- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Measurement of Energy-dependent Rhodamine 6G Efflux in Yeast Species
Published: Vol 7, Iss 15, Aug 5, 2017 DOI: 10.21769/BioProtoc.2428 Views: 8982
Reviewed by: Yanjie LiLip Nam LOHSadri Znaidi

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
An Assay to Determine NAD(P)H: Quinone Oxidoreductase Activity in Cell Extracts from Candida glabrata
Anamika Battu [...] Rupinder Kaur
Nov 5, 2021 3084 Views
Isolation of Mitochondria from Ustilago maydis Protoplasts
Juan Pablo Pardo [...] Lucero Romero-Aguilar
Jan 5, 2022 3600 Views
Abstract
Rhodamine 6G is a highly fluorescent dye often used to determine the transport activity of yeast membrane efflux pumps. The ATP-binding cassette transporter KlPdr5p confers resistance to several unrelated drugs in Kluyveromyces lactis. KlPdr5p also extrudes rhodamine 6G (R6G) from intact yeast cells in an energy-dependent manner. Incubation of yeast cells in the presence of 2-deoxy-D-glucose (inhibitor of glycolysis) and R6G (mitochondrial ATPase inhibitor) leads to marked depletion of intracellular ATP pool (Kolaczkowski et al., 1996). An active KlPdr5p mediated extrusion of R6G from intact yeast cells can be followed by direct measurement of the fluorescence of extruded R6G in the assay buffer.
Keywords: Rhodamine 6GBackground
Multidrug efflux pumps are widely distributed and can be found in all living species. They represent an important mechanism of antimicrobial resistance. The ability to quantify the activity of efflux pumps is necessary for understanding of their contribution to physiological processes and assessment of the validity of potential therapeutics (e.g., efflux inhibitors) (Blair and Piddock, 2016). Methods for efflux activity measurements largely rely on two different mechanisms. Some methods directly measure the substrate efflux, i.e., how much of the substrate is pumped out, and others measure substrate molecule accumulation inside the cell, the levels of which is then used to infer efflux indirectly. However, the latter is less sensitive due to variable membrane permeability that alters dye influx rates (Blair and Piddock, 2016). Accumulation of R6G in growing C. albicans cells inversely correlates with the level of the ABC transporter Candida drug resistance 1 (CDR1) mRNA expression, establishing levels of intracellular R6G accumulation can be therefore used for identification of azole-resistant strains (Maesaki et al., 1999). Historically, this was carried out by measurements of accumulated radiolabelled-substrates. More recently, fluorescence-based methods are being used. Accumulation of fluorescent dye in a single cell can also be measured by flow cytometry. The benefit of this approach lies in the ability to measure variation in efflux activity among individual cells.
The protocol of the above described method involves preloading the cell population with a fluorescent substrate prior to the efflux assay. After the loading step, substrate accumulates within the cells at maximum concentration. Cells are then washed to remove the substrate. Subsequently, glucose is supplemented to the culture as a source of energy, and the fluorescence signal of substrate is monitored. The method is suitable for use with any yeast species (Borecka-Melkusova et al., 2008).
Materials and Reagents
- Pipettes tips
- Sterile inoculation loop
- 50-ml polypropylene centrifuge tubes (TPP Techno Plastic Products, catalog number: 91050 )
- 1.5-ml microcentrifuge tubes
- 96-well flat bottom with lid MicroWell plates (Thermo Fisher Scientific, Thermo ScientificTM, catalog number: 167008 )
- Yeast strain to be analysed
- Yeast extract (Biolife Italiana, catalog number: 4122202 )
- Bacto peptone (Biolife Italiana, catalog number: 4122592 )
- HEPES, free acid (AMRESCO, catalog number: 0511 )
- Sodium hydroxide (NaOH), 1 mol/L (Merck, catalog number: 109137 )
- 2-deoxy-D-glucose (Sigma-Aldrich, catalog number: D6134 )
- Rhodamine 6G (Sigma-Aldrich, catalog number: R4127 )
- D-glucose (Biolife Italiana, catalog number: 4125012 )
- YEPD rich growth medium (see Recipes)
- 20 mM glucose (see Recipes)
- 50 mM HEPES/NaOH assay buffer (see Recipes)
- 2-deoxy-D-glucose in HEPES/NaOH buffer (see Recipes)
- 10 mM rhodamine 6G (see Recipes)
Equipment
- Pipettes
Xplorer® 15-300 µl (Eppendorf, catalog number: 4861000031 )
Research® plus 2-0 µl (Eppendorf, catalog number: 3120000038 ) - pH meter (Xylem, WTW, model: inoLab® pH 7110 )
- Incubation shaker Unitron (Infors, model: Plus AJ252 )
- Haemocytometer
- Centrifuge 5804R (Eppendorf, model: 5804 R )
- Centrifuge Mikro 200R (Hettich Lab Technology, model: MIKRO 200 R )
- Spectrofluorometer Varioscan Flash (Thermo Fisher Scientific, Thermo ScientificTM, catalog number: 5250500 )
- Autoclave
Procedure
- Transfer a yeast colony from an agar plate with a sterile inoculation loop, and inoculate into 10 ml YEPD medium (see Recipes). Incubate the culture in an incubation shaker at 150 rpm, at 28 °C for 20 h. (see Note 1)
- Determine the cell number using haemocytometer. Add 1 ml (total amount of 5 x 108 cells) from overnight culture into 100 ml fresh YEPD medium.
- Incubate the cells in an incubation shaker at 150 rpm for 1.5-2 h at 28 °C until they reach the exponential phase (1 x 107 cells/ml). Monitor the cell density by counting the cell number in a haemocytometer.
- Harvest the cells by centrifugation at 3,000 x g for 5 min at laboratory temperature and discard the supernatant.
- Wash the cell pellet twice with 50 ml of sterile distilled water (room temperature, RT), then with 50 ml HEPES/NaOH (RT, see Recipes) assay buffer, vortex. Collect the cells by centrifugation (3000 x g, 5 min, RT).
- Resuspend the cell pellet (total cell number 109) in 50 ml HEPES/NaOH assay buffer (RT) containing 2-deoxy-D-glucose (2 mM) and R6G (10 µM) (see Recipes and Note 2).
- Incubate the cell suspension in an incubation shaker at 150 rpm for 2 h at 28 °C.
- Harvest the cells by centrifugation (3,000 x g; 5 min; 4 °C) and wash the pellet with 50 ml sterile distilled water (4 °C) first, then with 50 ml HEPES/NaOH (4 °C) assay buffer.
- Resuspend the pellet in 50 ml HEPES/NaOH (4 °C) assay buffer to 108 cells/ml.
- Transfer 5 ml of the cell suspension into a new 50 ml centrifuge tube and add 500 µl of glucose from a 20 mM stock solution (see Recipes) to a final concentration of 2 mM. For negative control experiment, glucose is not supplemented in the culture medium (see Note 3).
- At specified time intervals (0, 5, 10, 15, 20, 30 min, etc.), transfer 400 µl of cell suspension to a microcentrifuge tube and harvest cells by centrifugation at 10,000 x g, 1 min, 4 °C.
- Transfer 100 µl of supernatant into microtiter plate well. Supernatant is sufficient for three technical replicates. Store the samples on ice until the end of experiment.
- Measure the R6G fluorescence of the samples at laboratory temperature using the spectrofluorometer (the excitation wavelength of 515 nm; emission wavelength of 555 nm).
Data analysis
- Calculate the R6G concentrations using a standard curve. The following concentrations of R6G were used to generate the standard curve: 0, 50, 100, 250, 500 and 1,000 nM.
- Plot the obtained R6G concentrations. The results are expressed as the mean ± SD for three independent experiments. A representative curve is shown in Figure 1.
- These data were fit and display a linear model y = ax, where the coefficient ‘a’ represents the rate constant of the reaction. Control experiments were done without the added glucose.
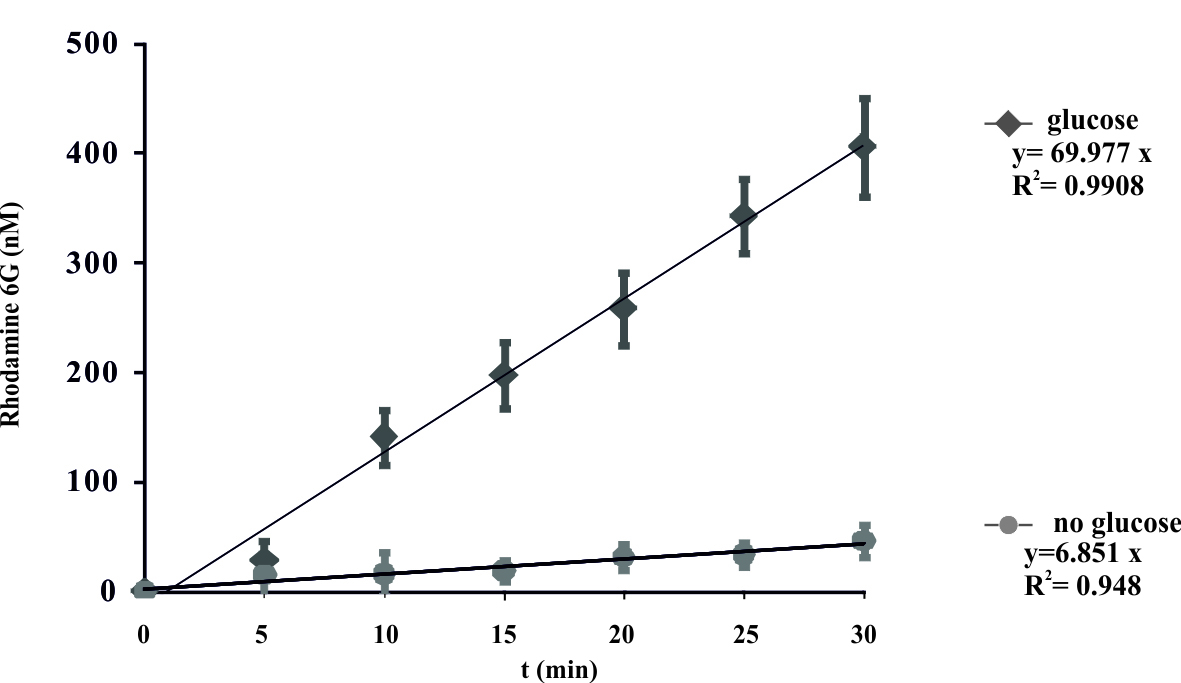
Figure 1. Energy-dependent ATP-binding cassette transporter-mediated efflux of R6G from yeast cells. De-energized cells were preloaded with R6G as described in the protocol. The efflux of R6G was followed by direct measurement of the fluorescence following the addition of glucose (2 mM) to suspension of K. lactis cells. The variation indicated for each time point is the standard error of the mean. The numbers on the x-axis represent time in minutes as independent variable, numbers on y-axis represent concentration of R6G (nM) in the supernatant. Glucose was added at time zero. (Konecna et al., 2016).
Notes
- Start the culture with fresh (up to 20 h old) inoculum. Starting concentration should be in the range 0.5-1 x 106 cells/ml.
- The Kluyveromyces lactis is usually grown at a temperature of 25-28 °C. The upper limit is approximately 40 °C.
- Freshly prepared rhodamine 6G solution is recommended.
- Control experiment without added glucose is recommended.
- To generate the calibration curve use three independent standard dilution sets of R6G weight.
- PBS buffer can be used instead of HEPES.
- Some spectrofluorometers require the use of Nunc FluoroNunc/LumiNunc plates (Nalge Nunc International, Rochester, NY).
Recipes
- YEPD rich growth medium
20 g/L 2% D-glucose
10 g/L 1% yeast extract
20 g/L 2% Bacto peptone
Sterilize using the autoclave (120 °C; 120 kPa for 20 min) - 20 mM glucose
36 g/100 ml
Sterilize using the autoclave (120 °C; 120 kPa for 20 min) - 50 mM HEPES/NaOH assay buffer
Dissolve 5.96 g HEPES in 500 ml distilled water, adjust pH to 7.0 with 1 N NaOH
Sterilize using the autoclave (120 °C; 120 kPa for 20 min) - 2-deoxy-D-glucose in HEPES/NaOH buffer
Dissolve 164.2 mg of 2-deoxy-D-glucose in 500 ml HEPES/NaOH pH 7 - 10 mM rhodamine 6G
Dissolve 4.8 mg R6G in 1 ml of sterile distilled water
Acknowledgments
This protocol was adapted from our previous studies (Goffa et al., 2014; Konecna et al., 2016). The work was supported by the Slovak Research and Development Agency grants APVV-0282-10 and VEGA 2/0111/15.
References
- Blair, J. M. and Piddock L. J. (2016). How to measure export via bacterial multidrug resistance efflux pumps. mBio 7: e00840-16.
- Borecka-Melkusova, S., Kozovska, Z., Hikkel, I., Dzugasova, V. and Subik, J. (2008). RPD3 and ROM2 are required for multidrug resistance in Saccharomyces cerevisiae. FEMS Yeast Res 8(3): 414-424.
- Goffa, E., Balazfyova, Z., Toth Hervay, N., Simova, Z., Balazova, M., Griac, P. and Gbelska, Y. (2014). Isolation and functional analysis of the KlPDR16 gene. FEMS Yeast Res 14(2): 337-345.
- Kolaczkowski, M., van der Rest, M., Cybularz-Kolaczkowska, A., Soumillion, J. P., Konings, W. N. and Goffeau, A. (1996). Anticancer drugs, ionophoric peptides, and steroids as substrates of the yeast multidrug transporter Pdr5p. J Biol Chem 271(49): 31543-31548.
- Konecna, A., Toth Hervay, N, Valachovic, M. and Gbelska, Y. (2016). ERG6 gene deletion modifies Kluyveromyces lactis susceptibility to various growth inhibitors. Yeast 33(12):621-632.
- Maesaki, S., Marichal, P., Vanden Bossche, H., Sanglard, D. and Kohno, S. (1999). Rhodamine 6G efflux for the detection of CDR1-overexpressing azole-resistant Candida albicans strains. J Antimicrob Chemother 44(1): 27-31.
Article Information
Copyright
© 2017 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Gbelska, Y., Toth Hervay, N., Dzugasova, V. and Konecna, A. (2017). Measurement of Energy-dependent Rhodamine 6G Efflux in Yeast Species. Bio-protocol 7(15): e2428. DOI: 10.21769/BioProtoc.2428.
Category
Microbiology > Microbial biochemistry > Protein > Activity
Biochemistry > Protein > Fluorescence
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link