- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Detection of Reactive Oxygen Species (ROS) in Cyanobacteria Using the Oxidant-sensing Probe 2’,7’-Dichlorodihydrofluorescein Diacetate (DCFH-DA)
Published: Vol 7, Iss 17, Sep 5, 2017 DOI: 10.21769/BioProtoc.2545 Views: 26231
Reviewed by: Dennis NürnbergPooja SaxenaAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

β-lactamase (Bla) Reporter-based System to Study Flagellar Type 3 Secretion in Salmonella
Fabienne F. V. Chevance and Kelly T. Hughes
Jun 20, 2023 1800 Views
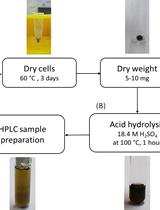
Determination of Poly(3-hydroxybutyrate) Content in Cyanobacterium Synechocystis sp. PCC 6803 Using Acid Hydrolysis Followed by High-performance Liquid Chromatography
Janine Kaewbai-ngam [...] Tanakarn Monshupanee
Aug 20, 2023 1865 Views
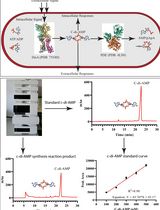
An HPLC-based Assay to Study the Activity of Cyclic Diadenosine Monophosphate (C-di-AMP) Synthase DisA from Mycobacterium smegmatis
Avisek Mahapa [...] Dipankar Chatterji
Dec 20, 2024 1815 Views
Abstract
Reactive oxygen species (ROS) are cell signaling molecules synthesized inside the cells as a response to routine metabolic processes. In stress conditions such as ultraviolet radiation (UVR), ROS concentration increases several folds in the cells that become toxic for the cell survival. Here we present the method for in vivo detection of ROS by using an oxidant-sensing probe 2’,7’-dichlorodihydrofluorescein diacetate (DCFH-DA) in cyanobacteria. This method provides reliable, simple, rapid and cost effective means for detection of ROS in cyanobacteria.
Keywords: Reactive oxygen speciesBackground
Cyanobacteria are the most ancient oxygenic photoautotrophs; they play an important role in the biomass production in both aquatic and terrestrial ecosystems and serve as source of various value-added products (Vaishampayan et al., 2001; Häder et al., 2007; Fischer, 2008). In recent years the depletion of the ozone layer has resulted in an increase in solar ultraviolet radiation (UVR) influx, which is harmful to all organisms residing on Earth including cyanobacteria (Holzinger and Lutz, 2006). The UVR harms cyanobacteria directly by acting on DNA/proteins or indirectly through oxidative damage from reactive oxygen species (ROS) (He and Häder, 2002). In plants, algal and mammalian cells various fluorescence and chemiluminescence methods have been used for detecting ROS (Crow, 1997; He and Häder, 2002; Soh, 2006; Wu et al., 2007; Palomero et al., 2008).
2’,7’-Dichlorodihydrofluorescein diacetate (DCFH-DA) is a non-fluorescent, cell-permeable dye which is hydrolyzed intracellularly into its polar, but non-fluorescent form DCFH on the action of cellular esterases and thus is retained in the cell. Oxidation of DCFH by the action of intracellular ROS and other peroxides turns the molecule into its highly fluorescent form 2’,7’-dichlorofluorescein (DCF) that can be detected by various fluorescent methods (He and Häder, 2002; Rastogi et al., 2010; Singh et al., 2014) (Figure 1). Although DCFH-DA is widely used for the detection of ROS, it should be noted, however, that the dye cannot be used as an indicator for a specific form of ROS (Marchesi et al., 1999).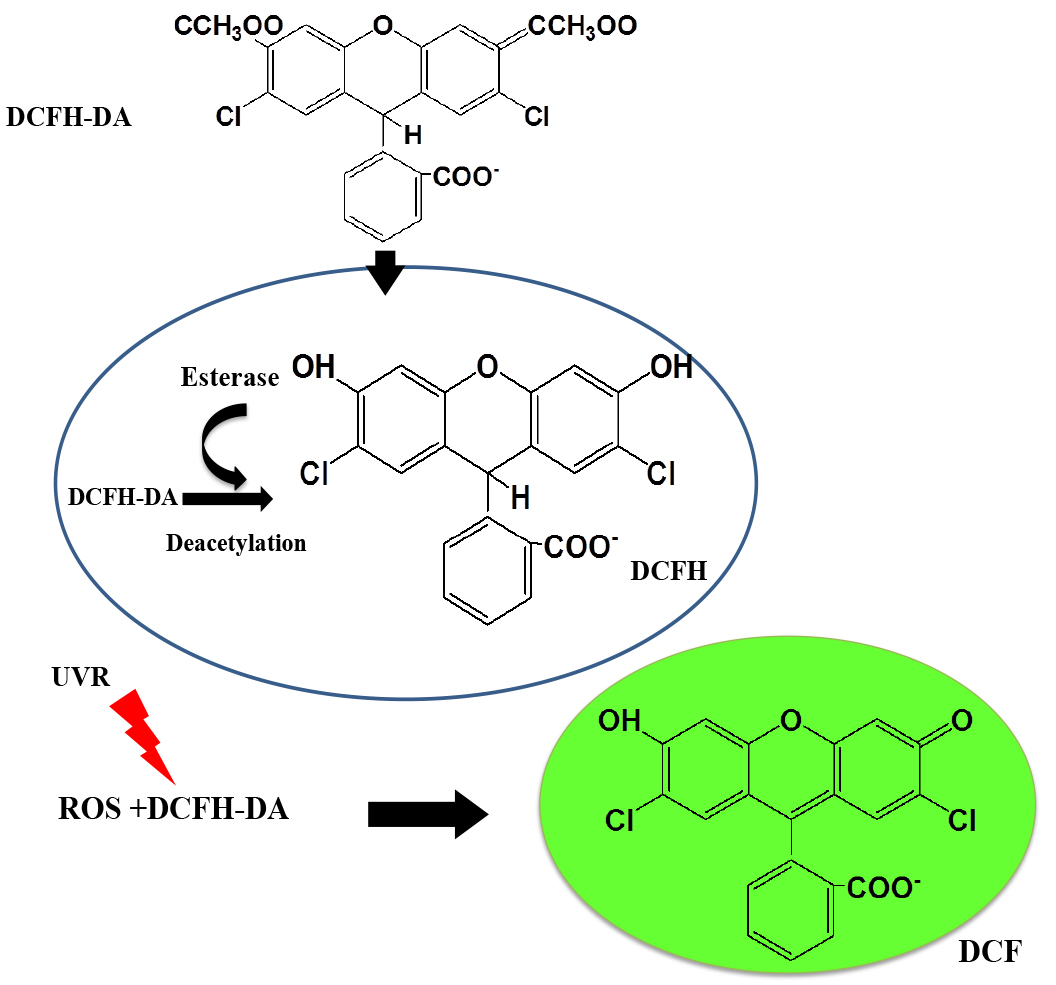
Figure 1. Mechanism of action of DCFH-DA probe inside the cell (Adapted from He and Häder, 2002)
Materials and Reagents
- 2 ml RNase, DNase free microcentrifuge tube (Thermo Fisher Scientific, InvitrogenTM, catalog number: AM12425 )
- Glass microscope slides (Fisher Scientific, catalog number: 12-544-4 )
- Glass microscope coverslips (Fisher Scientific, catalog number: S17525B )
- Millipore membrane filter (EMD Millipore, catalog number: HAWP04700 )
- Cuvette (fluorescence spectroscopy) 3 ml (Hellma, catalog number: 101-QS )
- Cyanobacterial cells e.g., Nostoc sp. strain HKAR-2
Note: Nostoc sp. strain HKAR-2, an autotrophic, filamentous and heterocystous cyanobacterium, was grown under axenic conditions in nitrogen-free liquid BGA medium (Safferman and Morris, 1964) at 20 ± 2 °C under continuous white light (12 ± 2 Wm-2) to an OD750 of 0.8 to 0.9 (exponential growth phase) which was measured using quartz cuvette in a spectrophotometer. - Nail varnish (Lakme)
- Potassium phosphate dibasic anhydrous (K2HPO4)
- Potassium phosphate monobasic (KH2PO4)
- 2’,7’-Dichlorodihydrofluorescein diacetate (DCFH-DA) (Sigma-Aldrich, catalog number: D6883 )
- 100% ethanol (Sigma-Aldrich, catalog number: 459836 )
- 50 mM phosphate buffer (see Recipes)
- 2 mM 2’,7’-dichlorodihydrofluorescein diacetate (DCFH-DA) stock solution (see Recipes)
Equipment
- Glass Petri-dishes (Corning, catalog number: 3160-102 )
- Measuring cylinder
- 295 nm UV cut-off filter (Ultraphan, Digefra, Munich, Germany) to facilitate the desired wavebands of UV-B (280-315 nm), UV-A (315-400 nm) and PAR (400-700 nm)
- UV-treatment chamber fitted with UV-B (Philips Ultraviolet-B TL 40 W: 12, Philips Lighting, model: TL 40W/12 RS SLV/25 ), UV-A (Philips Ultraviolet-A TL 40 W: 12, Philips Lighting, model: TL-K 40W/10-R UV-A ) and PAR (55.08 ± 9.18 μmol m-2 sec-1) (OSRAM L 36 W: 32 Lumilux de luxe warm white and Radium NL 36 W: 26 Universal white, Germany) lamps
- Glass rod (Fisher Scientific, catalog number: 11-380A )
- Magnetic stirrer (REMI ELECTROTECHNIK, model: 2 MLH )
- Refrigerated centrifuge (REMI ELECTROTECHNIK, model: CM-12 PLUS )
- Shaker
- Fluorescence microscope (Nikon eclipse Ni fluorescence microscope processed by NIS Elements (BR))
- Fluorescence spectrophotometer (Agilent Technologies, model: Cary Eclipse )
- Spectrophotometer (Hitachi High-Technologies, model: U-2900 , Double beam spectrophotometer)
- Quartz cuvette (3.5 ml) (Cole-Parmer, JENWAY, catalog number: 035 028 )
Software
- NIS-Elements (BR) imaging software (Nikon)
- SigmaPlot 11 software
Procedure
A flow chart of the sample preparation is shown in Figure 2.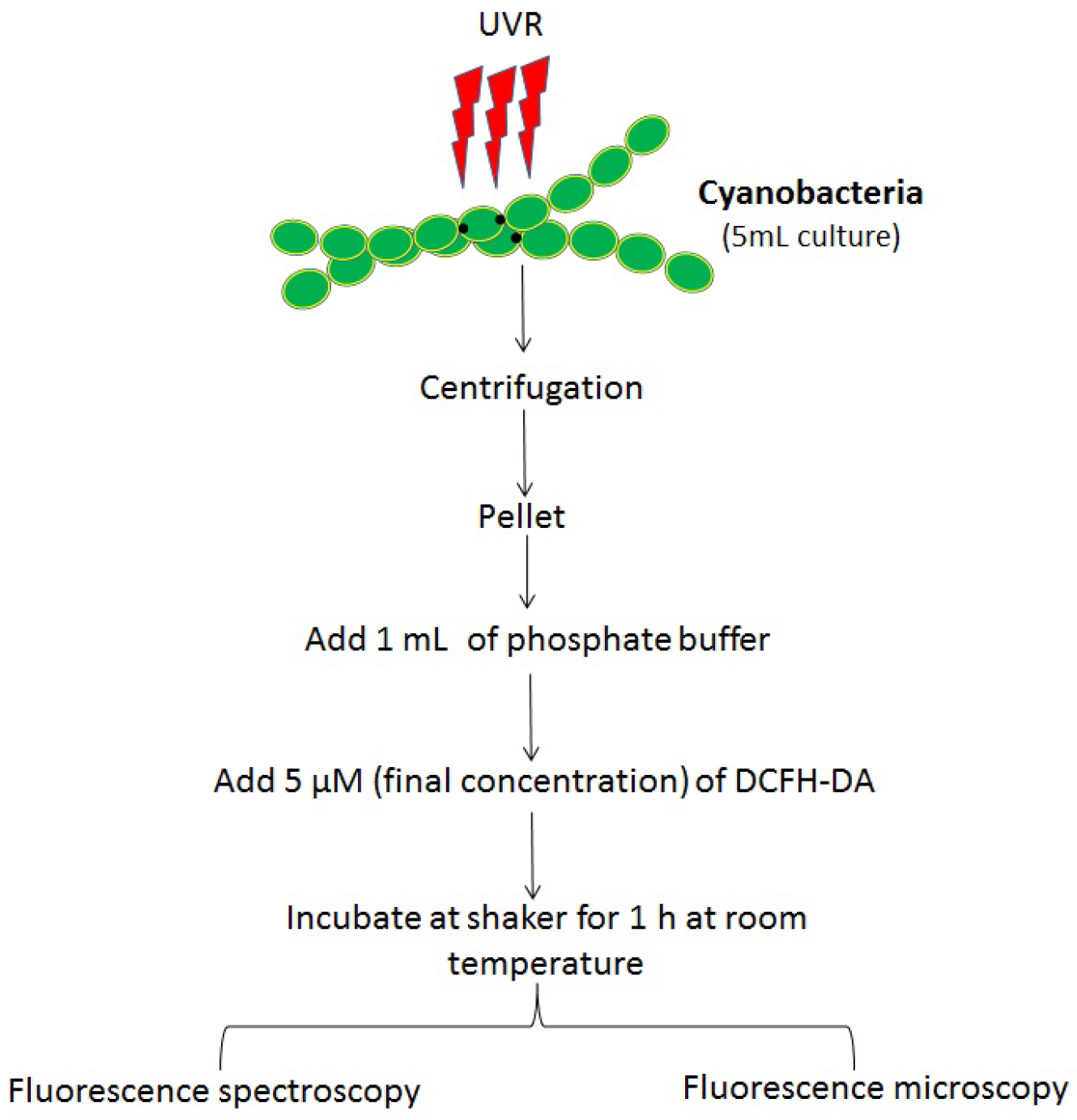
Figure 2. Flow chart showing various steps involved in the protocol
- Transfer 100 ml of cyanobacterial culture to sterile glass Petri-dishes with the help of a measuring cylinder. Cover the Petri dishes with a 295 nm UV cut-off filter and transfer them to a UV light treatment chamber.
- Irradiate the cyanobacteria with UV-A, UV-B and PAR and maintain the temperature of the chamber at 25 ± 2 °C to avoid a heating effect. Mix the culture with a glass rod at regular intervals or use magnetic stirrer (15-20 rpm) to avoid self-shading of the cells.
- After desired time intervals (hereafter 12 h and 24 h), take 5 ml of sample and harvest cells by centrifugation at 9,050 x g for 20 min at room temperature.
- Resuspend the cell pellet in 1 ml phosphate buffer (see Recipes).
- Add 2.5 µl of 2 mM 2’,7’-dichlorodihydrofluorescein diacetate (DCFH-DA) solubilized in ethanol (see Recipes) to the sample mixture.
- Incubate the sample mixture on a shaker (15 rpm) at room temperature in the dark for 1 h.
- After 1 h incubation
- For fluorescence microscopy: Take a clean glass slide and add 30 µl of culture. Cover the cells with a glass cover slip. Seal the slide with nail varnish to avoid drying out. Cells were visualized under a fluorescence microscope using an excitation wavelength of 488 nm and emission was detected in the range of 500-600 nm (Figure 3).
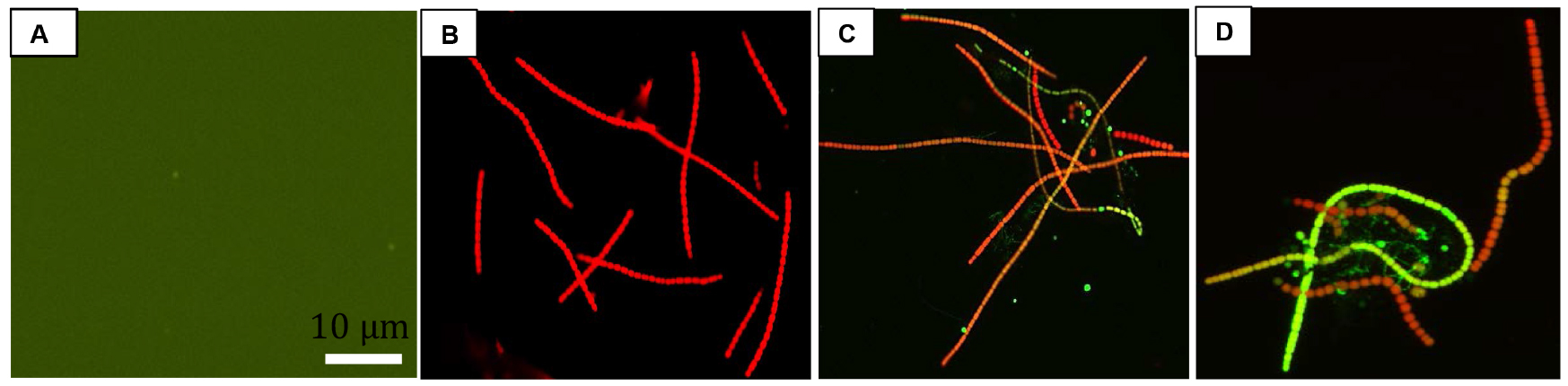
Figure 3. Fluorescence images of UV-A + UV-B + PAR exposed Nostoc sp. strain HKAR-2 showing green DCF fluorescence after reaction with ROS. Negative control (containing DCFH-DA only; showing the basal level of fluorescence) (A); fluorescence at 0 h (B); 12 h UV-A + UV-B + PAR exposure (C); 24 h UV-A + UV-B + PAR exposure (D). PAR: Photosynthetically active radiation. Scale bar =10 µm. - For fluorescence spectroscopy: 3 ml of liquid sample was added to a cuvette for fluorescence spectrophotometric analysis (Figure 4). The cuvette was placed in fluorescence spectrophotometer and the sample was excited at 485 nm. Emission was recorded in the range of 500-600 nm. The exposure time was limited to 600 msec to reduce the damage of cells. Fluorescence was measured in terms of emitted fluorescence intensity after different durations of stress exposure.
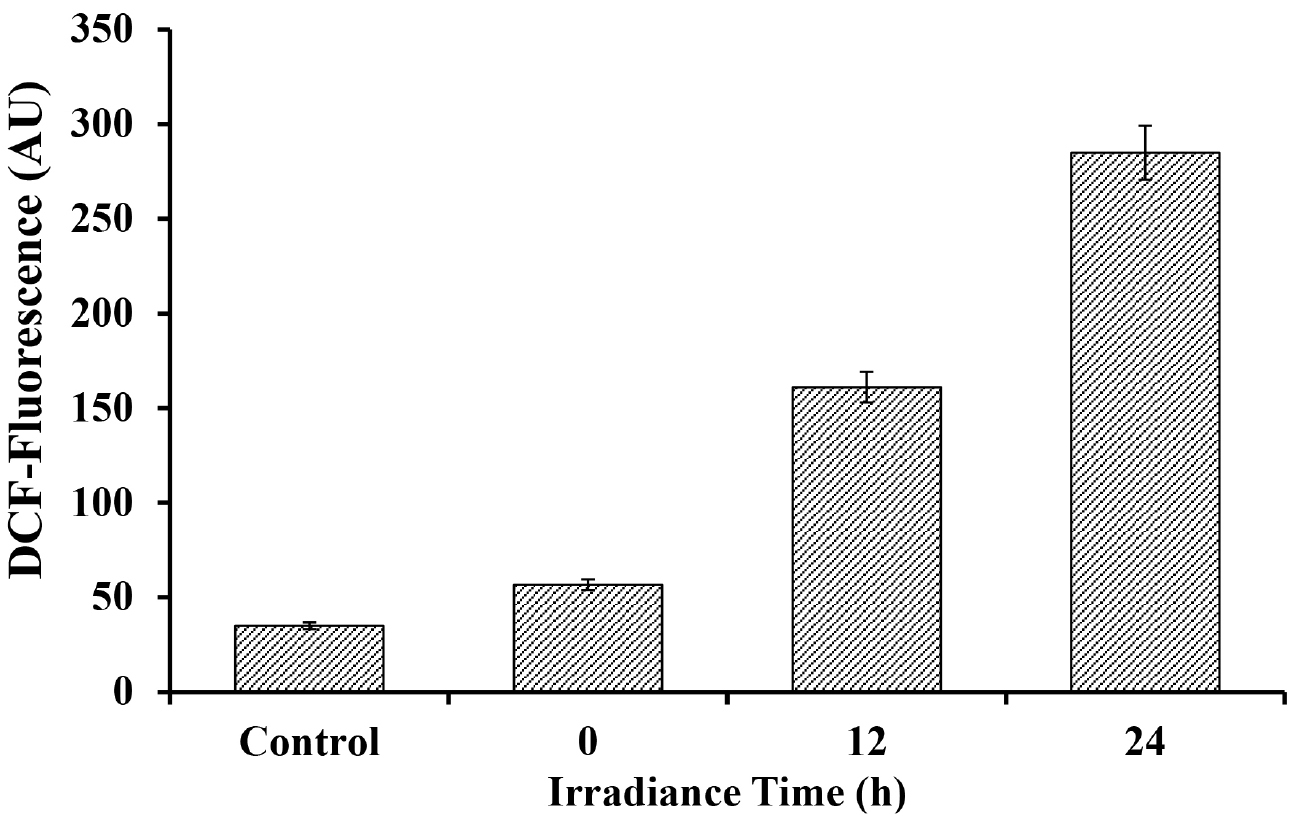
Figure 4. Fluorescence intensity of DCF after reaction with ROS generated due to varying duration of UV-A + UV-B + PAR exposure in Nostoc sp. strain HKAR-2 (means ± SD, n = 3). Control: Untreated sample. PAR: Photosynthetically active radiation.
Data analysis
The UVR irradiated cells were analyzed using a Nikon Eclipse Ni fluorescence microscope processed by NIS-Elements (BR) imaging software. The microscope was equipped with the following filter set: UV: (DAPI) EX 340 nm EM 488 nm, blue: (FITC) EX 495 nm EM 510 nm and green: (PI 550) EX 550 nm EM 650 nm. Cells were imaged in the epifluorescence mode with a 20x objective lens. The image analysis was performed by NIS-Elements (BR) imaging software provided by Nikon and images were saved in JPEG format. In addition, the fluorescence of the samples was measured by a fluorescence spectrophotometer (Cary Eclipse, Agilent Technologies) with an excitation wavelength of 485 nm and an emission band between 500 and 600 nm. The data of the fluorescence spectra were exported to excel and the fluorescence intensity values at 525 nm were extracted. A bar diagram was plotted with SigmaPlot 11 software. All fluorescence measurements were performed at room temperature. All results are presented as mean values of three replicates for fluorescence spectrophotometer analysis and random sites of filaments were used for fluorescence microscopy. All data were analysed by one-way analysis of variance (Brown, 2005). Once a significant difference was detected post hoc multiple comparisons were made by using the Tukey test. The level of significance was set at 0.05 for all tests. All statistical analyses were performed by using SigmaPlot 11 software.
Notes
- Direct exposure of light to UV treated samples should be avoided.
- The stock solution of 2’,7’-dichlorodihydrofluorescein diacetate (DCFH-DA) once prepared, was kept in -20 °C for further use and direct exposure to light should be avoided (it remains stable for more than 3 months).
- All solutions were filtered through 0.25 µm size Millipore membrane filter before use.
Recipes
- 50 mM phosphate buffer (400 ml)
8.7 g K2HPO4
6.8 g KH2PO4
Adjust the pH to 7.00
Filter through 0.25 µm size Millipore membrane filter before use - 2 mM (w/v) 2’,7’-dichlorodihydrofluorescein diacetate (DCFH-DA) stock solution
Dissolve 0.974 mg of 2’,7’-dichlorodihydrofluorescein diacetate (DCFH-DA) in 1 ml of absolute ethanol
Acknowledgments
Rajneesh and Jainendra Pathak are thankful to the Department of Biotechnology (DBT-JRF/13/AL/143/2158) and the Council of Scientific and Industrial Research (09/013/0515/2013-EMR-I), New Delhi, India, respectively, for the financial support in the form of fellowships. SP Singh acknowledges the DST-SERB and UGC for Early Career Research Award and UGC Start-Up Research Grant, respectively. We are also thankful to the Interdisciplinary School of Life Sciences (ISLS), BHU, Varanasi, India, for providing access to the fluorescence microscopy facility. This protocol was adapted from procedures published by Rastogi et al., 2010.
References
- Brown,A. M. (2005). A new software for carryingout one-way ANOVA post hoc tests. ComputMethods Programs Biomed 79(1): 89-95.
- Crow,J. P. (1997). Dichlorodihydrofluorescein anddihydrorhodamine 123 are sensitive indicators of peroxynitrite in vitro: implications for intracellularmeasurement of reactive nitrogen and oxygen species. NitricOxide 1(2): 145-157.
- Fischer,W. F. (2008). Life beforethe rise of oxygen. Nature 455: 1051-1052.
- Häder, D. P., Kumar, H. D., Smith,R. C. and Worrest, R. C. (2007). Effects of solar UV radiationon aquatic ecosystems and interactions with climate change. PhotochemPhotobiol Sci 6(3): 267-285.
- He, Y. Y. and Häder, D. P. (2002). UV-B-induced formation ofreactive oxygen species and oxidative damage of the cyanobacterium Anabaena sp.: protective effects of ascorbic acid and N-acetyl-L-cysteine. J Photochem Photobiol B 66(2): 115-124.
- Holzinger,A. and Lutz, C. (2006). Algae and UV irradiation:effects on ultrastructure and related metabolic functions. Micron 37(3): 190-207.
- Marchesi,E., Rota, C., Fann, Y. C., Chignell, C. F. and Mason, R. P. (1999). Photoreductionof the fluorescent dye 2’-7’-dichlorofluorescein: a spin trapping and directelectron spin resonance study with implications for oxidative stressmeasurements. FreeRadicBiol Med 26(1-2): 148-161.
- Palomero,J., Pye, D., Kabayo, T., Spiller, D. G. and Jackson, M. J. (2008). In situ detection and measurement ofintracellular reactive oxygen species in single isolated mature skeletal musclefibers by real time fluorescence microscopy. AntioxidRedox Signal 10(8): 1463-1474.
- Rastogi,R. P., Singh, S. P., Häder, D. P. and Sinha, R. P.(2010). Detection of reactive oxygenspecies (ROS) by the oxidant-sensing probe 2’,7’-dichlorodihydrofluoresceindiacetate in the cyanobacterium Anabaena variabilis PCC 7937. BiochemBiophys Res Commun 397(3): 603-607.
- Safferman, R. S. and Morris, M E. (1964). Growth characteristics of the blue-green algal virusLPP1. J Bacteriol 88: 771-775.
- Singh,S. P., Rastogi, R. P., Hader, D. P. and Sinha, R. P. (2014). Temporaldynamics of ROS biogenesis under simulated solar radiation in thecyanobacterium Anabaena variabilis PCC 7937. Protoplasma 251(5): 1223-1230.
- Soh,N. (2006). Recent advances in fluorescentprobes for the detection of reactive oxygen species. AnalBioanal Chem 386(3): 532-543.
- Vaishampayan,A., Sinha, R. P., Häder, D. P., Dey, T., Gupta, A. K., Bhan, U. and Rao, A. L. (2001). Cyanobacterialbiofertilizers in rice agriculture. Bot Rev 67:453-516.
- Wu, T. T. Chen, K. and Keaney, J. F. (2007). Use of 2’,7’-dichlorodihydrofluorescein diacetate in an easy and quantifiable assay forhypochlorous acid oxidation. FASEBJ 21: 882-889.
Article Information
Copyright
© 2017 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Rajneesh, , Pathak, J., Chatterjee, A., Singh, S. P. and Sinha, R. P. (2017). Detection of Reactive Oxygen Species (ROS) in Cyanobacteria Using the Oxidant-sensing Probe 2’,7’-Dichlorodihydrofluorescein Diacetate (DCFH-DA). Bio-protocol 7(17): e2545. DOI: 10.21769/BioProtoc.2545.
Category
Microbiology > Microbial biochemistry > Other compound
Biochemistry > Other compound > Reactive oxygen species
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link











