- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Isolation and Analysis of Stromal Vascular Cells from Visceral Adipose Tissue
Published: Vol 7, Iss 16, Aug 20, 2017 DOI: 10.21769/BioProtoc.2444 Views: 13942
Reviewed by: Jia LiAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
Selective Enrichment and Identification of Cerebrospinal Fluid-Contacting Neurons In Vitro via PKD2L1 Promoter-Driven Lentiviral System
Wei Tan [...] Qing Li
Nov 20, 2025 1378 Views
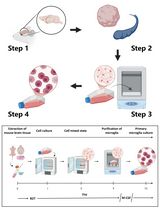
Revisiting Primary Microglia Isolation Protocol: An Improved Method for Microglia Extraction
Jianwei Li [...] Guohui Lu
Dec 5, 2025 1582 Views
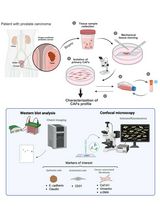
Non-Enzymatic Isolation of Cancer-Associated Fibroblasts From Human Prostate Tumor Explants
Giulia Gangarossa [...] Paola Chiarugi
Mar 5, 2026 155 Views
Abstract
The obesity epidemic is the underlying driver of the type 2 diabetes mellitus epidemic. A remarkable accumulation of various pro-inflammatory immune cells in adipose tissues is a hallmark of obesity and leads to pathogenesis of tissue inflammation and insulin resistance. Here, we describe a detailed protocol to isolate adipose tissue stromal vascular cells (SVCs), which enrich various immune cells of adipose tissues. These SVCs can be used to examine the population and activation status of immune cells by tracking their cell surface antigens, gene expression, and activation of specific signaling pathways.
Keywords: Adipose tissueBackground
Over the past several decades, obesity is now an epidemic and has become one of the most common causes of insulin resistance. Insulin resistance is the key etiology for the pathogenesis of metabolic syndrome. Prolonged status of metabolic syndrome drives the development of type 2 diabetes mellitus (T2DM) (Romeo et al., 2012; Johnson and Olefsky, 2013; Saltiel and Olefsky, 2017).
Chronic low-degree tissue inflammation, accompanied by enhanced immune cell infiltration, is a hallmark of obesity in both rodent and human and is a major causal factor for the pathogenesis of insulin resistance through promoting the inflammation status and interrupting the insulin signalling (Romeo et al., 2012; Johnson and Olefsky, 2013; Saltiel and Olefsky, 2017). The infiltrated immune cells such as pro-inflammatory macrophages and B cells play critical roles in modulating obesity-associated adipose tissue inflammation and insulin resistance (Weisberg et al., 2003; Winer et al., 2011). Chronic nutrient excess drives adipose tissue macrophages (ATMs) to undergo a unique phenotypic switch from anti-inflammatory M2-like activation in lean adipose tissue to a more pro-inflammatory M1-like activation state in obese tissues (Lumeng et al., 2007; Nguyen et al., 2007; Lumeng et al., 2008). Pro-inflammatory M1-like ATMs contribute to the development of tissue inflammation and systemic insulin resistance in obesity. Our recent study also demonstrates that leukotriene B4 (LTB4)-induced recruitment and activation of adipose tissue B2 (ATB2) cells can cause obesity-induced insulin resistance (Ying et al., 2017). In this protocol, we provide a step-by-step procedure to isolate stromal vascular cells from adipose tissue and characterize various immune cells in adipose tissues.
Materials and Reagents
- Pipette tips (USA Scientific)
- 100-mm Petri dish
- 50 ml Falcon tube (Corning, Falcon®, catalog number: 352070 )
- Nylon biopsy bag (Electron Microscopy Sciences, catalog number: 62324-35 )
- MicroAmp Optical 96-well reaction plate (Thermo Fisher Scientific, Applied BiosystemsTM, catalog number: N8010560 )
- Stromal vascular cells (SVCs)
- 70% ethanol
- BDTM stabilizing fixative buffer (BD, BD Biosciences, catalog number: 339860 )
- Phosphate-buffered saline (PBS)
- 2% fetal bovine serum (FBS)
- Antibody
- Rabbit monoclonal anti-GAPDH (Cell Signaling Technology, catalog number: 5174 )
- Rabbit monoclonal anti-Phospho-NF-κB p65 (Cell Signaling Technology, catalog number: 3033 )
- PE-Cyanine7 anti-mouse F4/80 (Thermo Fisher Scientific, eBioscienceTM, catalog number: 25-4801-82 )
- Alexa Fluor 488 anti-mouse CD11b (Thermo Fisher Scientific, eBioscience TM, catalog number: 53-0112-82 )
- APC anti-mouse CD11c (Thermo Fisher Scientific, eBioscienceTM , catalog number: 17-0114-82 )
- PE anti-mouse CD206 (BioLegend, catalog number: 141706 )
- eVolve-605 anti-mouse CD45 (Thermo Fisher Scientific, eBioscienceTM, catalog number: 83-0451-42 )
- APC anti-mouse CD19 (Thermo Fisher Scientific, eBioscienceTM , catalog number: 17-0193-82 )
- Rabbit monoclonal anti-GAPDH (Cell Signaling Technology, catalog number: 5174 )
- Trizol reagent (Thermo Fisher Scientific, InvitrogenTM , catalog number: 15596026 )
- Direct-zol RNA kits (Zymo Research, catalog number: R2070 )
- High-Capacity cDNA Reverse Transcription Kit (Thermo Fisher Scientific, Applied BiosystemsTM, catalog number: 4368813 )
- qPCR primers (Table 1)
Table 1. qPCR primer information
- SYBR Green PCR Master mix (Thermo Fisher Scientific, Applied BiosystemsTM, catalog number: 4309155 )
- Hanks’ balanced salt solution (HEPES) (Thermo Fisher Scientific, GibcoTM, catalog number: 15630080 )
- Collagenase II (Sigma-Aldrich, catalog number: C1764-50MG )
- Bovine serum albumin (BSA)
- Ammonium chloride (NH4Cl)
- Potassium bicarbonate (KHCO3)
- 5% EDTA
- Sodium azide (NaN3)
- Iscove’s Modified Dulbecco’s Medium (IMDM)
- Penicillin-streptomycin (Thermo Fisher Scientific, GibcoTM, catalog number. 15140122 )
- Digestion buffer (see Recipes)
- Red blood cell lysis buffer (see Recipes)
- FACS staining buffer (see Recipes)
- Complete culture medium (see Recipes)
Equipment
- Pipettes
- Mortar and pestle
- Curved scissors
- New Brunswick Scientific 12400 incubator shaker (Eppendorf, model: New BrunswickTM 124 )
- Eppendorf centrifuge 5810R (Eppendorf, model: 5810 R )
- TC20 automated cell counter (Bio-Rad Laboratories, model: TC20TM, catalog number: 1450102 )
- BD FACSCanto flow cytometry analyzer
- StepOnePlus Real-Time PCR System (Thermo Fisher Scientific, Applied BiosystemsTM, model: StepOnePlusTM , catalog number: 4376600)
- DNA Engine Peltier Thermal Cycler (Bio-Rad Laboratories, model: PTC-200 )
- NanoDrop 1000 Spectrophotometer (Thermo Fisher Scientific, model: NanoDropTM 1000 )
Software
- FlowJo
- GraphPad Prism
Procedure
- Isolation of stromal vascular cells (SVCs) from adipose tissue (Figure 1)
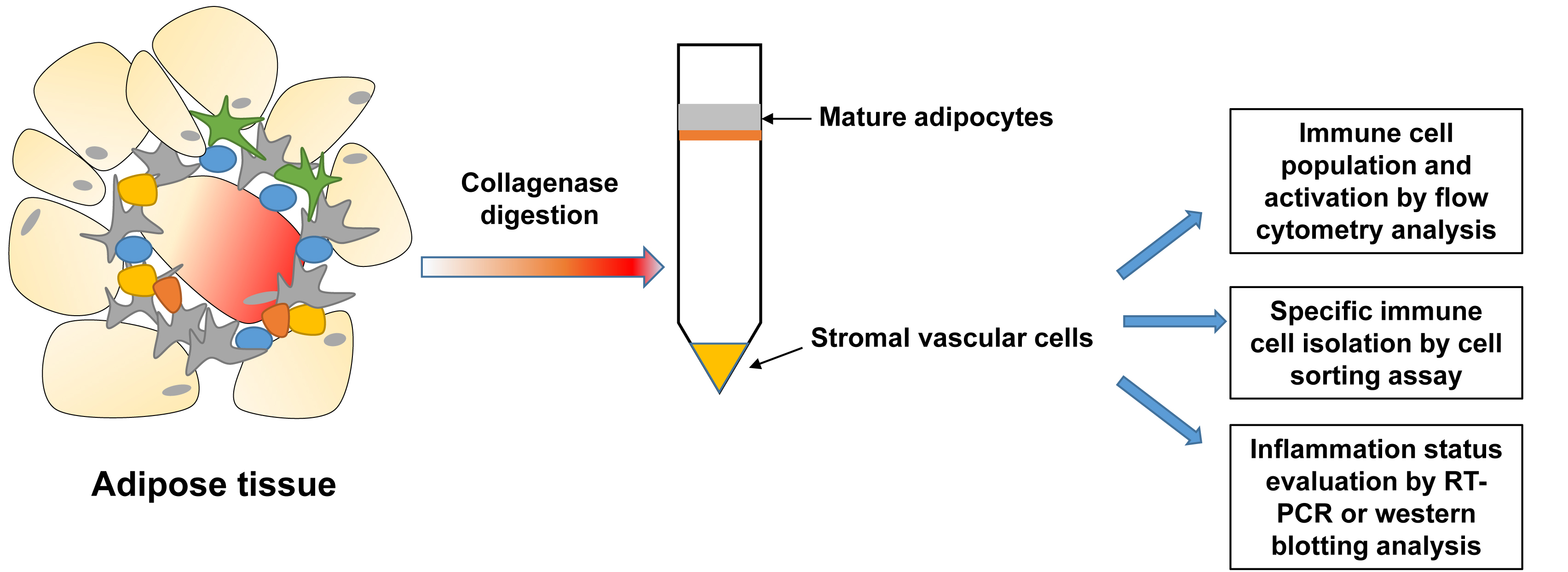
Figure 1. Scheme for isolation of stromal vascular cells from adipose tissue- Male mice with 18-20 weeks age were euthanized following the ICCUAC approval protocol.
- After thoroughly wet the fur with 70% ethanol and then open the thoracic cavity, epididymal fat tissues were collected and weighed.
- Mince fat tissues on a 100-mm Petri dish by using a curved scissor.
- Transfer minced tissues to a 50 ml Falcon tube containing digestion buffer (HBSS containing 2% BSA and 100 mM HEPES).
- Add 10 ml digestion buffer (see Recipes) per 1 g fat tissue.
- Incubate in an incubator shaker (secure the tubes horizontally) at 220 rpm for 10 min, 37 °C (no chunk tissue represents complete digestion).
- Add equal volume of complete culture medium and mix to cease the digestion.
- Filter through a nylon biopsy bag (pore size = 250 µm) into a new 50 ml Falcon tube.
- Centrifuge at 1,500 x g at 4 °C for 10 min.
- Decant the supernatant and re-suspend cell pellet in 1 ml of PBS/2% FBS.
- Add 2 ml of red blood cell lysis buffer (see Recipes) (1:2) and mix.
- Incubate on ice for 10 min.
- Add 3 ml of complete medium (1:1) and mix.
- Centrifuge at 1,000 x g for 5 min, 4 °C.
- Decant the supernatant.
- SVCs are resuspended with 0.5 ml of PBS/2% FBS and then counted using the TC20 automated cell counter. Cell concentration is adjusted to 2 x 106 cells per ml, and the cells are ready for the downstream experiments.
- Male mice with 18-20 weeks age were euthanized following the ICCUAC approval protocol.
- Flow cytometry analysis
- Prepare 0.5-1 x 106 SVCs and resuspend in FACS staining buffer (see Recipes).
- Stain SVCs with the fluorescent-conjugated antibodies (Materials and Reagents #11) following the manufacturer’s instruction.
- Incubate for 15 min at RT (light sensitive; keep in the dark).
- Add 1 ml FACS staining buffer.
- Centrifuge at 1,000 x g for 5 min, 4 °C.
- Decant the supernatant.
- Re-suspend with 100-200 µl FACS staining buffer.
Option: Cells can be stored in BDTM stabilizing fixative buffer up to 3 days at 4 °C and then used for FACS analysis. - Then the sample is ready for flow cytometry analysis (Figure 2) or specific immune cell isolation by cell sorter.
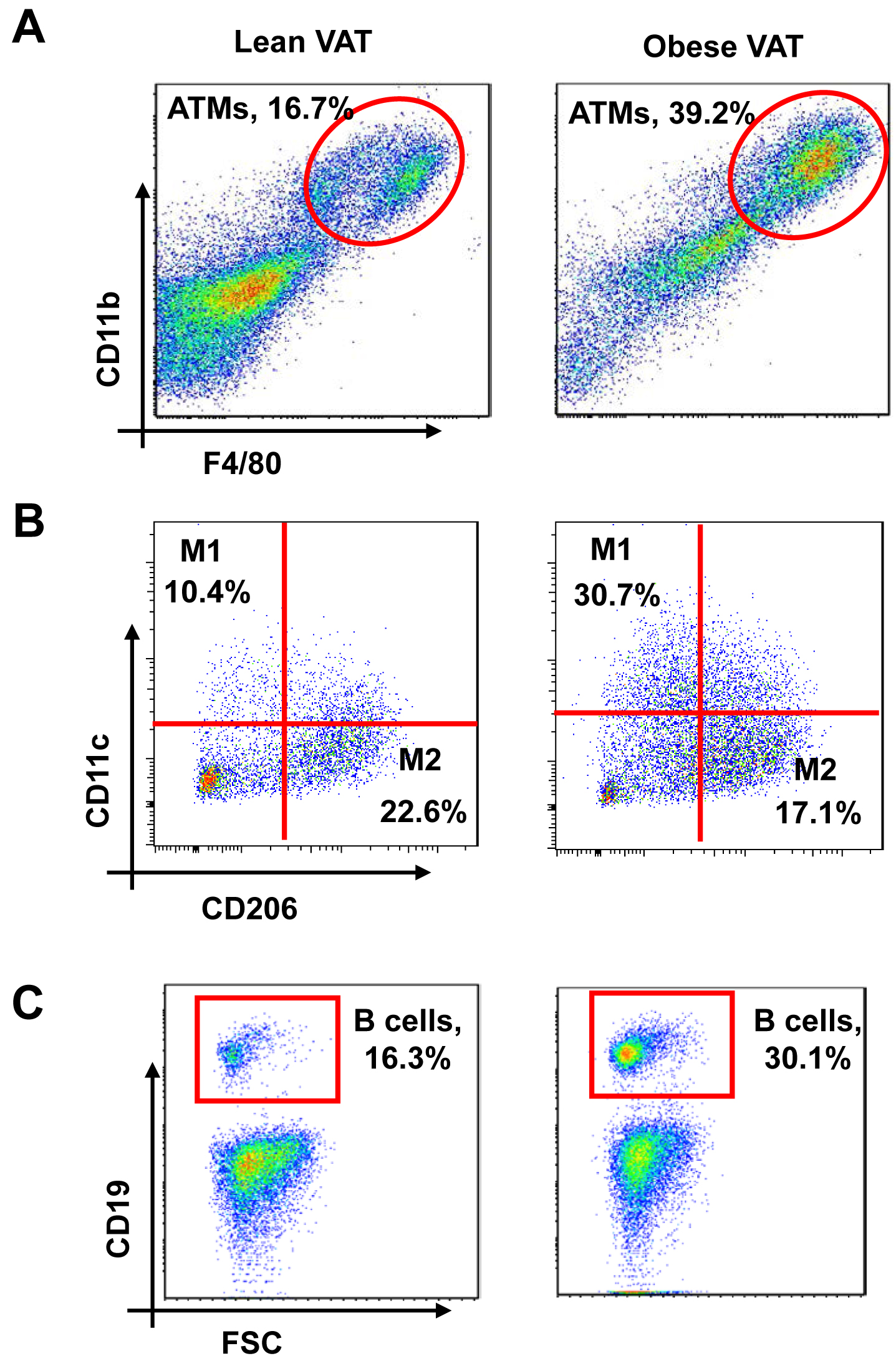
Figure 2. Flow cytometry analysis of immune cells of visceral adipose tissue (VAT). The SVCs were stained with fluorescent-conjugated antibodies against CD11b, F4/80, CD11c, CD206, CD19, and then examined by flow cytometry analysis. A. Adipose tissue macrophages (ATMs) were defined as CD11b+F4/80+ subpopulations and displayed as percentage of CD45+ cells. B. M1 and M2 ATMs were defined as CD11b+F4/80+CD11c+CD206- and CD11b+F4/80+CD11c-CD206+, respectively. These cell populations were shown as percentage of ATMs. C. B cells were defined as CD45+CD19+ and the cell population was presented as the percentage of CD45+ cells.
- Prepare 0.5-1 x 106 SVCs and resuspend in FACS staining buffer (see Recipes).
- RT-PCR
- SVCs are resuspended in Trizol (add 100 µl Trizol reagent to < 105 cells; add 300 µl Trizol reagent to < 106 cells) and incubated for 5 min at room temperature (RT).
- After centrifugation at 1,000 x g for 5 min (RT), transfer the supernatant to a new tube.
- Total RNAs are isolated using Direct-zol RNA kits.
- RNA concentration is measured by NanoDrop 1000 spectrophotometer.
- RNAs (500 ng) are converted to cDNA using High-Capacity cDNA Reverse Transcription Kit.
- The reverse transcription reactions are under the thermal cycling program: 25 °C for 10 min, then 37 °C for 120 min, then 85 °C for 5 min, and then 4 °C.
- The cDNA is mixed primers and SYBR Green PCR Master Mix in the MicroAmp Optical 96-well reaction plate.
- The real time PCR (qPCR) reaction is ran under the thermal cycling condition: 95 °C for 10 min, then 40 cycles (95 °C for 15 sec, and then 60 °C for 1 min).
- The RT-PCR results show that lean SVCs exhibit greater arginase 1 expression but less TNFα abundance than obese SVCs (Figures 3A and 3B).
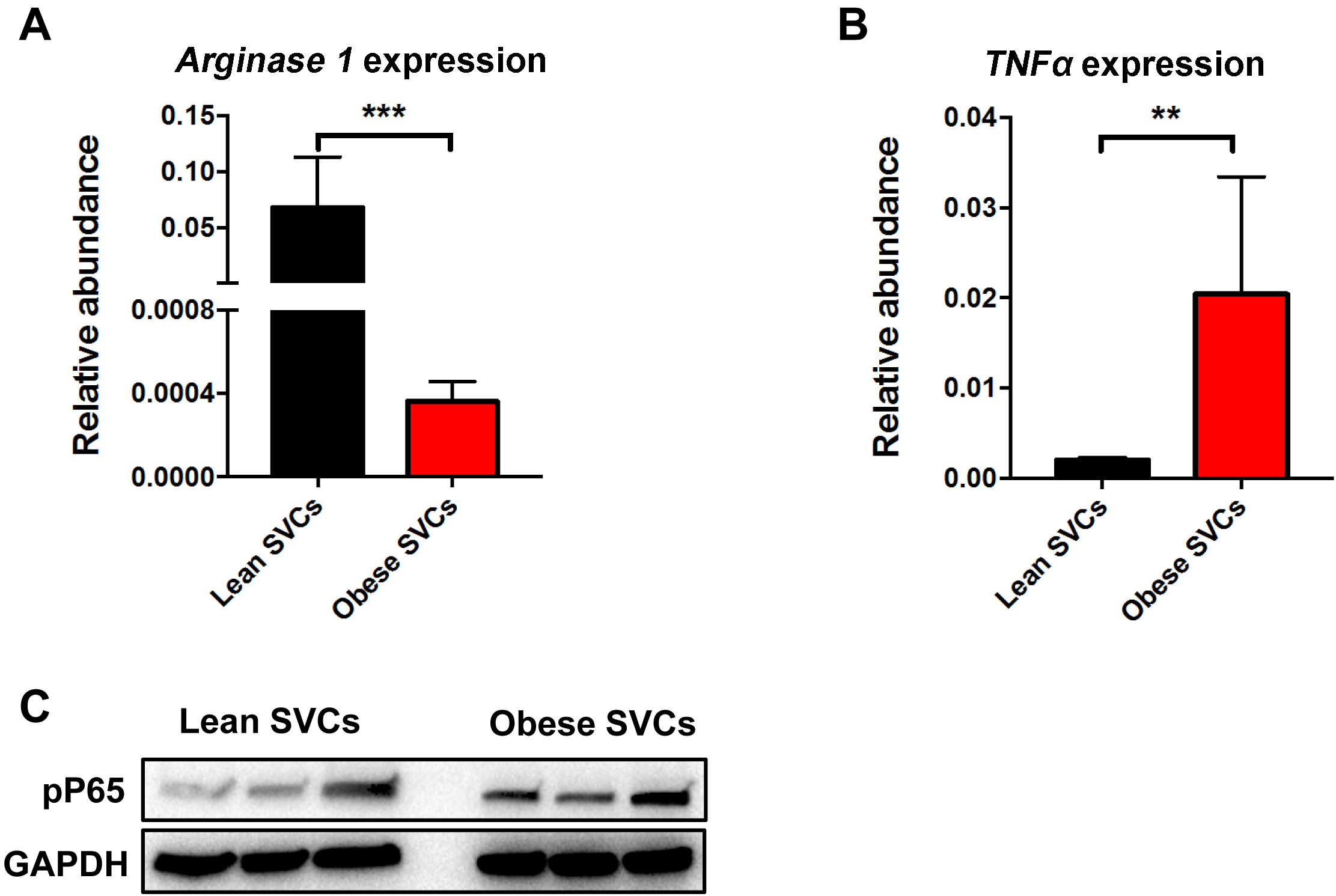
Figure 3. Inflammatory status of SVCs by RT-PCR analysis and Western blots. There was less arginase 1 (A) but more TNFα (B) abundance in obese SVCs, compared to the lean SVCs (qPCR primer information see Table 1). C. Activation of NF-κB signalling pathway of SVCs (without stimulation) was evaluated by Western blotting analysis (Antibodies information see Materials and Reagents #11). Data are presented as mean ± SEM. n = 3 per group. **P < 0.01, ***P < 0.001, Student’s t-test.
- SVCs are resuspended in Trizol (add 100 µl Trizol reagent to < 105 cells; add 300 µl Trizol reagent to < 106 cells) and incubated for 5 min at room temperature (RT).
- Western blots
See previous article by Zhang, 2011. The Western blotting analysis indicates that obese SVCs have higher level of phosphorylated P65 than lean SVCs (Figure 3C).
Recipes
- Digestion buffer (prepare freshly), 50 ml
50 ml HBSS
50 mg collagenase II
100 mg BSA
100 mM HEPES - Red blood cell lysis buffer (10x), 1 L
PBS (without calcium and magnesium)
8.3 g ammonium chloride (NH4Cl) (150 mM)
1.0 g potassium bicarbonate (KHCO3) (10 mM)
1.8 ml of 5% EDTA (0.1 mM) - FACS staining buffer, 100 ml
98 ml PBS (without calcium and magnesium)
2 ml FBS
0.1 g NaN3 - Complete culture medium
500 ml Iscove’s modified Dulbecco’s medium (IMDM)
50 ml FBS
5 ml penicillin-streptomycin (Thermo Fisher Scientific)
Acknowledgments
This protocol was adapted from Ying et al. (2017). This study was funded by the American Heart Association (16POST31350039 to W. Ying), the National Natural Science Foundation of China (81600610 to W. Ying).
References
- Johnson, A. M. and Olefsky, J. M. (2013). The origins and drivers of insulin resistance. Cell 152(4): 673-684.
- Lumeng, C. N., DelProposto, J. B., Westcott, D. J. and Saltiel, A. R. (2008). Phenotypic switching of adipose tissue macrophages with obesity is generated by spatiotemporal differences in macrophage subtypes. Diabetes 57(12): 3239-3246.
- Lumeng, C. N., Deyoung, S. M., Bodzin, J. L. and Saltiel, A. R. (2007). Increased inflammatory properties of adipose tissue macrophages recruited during diet-induced obesity. Diabetes 56(1): 16-23.
- Nguyen, M. T., Favelyukis, S., Nguyen, A. K., Reichart, D., Scott, P. A., Jenn, A., Liu-Bryan, R., Glass, C. K., Neels, J. G. and Olefsky, J. M. (2007). A subpopulation of macrophages infiltrates hypertrophic adipose tissue and is activated by free fatty acids via Toll-like receptors 2 and 4 and JNK-dependent pathways. J Biol Chem 282(48): 35279-35292.
- Romeo, G. R., Lee, J. and Shoelson, S. E. (2012). Metabolic syndrome, insulin resistance, and roles of inflammation–mechanisms and therapeutic targets. Arterioscler Thromb Vasc Biol 32(8): 1771-1776.
- Saltiel, A. R. and Olefsky, J. M. (2017). Inflammatory mechanisms linking obesity and metabolic disease. J Clin Invest 127(1): 1-4.
- Weisberg, S. P., McCann, D., Desai, M., Rosenbaum, M., Leibel, R. L. and Ferrante, A. W., Jr. (2003). Obesity is associated with macrophage accumulation in adipose tissue. J Clin Invest 112(12): 1796-1808.
- Winer, D. A., Winer, S., Shen, L., Wadia, P. P., Yantha, J., Paltser, G., Tsui, H., Wu, P., Davidson, M. G., Alonso, M. N., Leong, H. X., Glassford, A., Caimol, M., Kenkel, J. A., Tedder, T. F., McLaughlin, T., Miklos, D. B., Dosch, H. M. and Engleman, E. G. (2011). B cells promote insulin resistance through modulation of T cells and production of pathogenic IgG antibodies. Nat Med 17(5): 610-617.
- Ying, W., Wollam, J., Ofrecio, J. M., Bandyopadhyay, G., El Ouarrat, D., Lee, Y. S., Oh, D. Y., Li, P., Osborn, O. and Olefsky, J. M. (2017). Adipose tissue B2 cells promote insulin resistance through leukotriene LTB4/LTB4R1 signaling. J Clin Invest 127(3): 1019-1030.
- Zhang, H. (2011). Western blot for detecting phosphorylated STAT3. Bio Protoc Bio101: e125.
Article Information
Copyright
© 2017 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Vu, J. and Ying, W. (2017). Isolation and Analysis of Stromal Vascular Cells from Visceral Adipose Tissue. Bio-protocol 7(16): e2444. DOI: 10.21769/BioProtoc.2444.
Category
Immunology > Immune cell isolation > Stromal vascular cell
Cell Biology > Cell isolation and culture > Cell isolation
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link









