- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Ciberial Muscle 9 (CM9) Electrophysiological Recordings in Adult Drosophila melanogaster
Published: Vol 7, Iss 14, Jul 20, 2017 DOI: 10.21769/BioProtoc.2401 Views: 8590
Reviewed by: Jihyun KimAlexandros C. KokotosAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
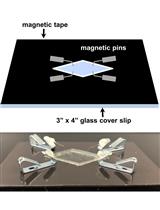
Estimation of the Readily Releasable Synaptic Vesicle Pool at the Drosophila Larval Neuromuscular Junction
Pragya Goel [...] Dion Dickman
Jan 5, 2019 8959 Views

Immuno-electrophysiology on Neuromuscular Junctions of Drosophila Third Instar Larva
Raffaella Klima [...] Fabian Feiguin
Feb 5, 2021 4159 Views
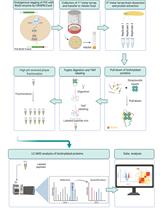
In vivo Characterization of Endogenous Protein Interactomes in Drosophila Larval Brain, Using a CRISPR/Cas9-based Strategy and BioID-based Proximity Labeling
Ezgi Uçkun [...] Ruth H. Palmer
Jul 5, 2022 4457 Views
Abstract
The complexity surrounding presynaptic recordings in mammals is a significant barrier to the study of presynaptic mechanisms during neurotransmission in the mammalian central nervous system (CNS). Here we describe an adult fly neuromuscular junction (NMJ), the ciberial muscle 9 (CM9) NMJ, which allows for the recording of both evoked (EPSPs) and spontaneous postsynaptic excitatory potentials (mEPSPs) at a mature glutamatergic synapse. Combined with CM9-specific genetic technologies, the CM9 NMJ provides a powerful experimental system to better understand the regulation of neurotransmitter release at a mature synapse.
Keywords: DrosophilaBackground
A significant hurdle in defining changes in presynaptic function during aging has been due to the lack of a simple model system for performing the electrophysiological recordings necessary to thoroughly characterize the release of neurotransmitter from the presynaptic nerve terminal. Existing rodent models suffer from the significant cost issues associated with aging studies and the technical difficulty of using electrophysiological recordings on single defined nerve terminals with consistent release parameters. To overcome these obstacles, we have pioneered a model synaptic system in the adult Drosophila for analyzing the effects of age on presynaptic function during neurotransmission, the CM9 NMJ located on the fly proboscis (Rawson et al., 2012; Mahoney et al., 2014; Mahoney et al., 2016) (Figure 4A). Briefly, the presynaptic arbor of the CM9 motor neuron (MN) converges upon the 15 muscle fibres of the CM9 muscle to form 35 individual distinct innervations (Rawson et al., 2012). The CM9 MN has been shown to be necessary for the contraction of the CM9 muscle and is the only source of glutamatergic input for the CM9 muscle (Kimura et al., 1986; Gordon and Scott, 2009). Given the highly-conserved nature of the mechanisms underlying synaptic vesicle (SV) release between flies and mammals, and the resemblance to the central synapses found in the mammalian CNS, this makes the CM9 NMJ a powerful model for investigating presynaptic function.
Materials and Reagents
- Sterile disposable filter (0.22 μm pore size, aPES membrane 19.6 cm2 CA membrane) (such as Corning® 250 ml Vacuum Filter/Storage Bottle System, Corning, catalog number: 430767 )
- Borosilicate glass capillary with filament (OD 1.50 mm, ID 0.86 mm) (Sutter Instrument, catalog number: BF150-86-10 )
- Borosilicate glass capillary without filament (OD 1.50 mm, ID 0.86 mm) (Sutter Instrument, catalog number: B150-86-10 )
- Silver wire (A-M Systems, catalog number: 782000 )
- Diamond coated bench stone (such as DMT 8 in Dia-Sharp bench stone)
- Drawn out P200 tip
- 10 ml syringe (Thermo Fisher Scientific, Thermo ScientificTM, catalog number: S7510-10 )
- Minutien pins (Fine Science Tools, catalog number: 26002-10 )
- Fine paint brush
- Flies of desired genotype and age
Note: UAS constructs can be driven within the CM9 motor neuron via the use of the E49-Gal4 driver (E49-Gal4 from Ulrike Heberlein’s Gal4 collection). - 50% Bleach
- Potassium chloride (KCl) (Sigma-Aldrich, catalog number: P9333-500G )
- Sodium chloride (NaCl) (Sigma-Aldrich, catalog number: S7653-250G )
- Calcium chloride (CaCl2) (Sigma-Aldrich, catalog number: 21115-250ML )
- Magnesium chloride (MgCl2) (Sigma-Aldrich, catalog number: 63069-500ML )
- Sodium bicarbonate (NaHCO3) (Sigma-Aldrich, catalog number: S5761-500G )
- Trehalose (Sigma-Aldrich, catalog number: T9531-25G )
- HEPES (Sigma-Aldrich, catalog number: H3375-25G )
- Sucrose (Sigma-Aldrich, catalog number: 84097-250G )
- Modified hemolymph like solution (HL3.1) (see Recipes)
Equipment
- Vannas spring scissors–2 mm cutting edge (Fine Science Tools, catalog number: 15000-03 )
- Tungsten Dissecting needle, 125 mm, Ultra fine (Roboz Surgical Instrument, catalog number: RS-6063 )
- Micro Dissecting needle holder, 5 ¼” (Roboz Surgical Instrument, catalog number: RS-6060 )
- Fine forceps (such as Dumont #5CO, Fine Science Tools, catalog number: 11295-20 and Dumont #3, Fine Science Tools, catalog number: 11231-30 )
- Dissecting stereoscopic zoom microscope (such as ZEISS, model: SteREO Discovery.V8 )
- Narrow Format Manipulator Systems (such as Sutter Instrument, model: ROE-200 )
- Horizontal Micropipette puller (such as programmable Flaming/Brown type micropipette puller, Sutter Instrument, model: P-97 ) with platinum filament
- Micro Forge electrode polisher (such as NARISHIGE, model: MF-900 )
- Upright confocal microscope (such as Olympus, model: BXS1WI ) with 10x and 40x water objectives
- Power lab 4/30 digital converter (ADInstruments, model: ML866 )
- Neuroprobe amplifier 1600 (A-M Systems, model: Model 1600 , catalog number: 680100)
- Stimulator (Digitimer, model: DS2A )
- Labchart 7 (ADInstruments)
Software
- Prism 6 (GraphPad software)
- Mini Analysis (comparable version 6.0.3, Synaptosoft Inc.)
Procedure
- Preparing silver chloride wire
Place silver wire in a 50% Bleach/50% ddH2O solution so that at least ¾ of the wire is immersed in the liquid. Leave the wire in the bleach/water solution for ~5 min to allow for sufficient chloridization to occur. - Preparing recording electrode–see JoVE video ‘Making patch pipettes and sharp electrodes with a programmable puller’ by Brown et al., 2008 for reference (see Notes).
- Insert a single silver-chloride wire into the electrode holder, ensuring that the end of the filament contacts the metal bottom of the holder.
- Carefully insert a glass capillary into the micropipette puller, and set puller program to create pipettes with adequate tip size and an internal resistance of 30 MΩ when filled with 3 M KCl. These parameters may be achieved by using the programmable P-97 Flaming/Brown puller (Figure 1B) (see Notes).
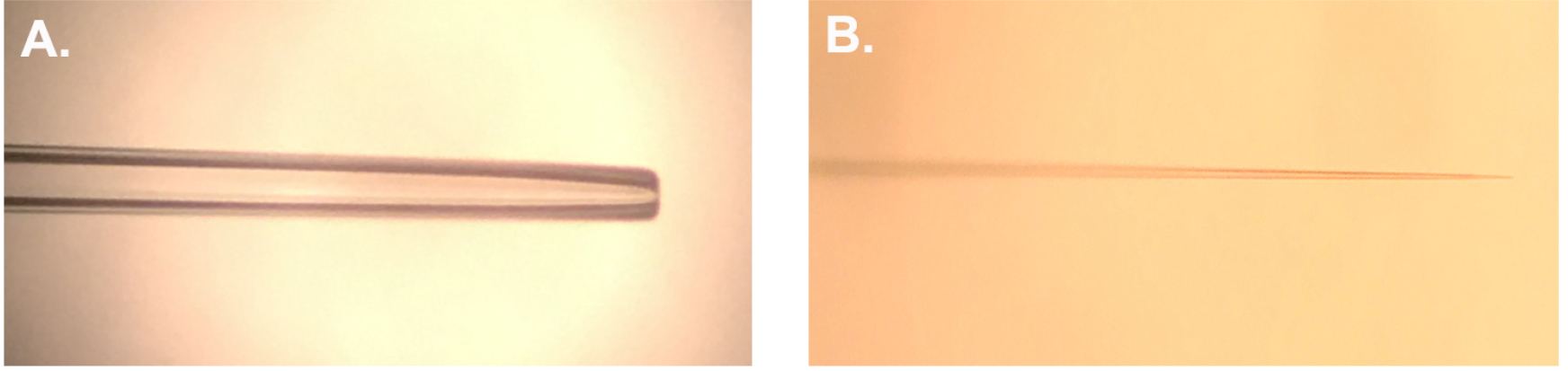

Figure 1. Prepared electrodes. A. Representative suction electrode polished to ~12-15 μm I.D. B. Representative recording electrode pulled using the parameters indicated in Notes. - Load the elongated tip syringe with 3 M KCl and dispense enough KCl into the glass capillary to sufficiently backfill the tip. Once the tip has sufficient KCl in place, fill the glass capillary with enough KCl to ensure the electrode is at least half full.
- Slide the silver-chloride wire into the glass capillary ensuring that it is immersed in the KCl.
- Insert recording electrode into the electrode micromanipulator that will be used for grounding (Figure 5.2).
- Preparing suction electrode
- Carefully insert a glass capillary tube, without filament, into the micropipette puller and set puller program to create pipettes with adequate tip size. Generally, the parameters used for creating the recording electrodes can also be used to make the suction electrode.
- Using a diamond coated bench stone (coarse side) break the tip sufficiently so that a flat opening of ~30 μm is created.
- Heat polish the electrode tip to reduce the I.D. to ~12-15 μm. Polishing is required to produce a sufficient taper to ensure that the CM9 MN does not come out of the electrode during recording and thus provides an adequate seal around the CM9 MN (Figure 1A).
- Once the suction electrode has been sufficiently polished, create a hole in the side of the electrode, about 1 cm down from the base of the tip, to allow for the placement of the silver wire. This is achieved by using the ‘fine’ side of the diamond coated bench stone to initially make a small hole. Once created, angle the suction electrode more acutely and continue scoring the glass. This will create a small gradient that will help to discourage bending of the silver wire required for stimulation (Figure 2).
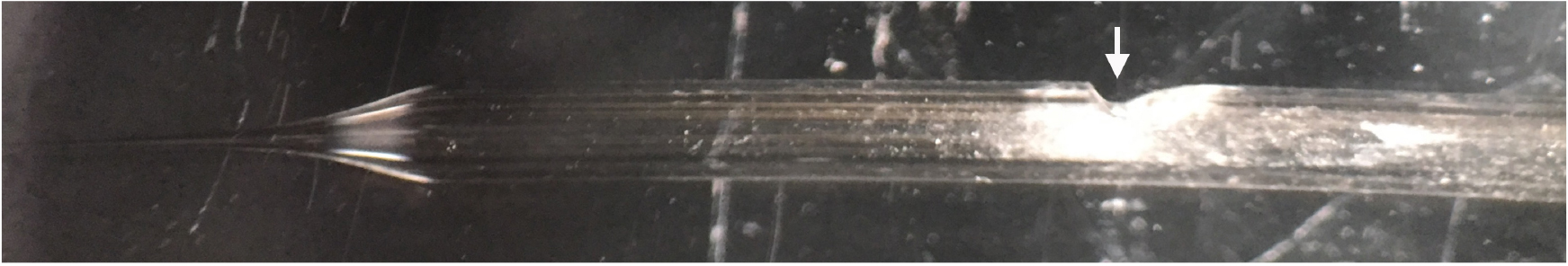

Figure 2. Representative image of the final suction electrode. Arrow indicates location of scored hole and gradient required to discourage bending of the silver wire required to stimulate the CM9 MN.
- Carefully insert a glass capillary tube, without filament, into the micropipette puller and set puller program to create pipettes with adequate tip size. Generally, the parameters used for creating the recording electrodes can also be used to make the suction electrode.
- Preparing fly for the experiment:
- Remove a single virgin female fly (see Notes) from food vial using a Pasteur pipette.
- Place the fly on ice until postural control is lost ~15/20 sec.
- Quickly transfer the fly to a small sylgard dissection surface and decapitate the head using a 30 G needle (Figure 3, right image).
- Place the back of the head on the dissection surface and pin the proboscis into an extended position (Figure 3, left image).
- Following successful pining (Figure 3, left image), cover the entire head in cold HL3 recording solution (see Recipes).
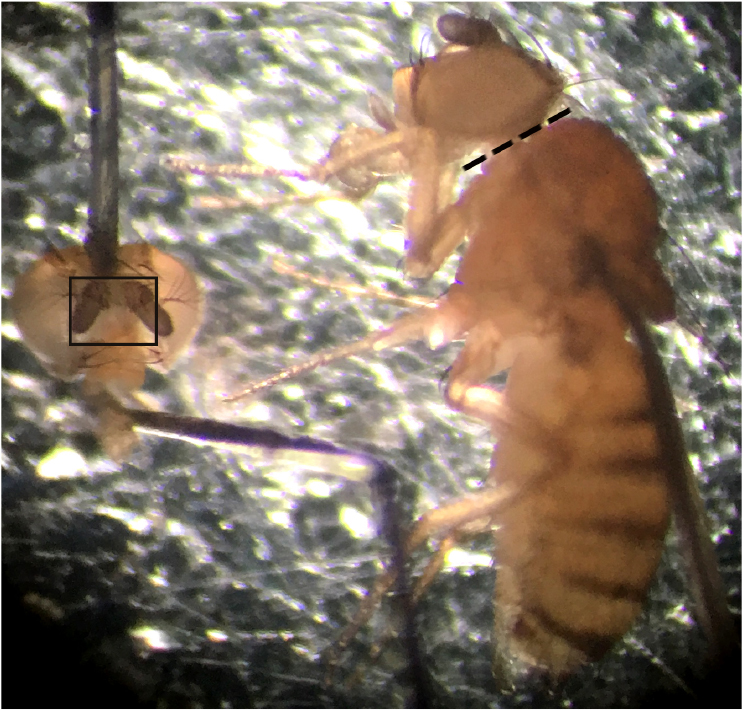

Figure 3. Representative image showing fly before (right) and after decapitation (left). Dashed line indicates where on the fly to cut to successfully decapitate the head. Image on the left shows how to correctly pin out the head prior to dissecting the cuticle–black box indicates area of cuticle to be removed in order to expose the CM9 group of muscles. - Dissect the anterior head cuticle containing the antennae from the preparation using a sharp tungsten needle to pierce the soft cuticle found underneath the faceplate and Vannas spring scissors to cut the cuticle. 5CO fine forceps are then used to remove any remaining tracheal air sacks that may remain after removing the faceplate. The use of 5CO forceps here to remove the remaining air sacks reduces the risk of damage to the nerve/muscles (Figure 3, left image and Figure 4B–black boxes).
- Re-pin the proboscis in a retracted position to increase the tension on the CM9 muscles (Figures 4B and 4C) (see Notes). The CM9 group of muscles run from the rostrum, identified by the brown/orange structure found at the end of the proboscis musculature located within the head cavity, to the bottom of the eye on either side of the head. Figure 4D indicates the translucency of the muscles and can be easily identified when compared to the more opaque fat bodies that remain within the head cavity post dissection.
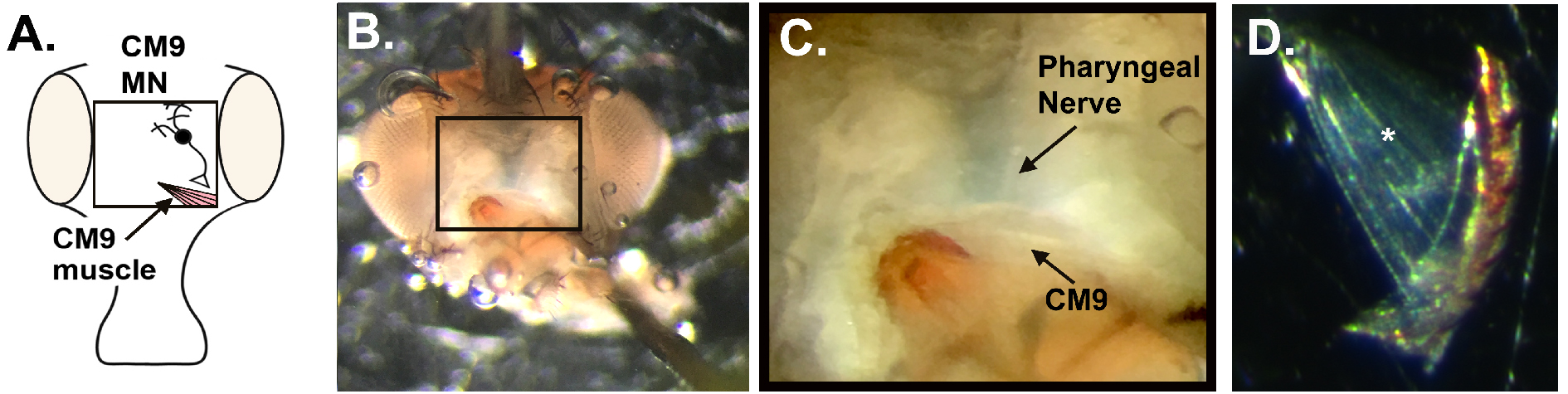
Figure 4. CM9 preparation. A. Diagram of Drosophila head indicating the approximate location of the CM9 muscle and motor neuron. B. Representative image of dissected fly head indicating the final positioning of the insect pins required to create sufficient tension on the muscles during recordings. C. Magnified image of CM9 muscles and MN (D) Dissected CM9 with * indicating the position of most cranial muscle fiber used for recordings. - Move the dissection dish to the slide holder on the microscope stand (Figure 5.4). Once in position, place both the grounding wire (Figure 5.5) and stimulation grounding wire (Figure 5.7) into the HL3 bath to ensure the formation of a complete electrical circuit.
- Using the 10x objective (Figure 5.3), slowly lower the suction electrode (Figure 5.6) into the HL3 bath and continue motion of the electrode until it comes into the plane of view of the preparation. Generally, as long as care is taken when lowering the electrode, the manipulator setting can be left on 1.
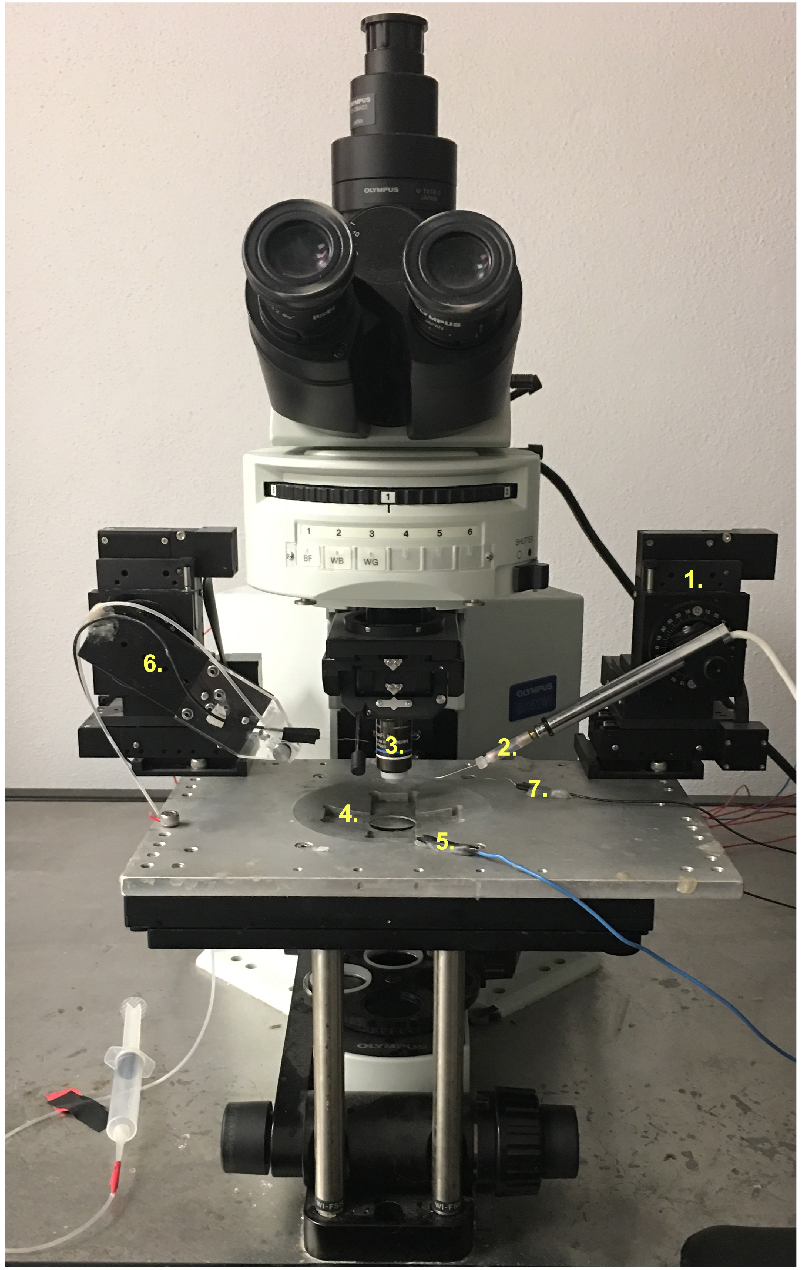

Figure 5. An overview of the rig setup. 1. Recording electrode head stage; 2. Recording. Electrode holder; 3. 10x/40x objective; 4. Microscope slide holder; 5. Grounding wire; 6. Suction electrode wire/holder; 7. Stimulator grounding wire. - Once positioned, suck a loop of the CM9 MN (Figure 4C–Pharyngeal nerve) into the polished suction electrode and fill with modified HL3 to create an en passant configuration (Figure 6A). Sometimes it will be necessary to switch to the 40x objective for the purposes of sucking up the CM9 nerve as it is not always easily observed under the 10x objective.
Note: There are 2 nerves that innervate the CM9 group of muscles but very little is known about the second smaller innervation and how it is involved in CM9 function. We do know that the second innervation is not a glutamatergic output due to the lack of positive vGLUT staining at the presynaptic nerve terminal. Care must be taken here to only suck up the CM9 MN and not the smaller innervation as this will impede the successful stimulation of the muscle.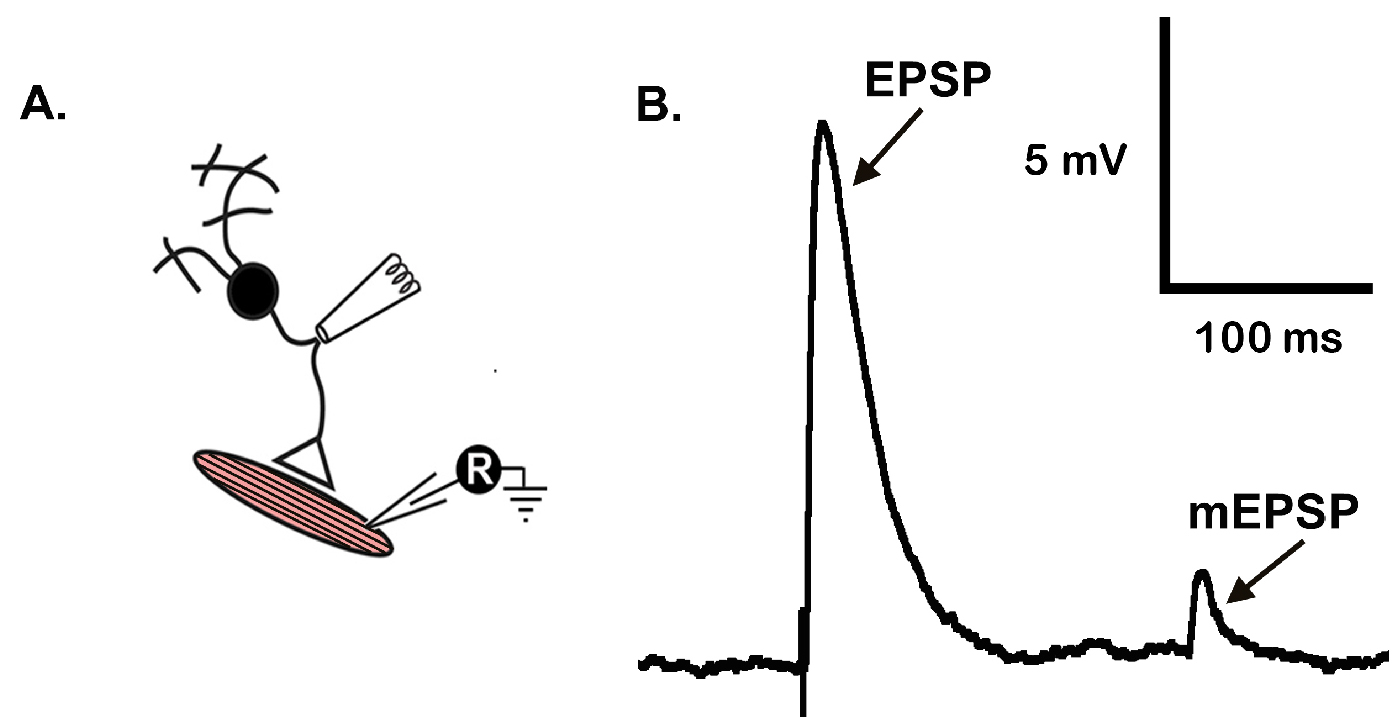
Figure 6. Representative mEPSP and EPSP traces from CM9 NMJ. A. Recording arrangement; B. Representative traces of an EPSP and mEPSP generated from a 7-day old animal.
- Remove a single virgin female fly (see Notes) from food vial using a Pasteur pipette.
- Intracellular recordings from CM9 muscle
- Slowly lower recording electrode into HL3 using coarse motion (setting 3/4) on the ROE-200 micromanipulator (Figure 5.1). Using the 10x objective, find the tip of the recording electrode and lower with caution until the electrode is in the plane of view of the preparation.
- Once the electrode is in the plane of view of the CM9 group of muscles finer motion is used to avoid impaling the muscle and breaking the electrode (setting 6/7 on ROE-200 micromanipulator). At this point, the objective should also be changed to 40x making sure to avoid hitting the electrodes already in position.
- Identify muscle to be recorded from, generally the most cranial CM9 muscle fiber accessible from the anterior position, and carefully impale the recording electrode into the muscle (Figure 4D).
- Slowly lower recording electrode into HL3 using coarse motion (setting 3/4) on the ROE-200 micromanipulator (Figure 5.1). Using the 10x objective, find the tip of the recording electrode and lower with caution until the electrode is in the plane of view of the preparation.
- Stimulating the CM9 MN
- Stimulate the CM9 at 0.5-5 V for 300 μsec. The observance of a presynaptic based action potential is confirmed by the observance of a distinct voltage threshold for EPSP appearance (Figure 6B).
- Generally, animals 7-35 days old have EPSPs amplitudes of ~9 mV (± 0.36) and ~12 mv (± 0.55) for 42 days old animals. mEPSP amplitudes remain relatively constant throughout the lifespan of the fly (~1.13 mV [± 0.43]) (see Mahoney et al., 2014 for complete values for 7-60 days old virgin females).
- Stimulate the CM9 at 0.5-5 V for 300 μsec. The observance of a presynaptic based action potential is confirmed by the observance of a distinct voltage threshold for EPSP appearance (Figure 6B).
Data analysis
- A Neuroprobe Amplifier Model 1600 is used in combination with a PowerLab 4/30 to amplify and digitize the data.
- LabChart 7 is used to record the data for at a rate of 1 Hz for 60 sec per fly taking care to record input resistance after the terminus of each 60 sec train.
- MiniAnalysis is used to analyze the amplitudes of both mEPSP and EPSPs (Figure 6B). From MiniAnalysis the values for EPSP and mEPSP amplitudes are tabulated in GraphPad Prism 6 from which appropriate statistical analysis is carried out to assess significance. Generally, all multiple comparisons are performed using a one-way ANOVA with a Bonferroni correction for multiple comparisons. All two-way comparisons are performed using a standard Student’s t-test.
Notes
- Fly husbandry: Virgin females are maintained at 25 °C and flipped every other day until needed for recording purposes. It is recommended that the flies used in the experiments do not grow in an over-populated environment (generally ~10-20 virgin females per vial).
- Muscle tension: To insure successful recordings, it is necessary to place sufficient tension on the CM9 group of muscles. This can be achieved by gently pushing the proboscis, via moving of the dissecting pin impaled proboscis, posteriorly towards the brain and sweeping the pin towards the base of the eye. Depending on which set of CM9 muscles are recorded from this will determine to which side the proboscis is repositioned.
- Electrode parameters: Heat–611, Pull–70, Vel–60, Del–130.
- A sharp recording electrode, similar to those used in this protocol, are used primarily to measure the electrical current passing through, or the voltage across, a neuronal/cellular membrane. These electrodes differ from classical whole cell patch pipettes in that the tip needs to be sufficiently sharp enough to penetrate the cellular membrane. On the other hand, the function of the suction electrode is to create a seal around the axon to allow for the detection of local circuit currents flowing around the axon as the action potential propagates, this can then be observed in the muscle via the use of the recording electrode. Depending on whether current clamp or voltage clamp mode is used this will determine the observance of either an EPSP (current clamp) or an EPSC (endplate postsynaptic current–voltage clamp).
Recipes
- Modified hemolymph like solution (HL3.1)
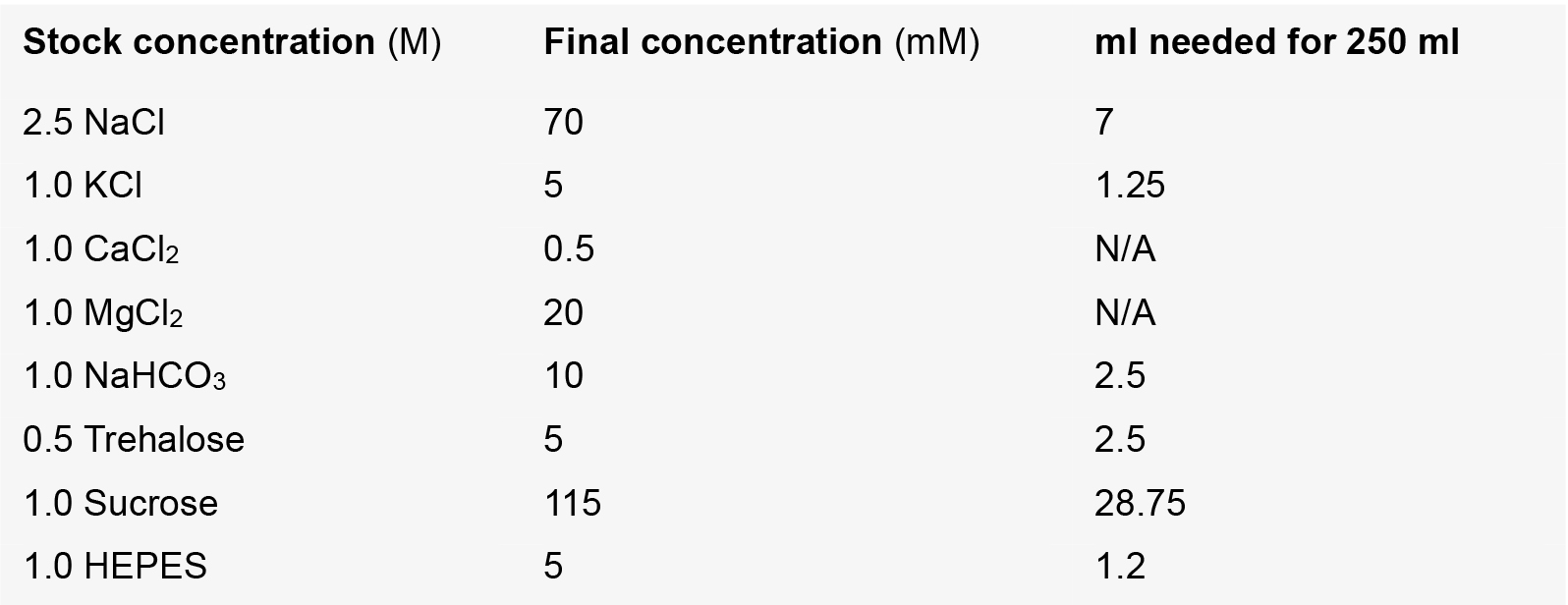
Note: Remaining volume needed for 250 ml is made up of ddH2O. Both CaCl2 and MgCl2 are added to 10 ml HL3 prior to recording. Adjust pH to 7.31 before use.
References
- Brown A. L., Johnson, B. E., Goodman, M. B. (2008). Making patch-pipettes and sharp electrodes with a programmable puller. J Vis Exp (20): e939.
- Gordon, M. D. and Scott, K. (2009). Motor control in a Drosophila taste circuit. Neuron 61(3): 373-384.
- Kimura, K. I., Shimozawa, T. and Tanimura, T. (1986). Isolation of Drosophila mutants with abnormal proboscis extension reflex. J Exp Zool 239(3): 393-399.
- Mahoney, R.E., Azpurua, J. and Eaton, B.A. (2016). Insulin signaling controls neurotransmission via the 4eBP-dependent modification of the exocytotic machinery. Elife 5.
- Mahoney, R. E., Rawson, J. M. and Eaton, B. A. (2014). An age-dependent change in the set point of synaptic homeostasis. J Neurosci 34(6): 2111-2119.
- Rawson, J.M., Kreko, T., Davison, H., Mahoney, R., Bokov, A., Chang, L., Gelfond, J., Macleod, G. T. and Eaton, B. A. (2012). Effects of diet on synaptic vesicle release in dynactin complex mutants: a mechanism for improved vitality during motor disease. Aging Cell 11(3): 418-427.
Article Information
Copyright
© 2017 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Readers should cite both the Bio-protocol article and the original research article where this protocol was used:
- Eaton, B. A. and Mahoney, R. E. (2017). Ciberial Muscle 9 (CM9) Electrophysiological Recordings in Adult Drosophila melanogaster. Bio-protocol 7(14): e2401. DOI: 10.21769/BioProtoc.2401.
- Mahoney, R. E., Rawson, J. M. and Eaton, B. A. (2014). An age-dependent change in the set point of synaptic homeostasis. J Neurosci 34(6): 2111-2119.
Category
Neuroscience > Cellular mechanisms > Synaptic physiology
Cell Biology > Cell signaling > Intracellular Signaling
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link









