- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Determination of Survival of Wildtype and Mutant Escherichia coli in Soil
Published: Vol 7, Iss 14, Jul 20, 2017 DOI: 10.21769/BioProtoc.2414 Views: 8863
Reviewed by: Esteban Paredes-OssesAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
Analysis of Heterocyst and Akinete Specific Glycolipids in Cyanobacteria Using Thin-layer Chromatography
Ritu Garg [...] Iris Maldener
Mar 20, 2022 2490 Views
Detecting Photoactivatable Cre-mediated Gene Deletion Efficiency in Escherichia coli
Yuta Koganezawa [...] Miki Umetani
Jun 5, 2023 1971 Views

Shipment of Cyanobacteria by Agarose Gel Embedding (SCAGE)—A Novel Method for Simple and Robust Delivery of Cyanobacteria
Phillipp Fink [...] Karl Forchhammer
Dec 5, 2024 1426 Views
Abstract
E. coli resides in the gastrointestinal tract of humans and other warm-blooded animals but recent studies have shown that E. coli can persist and grow in various external environments including soil. The general stress response regulator, RpoS, helps E. coli overcome various stresses, however its role in soil survival was unknown. This soil survival assay protocol was developed and used to determine the role of the general stress response regulator, RpoS, in the survival of E. coli in soil. Using this soil survival assay, we demonstrated that RpoS was important for the survival of E. coli in soil. This protocol describes the development of the soil survival assay especially the recovery of E. coli inoculated into soil and can be adapted to allow further investigations into the survival of other bacteria in soil.
Keywords: Soil survivalBackground
Escherichia coli is a Gram-negative, facultative anaerobe, belonging to the Enterobacteriaceae family, which inhabits the intestinal tract of humans, warm-blooded animals and reptiles (Berg, 1996; Gordon and Cowling, 2003). It can be transferred through water and sediments via faeces and is used as an indicator of faecal contamination in drinking and recreational water. The use of E. coli as a faecal indicator is based, at least in part, on the assumption that it exists transiently outside of the host gastrointestinal tract (Ishii and Sadowsky, 2008) and does not survive for a long time in the external environment. However, several studies have isolated E. coli from various natural environments such as municipal wastewater, freshwater, beach water, beach sand and soils (Jiménez et al., 1989; Brennan et al., 2010; Chiang et al., 2011; Byappanahalli et al., 2012; Zhi et al., 2016). The capacity of these E. coli strains to survive for long periods of time and grow in the external environment raises questions about the validity of its continued use as indicator of water quality (Brennan et al., 2010). To understand the genetic mechanism underlying the survival and persistence of E. coli in soil, we developed a soil survival assay to evaluate the role of the different genetic factors on soil survival. We investigated the role of the general stress response regulator, RpoS, in the survival of long-term soil persistent E. coli in soil. The ability of the rpoS mutant (COB583ΔrpoS) to survive in soil was compared with the wildtype strain (COB583) and RpoS was demonstrated to be important for the survival and long-term persistence of E. coli in soil (Somorin et al., 2016). Here, we present the detailed protocol for the soil survival assay and the recovery of E. coli from soil.
Materials and Reagents
Note: All reagents used were from the specified manufacturers (catalog numbers indicated). Nonetheless, the same reagents from different manufacturers are expected to produce similar results.
- Spatula
- Weighing boat (Sparks Lab Supplies, catalog number: BAL1822 )
- Gloves (Sparks Lab Supplies, catalog number: SAF5534X20 )
- Sterile micropipette tips (100-1,000 µl) (Abdos Labtech, catalog number: P10102 )
- Sterile micropipette tips (2-200 µl) (Abdos Labtech, catalog number: P10101 )
- Microcentrifuge tube (1.5 ml) (Abdos Labtech, catalog number: P10202 )
- Centrifuge tube (15 ml graduated) (Abdos Labtech, catalog number: P10402 )
- Cuvette (LP ITALIANA, catalog number: 112117 )
- 2 mm laboratory test sieve (B5410/1986, Endecotts, catalog number: 200SIW2.00 )
- 96-well plate (TC microwell 96U) (Thermo Fisher Scientific, Thermo ScientificTM, catalog number: 163320 )
- 90 mm Petri dishes (Abdos Labtech, catalog number: P10901 )
- Silty-Loam soil
- E. coli strains
- E. coli COB583 (Somorin et al., 2016)
- E. coli COB583ΔrpoS (Somorin et al., 2016)
- E. coli COB583 (Somorin et al., 2016)
- Luria-Bertani (LB) broth (Sigma-Aldrich, catalog number: L3022-1KG )
- Agar No. 2 (Lab M, catalog number: MC006 )
- MacConkey agar (Sigma-Aldrich, catalog number: M7408 )
- Distilled water (Milli-Q) (EMD Millipore)
- Phosphate buffered saline (PBS) (Sigma-Aldrich, catalog number: P4417-100TAB )
- LB broth (see Recipes)
- LB agar plates (see Recipes)
- PBS buffer (see Recipes)
- MacConkey agar plates (see Recipes)
Equipment
- 250 ml conical flask (VWR, catalog number: 214-1132 )
- -80 °C freezer
- Styrofoam 15 ml tube holder
- Bunsen burner
- Orbital shaker (Gallenkamp)
- Discovery comfort multichannel pipette (20-200 µl; HTL)
- Discovery comfort multichannel pipette (5-20 µl; HTL)
- Nichipet EX pipette (200-1,000 µl; Nichiryo)
- Nichipet EX Pipetman classic pipettes (20-200 µl; Nichiryo)
- Vortex mixer (Reax top; Heidolph)
- Biomate 3 spectrophotometer (Thermo Fisher Scientific, catalog number: 335904 )
Note: This product has been discontinued.
- Weighing scale (Sartorius, catalog number: BL120S )
Note: This product has been discontinued.
- Centrifuge (Eppendorf, model: 5418 )
- Labo autoclave (SANYO, model: MLS-3020U )
Procedure
- Measurement of cell concentration by optical density (OD600 nm)
- Streak out Escherichia coli COB583 wildtype and the rpoS deletion mutant (COB583ΔrpoS) from the stock culture kept in the -80 °C freezer onto LB agar (see Recipes) around a Bunsen burner and incubate them overnight (~16 h) at 37 °C.
- Inoculate one colony of each strain with a disposable sterile inoculating loop into 10 ml LB in a sterile conical flask around a Bunsen burner and incubate them at 37 °C for 6 h. Perform this step in triplicate for each strain.
- From the 6 h culture, take 100 µl and add it to 900 µl LB and determine the OD600 nm.
- Determine the volume of the 6 h culture required to give a starting OD600 nm of 0.05 in a 25 ml LB as follows:
C1V1 = C2V2
C1 = OD600 nm of the 6 h culture; V1 = volume of the 6 h culture
C2 = final OD600 nm required; V2 = final volume required
V1 = 0.05 x 25 ml/C1
- Make the cultures with starting OD600 nm of 0.05 in 25 ml LB for each strain and their respective replicates in sterile 250 ml conical flasks and incubate the flasks overnight on a shaker at 37 °C.
- On the following day, dilute the overnight cultures 1:10 in LB and determine the OD600 nm.
- Pipette 1 ml of each overnight culture into two separate sterile 1.5 ml tubes for each strain and their respective replicates.
- Centrifuge the cultures at 9,000 x g for 10 min at room temperature (23-25 °C) and remove the supernatants with sterile pipette.
- Wash the cell pellets by re-suspending them in sterile PBS (see Recipes) and centrifuge at 9,000 x g for 10 min. Repeat the washing step two more times.
- Resuspend the washed cell pellets in 1 ml of sterile PBS by pipetting up and down and vortexing and dilute 1:2; 1:5; 1:10; 1:20; 1:100; and 1:1,000 in sterile PBS to make 1 ml in sterile microcentrifuge tubes.
- Transfer each diluted cell suspension into a cuvette and determine the OD600 nm of the diluted cell suspensions of the E. coli strains and their respective replicates in a spectrophotometer three times and determine the mean value.
- Make serial dilutions of each cell suspension and spot 10 µl onto LB plates in triplicate and incubate the plates at 37 °C overnight.
- Count the colonies and use the numbers derived to estimate the cell count (CFU ml-1) and determine the mean cell count from the triplicate experiments.
- Plot the mean cell count against the mean OD600 nm of the respective dilutions of each strain in three independent experiments to obtain calibration curves from which desired cell numbers can be obtained. The standard curve of graph of the cell count versus OD600 nm is presented in Figure 1.
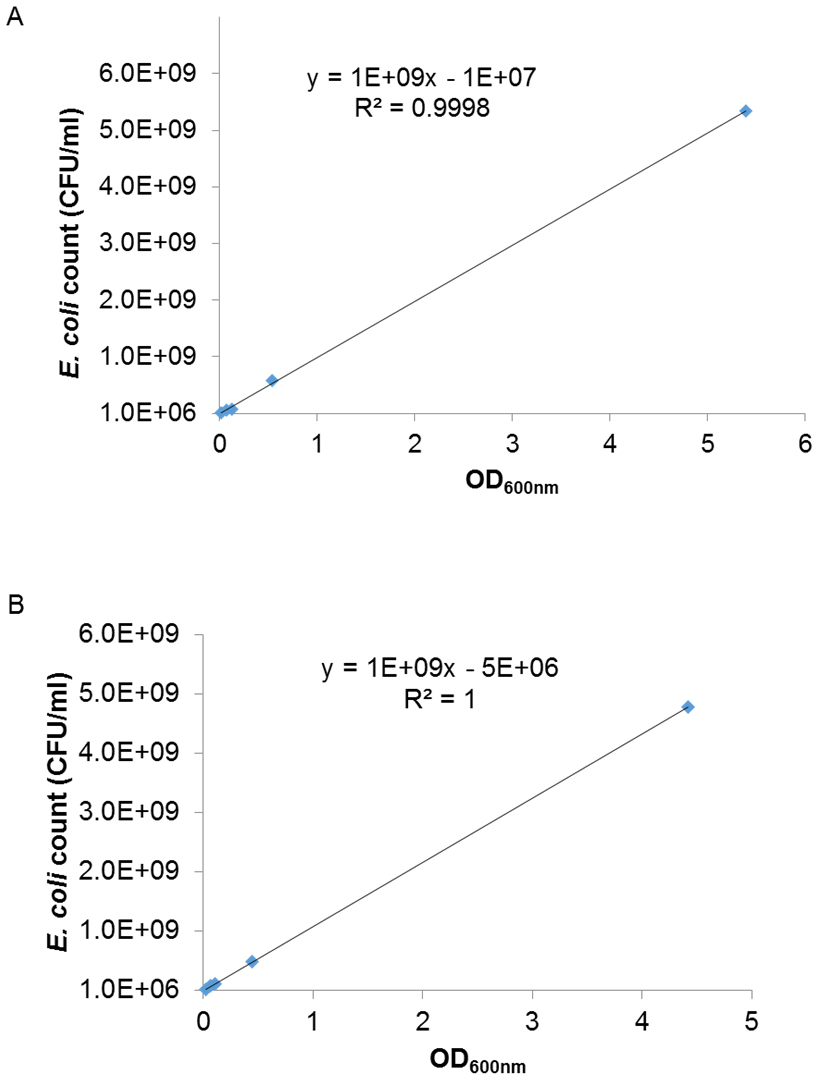
Figure 1. Standard curve of cell numbers versus OD600 nm of COB583 (A) and COB583ΔrpoS (B)
- Streak out Escherichia coli COB583 wildtype and the rpoS deletion mutant (COB583ΔrpoS) from the stock culture kept in the -80 °C freezer onto LB agar (see Recipes) around a Bunsen burner and incubate them overnight (~16 h) at 37 °C.
- Development of soil survival assay
- Inoculate one colony each of COB583 and COB583ΔrpoS into 25 ml LB in triplicate and incubate overnight at 37 °C.
- Dilute the resulting cultures 1:10 in LB and determine the OD600 nm.
- Add 1 ml of the overnight culture to two sterile Eppendorf tubes for each replicate of the two strains.
- Wash the cultures as described above (steps A8 and A9).
- Pool the resulting cell pellets and resuspend in 2 ml of sterile PBS in a 15 ml tube.
- Make 1:10 dilution (1 ml) of the each washed cell suspension to determine the OD600 nm.
- Based on the OD600 nm, prepare a normalised cell suspension with OD600 nm equivalent to 2 x 108 CFU ml-1 from each replicate.
- Make serial dilution of each normalised bacterial suspension and spot 10 µl of each dilution on LB agar in triplicate. Allow the spots to air-dry and incubate the plates overnight at 37 °C. Count the colonies and determine the mean cell counts.
- Sieve the soil to be used with 2 mm laboratory test sieve and weigh 1 g of the sieved soil into a 15 ml tube.
- Take 50 µl of each normalised bacterial suspension for each strain and inoculate it into the soil (to give ~1 x 107 CFU g-1 of soil). The uninoculated control should have 50 µl of sterile PBS added to it. The experiment was set up in triplicate for each strain.
- Cover the tubes with the lid and invert them 10 times by hand for proper mixing.
- Leave the tubes for about 10 min at room temperature (23-25 °C).
- Perform the recovery of the inoculated E. coli strains by adding 2 ml of sterile PBS to each 15 ml tube. Cover the tube with its lid, invert it three times and immediately vortex for 20 sec twice.
- Leave the tube on the workbench and allow the larger soil particles in the resulting soil slurry to settle out (2-10 min, depending on soil texture) (Figure 2).
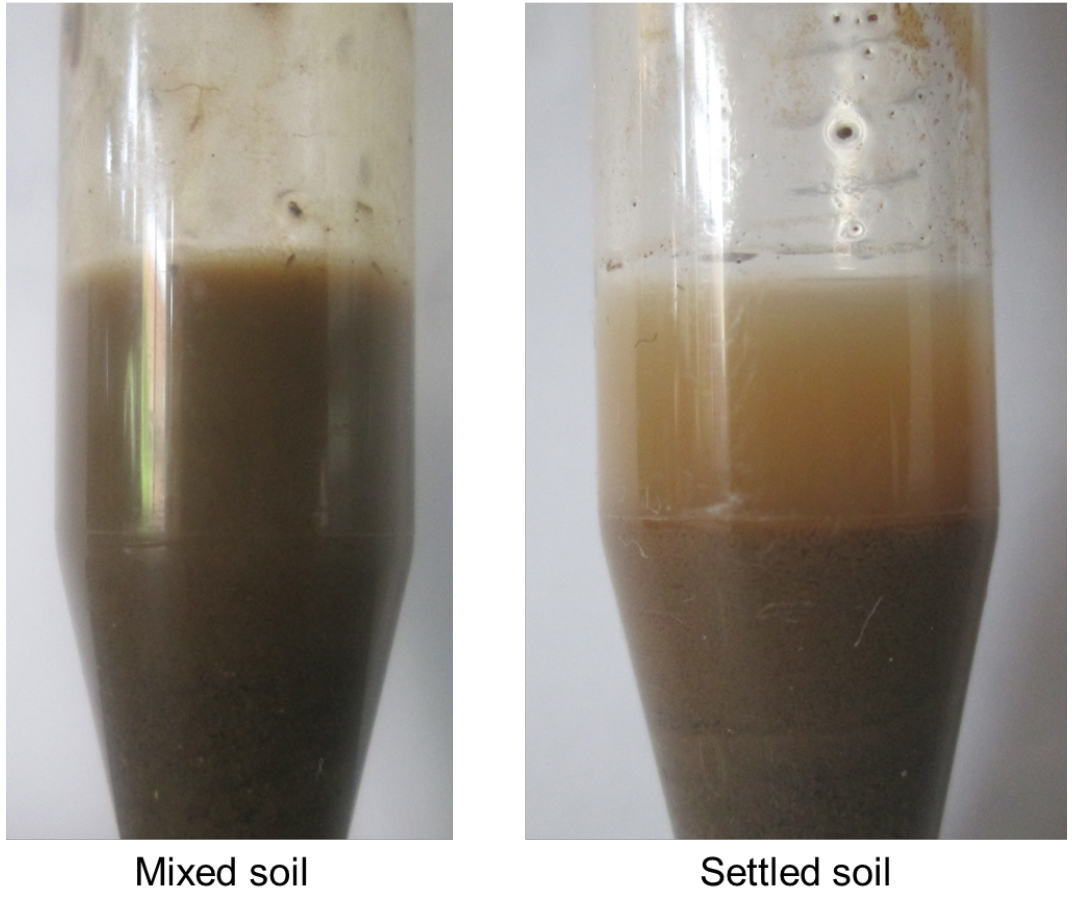

Figure 2. Experimental set-up for the soil survival assay with the soil slurry just after mixing and after settling out
- Take 20 µl from the supernatant of each soil slurry, place them in separate wells in a 96-well plate and make serial dilution (up to 10-6) from them.
- Take 10 µl of each dilution and spot onto MacConkey agar plates (see Recipes) in triplicate, and incubate plates overnight at 37 °C.
- Count the colonies and use the numbers derived to estimate the mean recovered cell count (CFU ml-1) from the triplicate experiments. The percentage recovery of cells was determined.
- Inoculate one colony each of COB583 and COB583ΔrpoS into 25 ml LB in triplicate and incubate overnight at 37 °C.
Data analysis
Cell count for each replicate is represented by the mean cell counts obtained from the three spots in each replicate. The mean and standard deviation of the cell counts from the normalised cell suspension (inoculum) and recovered cell counts were obtained from three independent replicates of each strain. The percentage of recovery ranges from 30-79% and the recovered cells from this procedure is presented in Figure 3.
![]()
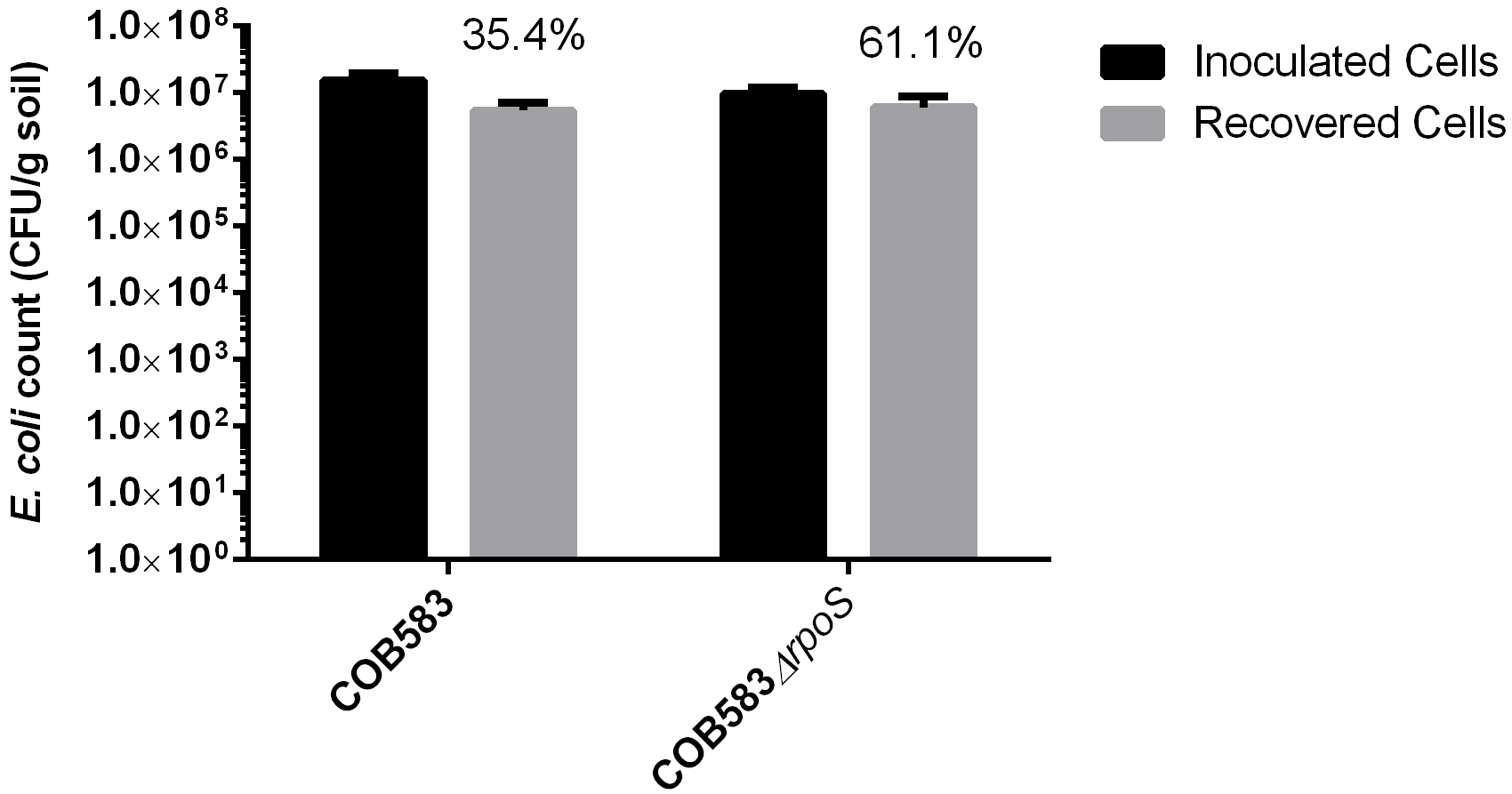
Figure 3. Recovery of Escherichia coli from soil. Percentage recovery is provided above the recovered cells.
Notes
- The optical density of the inoculum should be confirmed to be accurate before inoculating into soil.
- Care should be taken when inoculating the soil to ensure the pipette tips do not contact the soil directly, as it prevents the release of the entire content of the pipette tip.
- Care should be taken when spotting the cell suspension on agar plates so as not to puncture the agar with the pipette tips. This could vary the cell numbers arising from each spot.
- The soil to be used should be checked and confirmed not to contain E. coli prior to the soil survival assay so that recovered E. coli will only be attributable to inoculated E. coli. Serial dilutions of 1 g of the soil sieved through a 2 mm mesh should be plated on MacConkey agar and incubated at 37 °C to determine this.
- For long-term soil survival experiments, the tubes should be slightly capped to allow air exchange and incubated statically at the desired temperature. Multiple tubes should be set up for experiments requiring sampling at different time points since inoculated soils are destructively sampled (i.e., PBS was added and could not be reused at another time point) for E. coli recovery.
- No significant difference was seen when soil slurry was left to settle for 2 and 10 min before aliquots were taken for serial dilution and plating for cell count determination.
Recipes
- LB broth
Add 20 g LB powder to 1 L of distilled water and autoclave
- LB agar plates
Add 20 g LB powder and 15 g agar No. 2 powder to 1 L of distilled water and autoclave. When cool to touch (~ 45 °C), pour about 25 ml of the molten agar into Petri dishes and allow to solidify
- PBS buffer
Dissolve one phosphate-buffered saline tablet (Sigma-Aldrich) in 200 ml of distilled water and autoclave, giving a PBS buffer (pH 7.4)
- MacConkey agar plates
Add 50 g MacConkey agar powder to 1 L of distilled water and autoclave. When cool to touch (~45 °C), pour about 25 ml of the molten agar into Petri dishes and allow to solidify
Acknowledgments
The protocol was adapted from a previously published work (Somorin et al., 2016). The authors thank Camila Thorne for her help and advice with setting up the soil survival assay. This work was supported by an NUI Galway College of Science Ph.D. Fellowship and Thomas Crawford Hayes research award to Y.S.
References
- Anderson, K. L., Whitlock, J. E. and Harwood, V. J. (2005). Persistence and differential survival of fecal indicator bacteria in subtropical waters and sediments. Appl Environ Microbiol 71(6): 3041-3048.
- Berg, R. D. (1996). The indigenous gastrointestinal microflora. Trends Microbiol 4(11): 430-435.
- Brennan, F. P., O'Flaherty, V., Kramers, G., Grant, J. and Richards, K. G. (2010). Long-term persistence and leaching of Escherichia coli in temperate maritime soils. Appl Environ Microbiol 76(5): 1449-1455.
- Byappanahalli, M. N., Whitman, R. L., Shively, D. A., Sadowsky, M. J. and Ishii, S. (2006). Population structure, persistence, and seasonality of autochthonous Escherichia coli in temperate, coastal forest soil from a Great Lakes watershed. Environ Microbiol 8(3): 504-513.
- Byappanahalli, M. N., Yan, T., Hamilton, M. J., Ishii, S., Fujioka, R. S., Whitman, R. L. and Sadowsky, M. J. (2012). The population structure of Escherichia coli isolated from subtropical and temperate soils. Sci Total Environ 417-418: 273-279.
- Chiang, S. M., Dong, T., Edge, T. A. and Schellhorn, H. E. (2011). Phenotypic diversity caused by differential RpoS activity among environmental Escherichia coli isolates. Appl Environ Microbiol 77(22): 7915-7923.
- Gordon, D. M. and Cowling, A. (2003). The distribution and genetic structure of Escherichia coli in Australian vertebrates: host and geographic effects. Microbiology 149(Pt 12): 3575-3586.
- Jimenez, L., Muniz, I., Toranzos, G. A. and Hazen, T. C. (1989). Survival and activity of Salmonella typhimurium and Escherichia coli in tropical freshwater. J Appl Bacteriol 67(1): 61-69.
- Ishii, S., Ksoll, W. B., Hicks, R. E. and Sadowsky, M. J. (2006). Presence and growth of naturalized Escherichia coli in temperate soils from Lake Superior watersheds. Appl Environ Microbiol 72(1): 612-621.
- Ishii, S. and Sadowsky, M. J. (2008). Escherichia coli in the environment: Implications for water quality and human health. Microbes Environ 23(2): 101-108.
- Somorin, Y., Abram, F., Brennan, F. and O'Byrne, C. (2016). The general stress response is conserved in long-term soil-persistent strains of Escherichia coli. Appl Environ Microbiol 82(15): 4628-4640.
- Zhi, S., Banting, G., Li, Q., Edge, T. A., Topp, E., Sokurenko, M., Scott, C., Braithwaite, S., Ruecker, N. J., Yasui, Y., McAllister, T., Chui, L. and Neumann, N. F. (2016). Evidence of naturalized stress-tolerant strains of Escherichia coli in municipal wastewater treatment plants. Appl Environ Microbiol 82(18): 5505-5518.
Article Information
Copyright
© 2017 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Somorin, Y. and O'Byrne, C. P. (2017). Determination of Survival of Wildtype and Mutant Escherichia coli in Soil. Bio-protocol 7(14): e2414. DOI: 10.21769/BioProtoc.2414.
Category
Microbiology > Microbial physiology > Adaptation
Microbiology > Microbial cell biology > Cell viability
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










