- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Exopolysaccharide Quantification for the Plant Pathogen Ralstonia solanacearum
Published: Vol 7, Iss 10, May 20, 2017 DOI: 10.21769/BioProtoc.2289 Views: 8532
Reviewed by: Arsalan DaudiHiroyuki HiraiKanika Gera

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
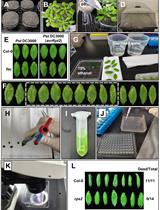

Bacterial Infection and Hypersensitive Response Assays in Arabidopsis-Pseudomonas syringae Pathosystem
Minhang Yuan and Xiu-Fang Xin
Dec 20, 2021 5367 Views
Protocol for Inoculation of PGPR Staphylococcus sciuri to Seeds and Seedlings of Rice and Tomato Plants for Increased Root and Shoot Growth
Girija Somna [...] Dinakar Challabathula
Mar 20, 2025 2264 Views
In Silico Prediction and In Vitro Validation of Bacterial Interactions in the Plant Rhizosphere Using a Synthetic Bacterial Community
Arijit Mukherjee [...] Sanjay Swarup
Nov 5, 2025 1705 Views
Abstract
Soluble exopolysaccharide is a major virulence factor produced by the plant pathogen Ralstonia solanacearum. Its massive production during plant infection is associated with the arrest of water flow in xylem vessels leading eventually to plant death. The composition of this heavy macromolecule includes mainly N-acetylgalactosamine. Here we describe a colorimetric method for quantitative determination of the soluble exopolysaccharide present in culture supernatant of R. solanacearum.
Keywords: Ralstonia solanacearumBackground
The plant pathogen Ralstonia solanacearum produces exopolysaccharide under the control of quorum sensing system, i.e., at high cell density, above 5 x 107 cell ml-1 (Flavier et al., 1997). The sugar content of the exopolysaccharide includes galactosamine, glucose, and rhamnose in the ratio of 10:2.5:1 (Drigues et al., 1985). A protocol for a reliable extraction and quantification of the exopolysaccharide from culture supernatant was initially developed by Brumbley and Denny (1990) and was updated recently by Peyraud et al. (2016). The quantification is based on the determination of hexoseamine content of the macromolecule using an adapted Elson and Morgan assay (Elson and Morgan, 1933; Gatt and Berman, 1966). Exopolysaccharides containing N-acetyl-D-galactosamine are produced by diverse Gram-negative or Gram-positive bacteria (Vaningelgem et al., 2004; Balzaretti et al., 2017) and also some fungi (Lee et al., 2015), so this protocol may also be applicable to such organisms.
Materials and Reagents
- Millex®-GP 33 mm syringe filter with a polyethersulfone membrane and 0.22 µm pore size (EMD Millipore, catalog number: SLGP033RB )
- Polypropylene microcentrifuge tubes of 2.0 ml Eppendorf® Safe-Lock (Eppendorf, catalog number: 0030120094 )
- Paper towel
- Aluminum foil
- Pipette tips
- 15-ml conical polypropylene Greiner centrifuge tubes (Greiner Bio One International, catalog number: 188271 )
- 50-ml conical polypropylene Greiner centrifuge tubes (Greiner Bio One International, catalog number: 227261 )
- Bridges for microcentrifuge tube (Milian, catalog number: 045427 )
- Gloves
- R. solanacearum strain GMI1000 (Salanoubat et al., 2002). This strain can be retrieved from the CIRM Biological Resource Center (http://www6.inra.fr/cirm_eng/CFBP-Plant-Associated-Bacteria) in the CIRM-BP collection (code CFBP6924)
- Acetone > 99.8% AnalaR NORMAPUR® (VWR, catalog number: 20066.296 )
- Milli-Q water obtained at a resistivity of 18.2 MOhm cm at 25 °C
- Acetyl acetone > 99% AnalaR NORMAPUR® (VWR, catalog number: 20092.23 0)
- Ethanol, 99.8% (Sigma-Aldrich, catalog number: 02851 )
- 4-(dimethylamino)benzaldehyde, 98% (Erlich’s reagent) (Sigma-Aldrich, catalog number: 109762 )
- Sodium L-glutamic acid monohydrate (Sigma-Aldrich, catalog number: 49621 )
- Iron(II) sulfate heptahydrate (FeSO4·7H2O) (Sigma-Aldrich, catalog number: F8263 )
- Ammonium sulfate, (NH4)2SO4 (Sigma-Aldrich, catalog number: A4418 )
- Magnesium sulfate heptahydrate (MgSO4·7H2O) (Sigma-Aldrich, catalog number: M5921 )
- Potassium phosphate monobasic (KH2PO4) (Sigma-Aldrich, catalog number: P5655 )
- Potassium hydroxide (KOH) (Sigma-Aldrich, catalog number: 60377 )
- Sodium chloride (NaCl) > 99.5% AnalaR NORMAPUR® (VWR, catalog number: 27810.295 )
- N-acetylgalactosamine, 98% (Sigma-Aldrich, catalog number: A2795 )
- Hydrochloric acid (HCl), 37%, AnalaR NORMAPUR® (VWR, catalog number: 20252.29 0)
- Sodium carbonate (Na2CO3) > 99.9 AnalaR NOR MAPUR® (VWR, catalog number: 27771.29 0)
- Minimal medium (see Recipes)
- 5 M NaCl solution (see Recipes)
- Standard samples of N-acetyl galactosamine (see Recipes)
- N-acetylgalactosamine standard stock solution (see Recipes)
- 2% acetyl acetone, 1.5 M Na2CO3 solution (see Recipes)
- Erlich’s reagent (see Recipes)
Equipment
- Dry bath heater with dual block Corning® (Corning, model: Corning® LSETM Digital Dry Bath Heater, catalog number: 6786-DB )
- Fume hood
- Eppendorf® microcentrifuge 5415 R (Eppendorf, model: 5415 R )
- Mini microcentrifuges Corning® (Corning, model: Corning® LSETM Mini Microcentrifuge, catalog number: 6766 )
- Vortex for microcentrifuge tubes
- UltrospecTM 2100 pro UV/Visible spectrophotometer (GE Healthcare, model: UltrospecTM 2100 pro UV/Visible Spectrophotometer , catalog number: 80211221)
- 10 ml volumetric flask
Software
- Appropriate software (SciDavis, Excel, etc.)
Procedure
- Culture and sample preparation
- Run a liquid R. solanacearum culture. Culture is performed in an Erlenmeyer containing 100 ml liquid minimal medium inoculated at 0.01 optical density at 600 nm, agitated at 180 rpm at a temperature of 28 °C.
- Collect 1 ml culture broth in the exponential or stationary phase. The cell culture must be at least above an optical density of 0.1 (ideally around 0.6) at 600 nm or 5 x 108 cell ml-1 to ensure passing the quorum sensing threshold and thus production of exopolysaccharide.
- Filter the cell broth with a 0.22 µm filter and collect the supernatant in a 2.0 ml microcentrifuge tube.
- Store the sample at -20 °C or run the extraction right away.
- Run a liquid R. solanacearum culture. Culture is performed in an Erlenmeyer containing 100 ml liquid minimal medium inoculated at 0.01 optical density at 600 nm, agitated at 180 rpm at a temperature of 28 °C.
- Exopolysaccharides extraction
- To precipitate exopolysaccharide, mix 0.2 ml supernatant + 0.004 ml 5 M NaCl solution + 0.8 ml acetone in a 2.0 ml microcentrifuge tube. Mix well with a vortex for 10 sec and place the sample at 4 °C overnight (around 12 h).
- Centrifuge the sample for 10 min at 16,000 x g (13,000 rpm) and at 4 °C and then discard the supernatant. Invert the tube on a paper towel to drain the pellet well during 15 min. Then, add 0.2 ml Milli-Q water to dissolve the pellet.
- The sample can be stored at 4 °C for few days (optional) until quantification.
- On the same day that the assay is to be performed, heat samples at 65 °C on dry bath heater for 10 min. Vortex well and then spin for 5 min at 16,000 x g (13,000 rpm), 4 °C. Collect the supernatant and place it in a new 2.0 ml microcentrifuge tube. The preparation is now referred to as the EPS (exopolysaccharide) sample.
- To precipitate exopolysaccharide, mix 0.2 ml supernatant + 0.004 ml 5 M NaCl solution + 0.8 ml acetone in a 2.0 ml microcentrifuge tube. Mix well with a vortex for 10 sec and place the sample at 4 °C overnight (around 12 h).
- Exopolysaccharide quantification
- Turn on the dry bath heater positioned in a fume hood. Temperature needs to be at 115 °C before adding the samples.
- Use the 200 μl of the EPS sample or you can gather EPS samples up to 450 ml. The chosen volume of the sample i is referred to be Vi in the data treatment step, see below.
- Add 0.15 ml concentrated HCl (37%) to i) the EPS sample or gathered EPS sample up to 0.45 ml, ii) the standard samples, see Table 1. Add enough Milli-Q water to make 0.6 ml total. Mix with a vortex and close the tubes tightly by adding bridges.
Table 1. Preparation of N-acetylgalactosamine standard samples. We recommend performing a standard curve at each run of the experiment, or at least running the 10 µg standard sample and the blank, see Table 1. The optical density at 530 nm in the 10 µg standard sample should be around 0.2.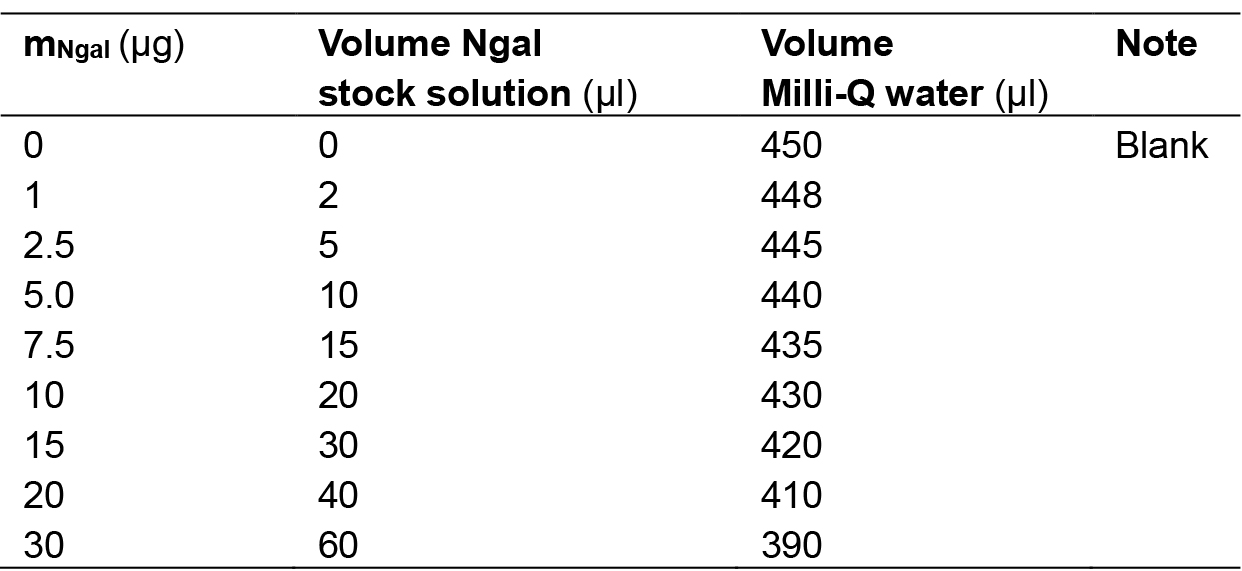
- Transfer the tubes to the dry bath heater. Maintain temperatures in the dry bath heater at 115 °C for 30 min. Cover the tubes and the dry bath heater with an aluminum foil if necessary.
- Transfer the tubes to a rack and cool to room temperature. Then centrifuge for 5 sec the tubes at 2,000 x g (6,000 rpm) using a mini microcentrifuge so the liquid on the top falls back.
- SLOWLY add, drop by drop using a pipette, 0.4 ml of 2 M Na2CO3 solution to each tube. Addition of this solution to the sample tubes will cause a very bubbly reaction. Mix gently but completely.
- Add 0.5 ml of a freshly made solution containing 2% acetyl acetone in 1.5 M Na2CO3. Vortex each tube, then open them to remove last CO2 released before heat in order to avoid opening of the tubes. Well close the tubes by adding bridges. Heat at 100 °C in a dry bath heater for 20 min.
- Transfer the tubes to a rack and cool to room temperature. Then centrifuge the tubes quickly so the liquid on the top falls back.
- Transfer liquid into a 15 ml tube and then add 1.0 ml 99.8% ethanol. Mix well.
- SLOWLY add 0.5 ml Erlich’s reagent solution. Addition of this solution to the sample tubes will cause a very bubbly reaction, like champagne. Once bubbles slow down, first mix gently and then vortex well.
- Let the tubes stand for about 30 min in the dark or with aluminum fold around. Then, read optical density of the solution at 530 nm. Use the water sample of the standard samples as blank, see Table 1.
- Turn on the dry bath heater positioned in a fume hood. Temperature needs to be at 115 °C before adding the samples.
Data analysis
- Plot the measured OD at 530 nm (OD530) by N-acetylgalactosamine mass (mNgal in µg) in the standard samples using appropriate software (SciDavis, Excel, etc.)
- Determine the regression equation with the following equation and the value of the correlation factor, α:

- Determine the concentration of [Ngal] in mM in the sample i using the following equation:

where,
ω is the mass weight of N-acetylgalactosamine, 221.2090 g mol-1,
Vi is the volume of the sample supernatant i, usually from 0.2 ml to 0.450 ml depending on the step C2.
Notes
- We recommend extreme carefulness during this essay since i) the protocol requires usage of concentrated HCl, ii) the acidic solutions are heated over 100 °C and the Eppendorf tube may open, iii) mixing the Na2CO3 and acidic solutions leads to a strong bubbling, and thus liquid may overflow out of the tubes. Hence, working under a fume hood and using gloves is required. Hence, we recommend usage of microcentrifuge tube bridges to avoid opening of the tubes during heat bath.
- Using two dry bath heaters with 2 heat blocks each is convenient to avoid temperature adjustment between the different protocol steps and also for running the standard samples plus the replicates.
Recipes
- Minimal medium
20 mM sodium L-glutamic acid monohydrate (carbon and energy source)
Salt composition:
1.25 x 10-4 g/L FeSO4·7H2O,
0.5 g/L (NH4)2SO4
0.05 g/L MgSO4·7H2O
3.4 g/L KH2PO4
The pH is adjusted to 7 with 10 N KOH
Note: Sodium L-glutamic acid monohydrate is the sole source of carbon and energy in Minimal medium (Plener et al., 2010). - 5 M NaCl solution (10 ml)
2.92 g NaCl
Milli-Q water
Dissolve 2.92 g NaCl in Milli-Q water using a 10 ml volumetric flask - N-acetylgalactosamine standard stock solution at 500 µg ml-1 concentration (100 ml)
50 mg N-acetylgalactosamine
100 ml Milli-Q water
Dissolve 50 mg N-acetylgalactosamine in Milli-Q water using a 100 ml volumetric flask. Prepare 1 ml aliquots in 2.0 ml microcentrifuge tubes and store at -20 °C until use - Standard samples of N-acetyl galactosamine (10 samples with different concentration)
N-acetyl galactosamine standard stock solution at 500 µg ml-1 concentration
Milli-Q water
Prepare the standards samples by diluting various quantity of the N-acetylgalactosamine standard stock solution at 500 µg ml-1 concentration with Milli-Q water to reach 0.450 ml final volume, see Table 1 - 2% acetyl acetone, 1.5 M Na2CO3 solution (20 ml)
3.18 g of Na2CO3
19.6 ml Milli-Q water
0.4 ml acetyl acetone
Dissolve 3.18 g of Na2CO3 in 19.6 ml Milli-Q water in a 50 ml Falcon tube. Then, add 0.4 ml acetyl acetone - Erlich’s reagent (12 ml, for 24 samples)
0.4 g 4-(dimethylamino)benzaldehyde (Erlich’s reagent)
6.0 ml 99.8% ethanol
6.0 ml HCl (37%)
Under a fume hood, dissolve the 4-(dimethylamino)benzaldehyde into 6 ml ethanol 99.8%. Then, carefully, add 6.0 ml of 37% HCl. Make fresh, just before use and protect from the light
Acknowledgments
This work was supported by EMBO (Long-Term Fellowship ALTF 1627-2011), Marie Curie Actions (EMBOCOFUND2010, GA-2010-267146), the Institut National de la Recherche Agronomique (Plant Health Division grant AAP SPE 2012) and the French Laboratory of Excellence project TULIP (ANR-10-LABX-41; ANR-11-IDEX-0002-02).
References
- Balzaretti, S., Taverniti, V., Guglielmetti, S., Fiore, W., Minuzzo, M., Ngo, H. N., Ngere, J. B., Sadiq, S., Humphreys, P. N. and Laws, A. P. (2017). A novel rhamnose-rich hetero-exopolysaccharide isolated from Lactobacillus paracasei DG activates THP-1 human monocytic cells. Appl Environ Microbiol 83(3) pii: e02702-16.
- Brumbley, S. M. and Denny, T. P. (1990). Cloning of wild-type Pseudomonas solanacearum phcA, a gene that when mutated alters expression of multiple traits that contribute to virulence. J Bacteriol 172(10): 5677-5685.
- Drigues, P., Demery-Lafforgue, D., Trigalet, A., Dupin, P., Samain, D. and Asselineau, J. (1985). Comparative studies of lipopolysaccharide and exopolysaccharide from a virulent strain of Pseudomonas solanacearum and from three avirulent mutants. J Bacteriol 162(2): 504-509.
- Elson, L. A. and Morgan, W. T. (1933). A colorimetric method for the determination of glucosamine and chondrosamine. Biochem J 27(6): 1824-1828.
- Flavier, A. B., Clough, S. J., Schell, M. A. and Denny, T. P. (1997). Identification of 3-hydroxypalmitic acid methyl ester as a novel autoregulator controlling virulence in Ralstonia solanacearum. Mol Microbiol 26(2): 251-259.
- Gatt, R. and Berman, E. R. (1966). A rapid procedure for the estimation of amino sugars on a micro scale. Anal Biochem 15(1): 167-171.
- Lee, M. J., Liu, H., Barker, B. M., Snarr, B. D., Gravelat, F. N., Al Abdallah, Q., Gavino, C., Baistrocchi, S. R., Ostapska, H., Xiao, T., Ralph, B., Solis, N. V., Lehoux, M., Baptista, S. D., Thammahong, A., Cerone, R. P., Kaminskyj, S. G., Guiot, M. C., Latge, J. P., Fontaine, T., Vinh, D. C., Filler, S. G. and Sheppard, D. C. (2015). The fungal exopolysaccharide galactosaminogalactan mediates virulence by enhancing resistance to neutrophil extracellular Traps. PLoS Pathog 11(10): e1005187.
- Peyraud, R., Cottret, L., Marmiesse, L., Gouzy, J. and Genin, S. (2016). A resource allocation trade-off between virulence and proliferation drives metabolic versatility in the plant pathogen Ralstonia solanacearum. PLoS Pathog 12(10): e1005939.
- Plener, L., Manfredi, P., Valls, M. and Genin, S. (2010). PrhG, a transcriptional regulator responding to growth conditions, is involved in the control of the Type III secretion system regulon in Ralstonia solanacearum. J Bacteriol 192(4): 1011-1019.
- Salanoubat, M., Genin, S., Artiguenave, F., Gouzy, J., Mangenot, S., Arlat, M., Billault, A., Brottier, P., Camus, J. C., Cattolico, L., Chandler, M., Choisne, N., Claudel-Renard, C., Cunnac, S., Demange, N., Gaspin, C., Lavie, M., Moisan, A., Robert, C., Saurin, W., Schiex, T., Siguier, P., Thebault, P., Whalen, M., Wincker, P., Levy, M., Weissenbach, J. and Boucher, C. A. (2002). Genome sequence of the plant pathogen Ralstonia solanacearum. Nature 415(6871): 497-502.
- Vaningelgem, F., Zamfir, M., Mozzi, F., Adriany, T., Vancanneyt, M., Swings, J. and De Vuyst, L. (2004). Biodiversity of exopolysaccharides produced by Streptococcus thermophilus strains is reflected in their production and their molecular and functional characteristics. Appl Environ Microbiol 70(2): 900-912.
Article Information
Copyright
© 2017 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Peyraud, R., Denny, T. P. and Genin, S. (2017). Exopolysaccharide Quantification for the Plant Pathogen Ralstonia solanacearum. Bio-protocol 7(10): e2289. DOI: 10.21769/BioProtoc.2289.
Category
Plant Science > Plant immunity > Disease bioassay
Plant Science > Plant immunity > Host-microbe interactions
Cell Biology > Cell isolation and culture > Cell differentiation
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










