
- Incubate the dehydrated samples in histoclear over night at 60 degree and then infiltrate with paraffin at 60 degree over 2 days, replacing histoclear with paraffin every 12 h.
- After embedding in paraffin, cut the embryo in 7 μm thick sections with a rotary microtome and place on RNase-free glass slides and store in darkness at 4 degree under dehydrating conditions with drierites.
- Deparaffinize sections in histoclear at room temperature for two changes of 10 min and air-dried before LCM.
- Dissect rice embryo from the seeds under the microscope.
- LCM
- Perform laser-capture microdissection using the Pix-Cell LCM system. After deparaffinizing and drying the tissues, laser-capture microdissect the interesting cells according to the manufacturer’s instructions.
- Based on the cell size diameters, the embryo organ cell types were isolated using 15 μm laser beam, laser power settings were 100 mW, and laser pulse durations were 2.5 ms. Embryo organ cell types were successfully identified and removed from heterogenous tissue by comparison of the difference among the images of the tissue before captured, the images of the tissue after removal of the harvested cells and the images of the cells captured on the cap. Generally, between 5 to 8 slides were processed each LCM caps and non-specific tissue were removed from the LCM cap using a Post-It note. See figure below.
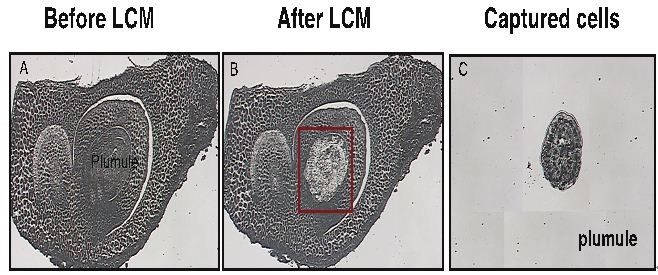
- Perform laser-capture microdissection using the Pix-Cell LCM system. After deparaffinizing and drying the tissues, laser-capture microdissect the interesting cells according to the manufacturer’s instructions.
Recipes
- 75% (v/v) ethanol and 25% (v/v) acetic acid
- Gradient series of ethanol solutions in H2O or histoclear
Acknowledgments
This protocol is adapted from Kerk et al. (2003) and Jiao et al. (2009).
References
- Jiao, Y., Tausta, S. L., Gandotra, N., Sun, N., Liu, T., Clay, N. K., Ceserani, T., Chen, M., Ma, L., Holford, M., Zhang, H. Y., Zhao, H., Deng, X. W. and Nelson, T. (2009). A transcriptome atlas of rice cell types uncovers cellular, functional and developmental hierarchies. Nat Genet 41(2): 258-263.
- Kerk, N. M., Ceserani, T., Tausta, S. L., Sussex, I. M. and Nelson, T. M. (2003). Laser capture microdissection of cells from plant tissues. Plant Physiol 132(1): 27-35.
- Nelson, T., Tausta, S. L., Gandotra, N., and Liu, T. (2006). Laser microdissection of plant tissue: what you see is what you get. Annu Rev Plant Biol 57: 181-201.
Article Information
Copyright
© 2012 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Liu, T. (2012). Isolation of Rice Embryo Single Cell Type using Laser Capture Microdissection (LCM) . Bio-protocol 2(18): e259. DOI: 10.21769/BioProtoc.259.
Category
Plant Science > Plant cell biology > Cell isolation
Plant Science > Plant cell biology > Tissue analysis
Cell Biology > Single cell analysis > Laser capture microdissection
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link








