- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Locomotor Assay in Drosophila melanogaster
Published: Vol 7, Iss 10, May 20, 2017 DOI: 10.21769/BioProtoc.2283 Views: 10296
Reviewed by: Xi FengManuel SarmientoAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

Caste Transition and Reversion in Harpegnathos saltator Ant Colonies
Comzit Opachaloemphan [...] Hua Yan
Aug 20, 2023 1576 Views

A Method for Studying Social Signal Learning of the Waggle Dance in Honey Bees
Shihao Dong [...] Ken Tan
Aug 20, 2023 1382 Views
Habituation of Sugar-Induced Proboscis Extension Reflex and Yeast-Induced Habituation Override in Drosophila melanogaster
Swati Trisal [...] Mani Ramaswami
Dec 5, 2023 1450 Views
Abstract
This protocol describes a simple locomotor assay in Drosophila melanogaster. In brief, the locomotor of each single fly in the culture dish is recorded by a web camera. The moving time, walking length, speed and the locomotor trails of the single fly could be quantitatively analyzed.
Keywords: LocomotorBackground
This protocol was implemented in the previously published study (Liu et al., 2016). In that study, this assay was combined with the optogenetic system by simply providing the proper excitation light.
Materials and Reagents
- 3.5 cm culture dishes with black paper inside
- Drosophila strains to be analyzed
- Ice for anesthesia
Equipment
- Empty vial for cold anesthesia
- Fluorescent lamp (Bannet T5 8 W, China)
- Infrared LEDs (TUOENS, model: TS-6036A )
- Web camera (Omiky, model: CEL USB 2.0 50.0M PC Camera , catalog number: CEL9002255) with the IR filter removed; Replacing the IR filter with the floppy disk (the black floppy disk contained in the hard shell) to filter the visible light
- Behavior room with the temperature and humidity controlled
Software
- MATLAB (MathWorks, R2013a)
Procedure
- Set the environmental illuminance (provided by the fluorescent lamp) to be 1,000-1,300 Lux, the temperature to be 24 ± 1 °C and the humidity to be 40-60%.
- Transfer the flies into the empty vial and anesthetize them on ice for 10 min.
- Transfer one fly to each culture dish, and let the flies recover from the anesthesia for at least 10 min.
- Record the locomotion of the flies for 2 min with the web camera equipped with a visible light filter (Figures 1A and 1D; Video1). The fly is light up and the background is dark (Figure 1B). The angle of the infrared LEDs should be adjusted before recording to reduce the reflection on the dish.
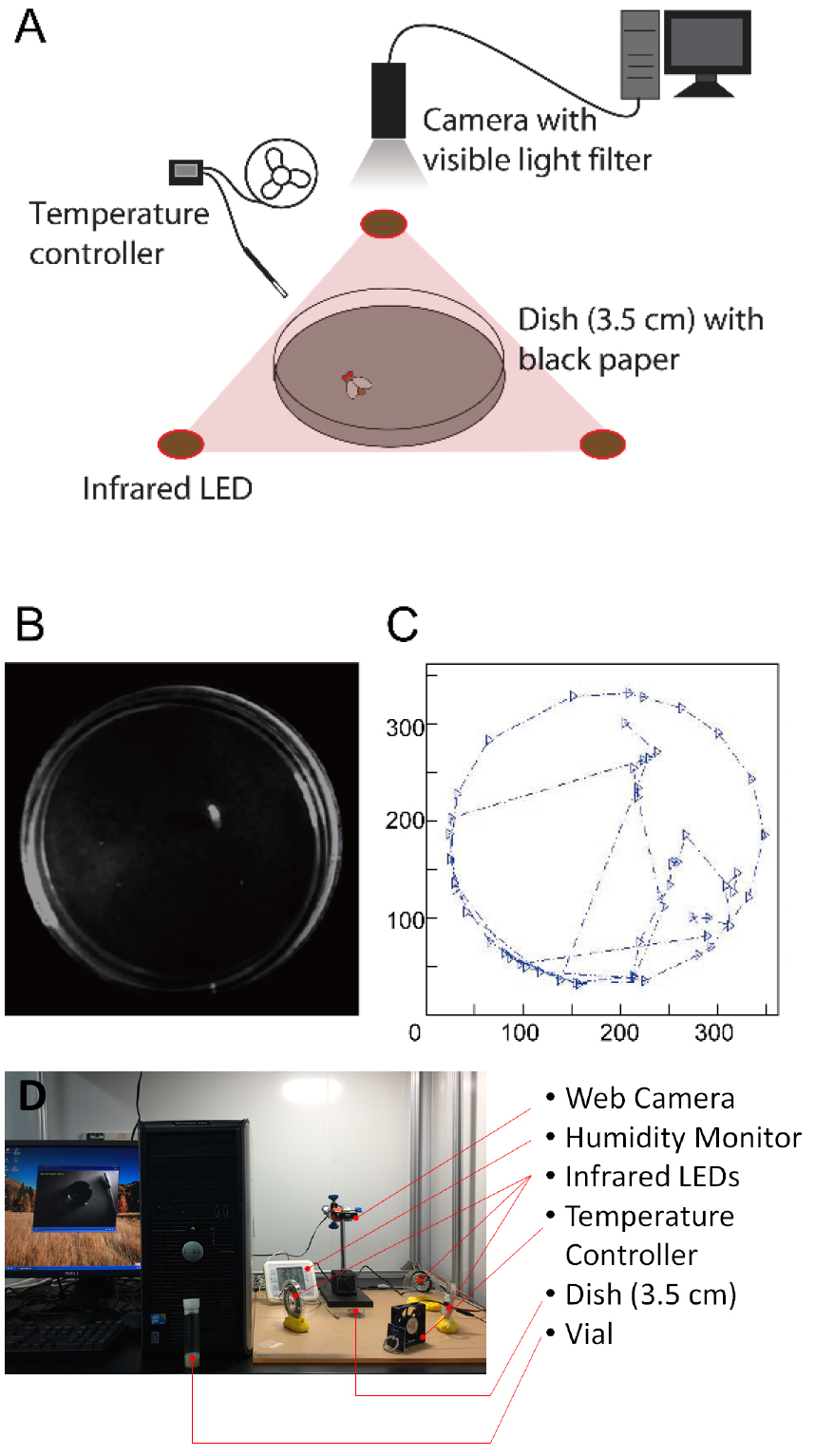
Figure 1. Locomotor assay in Drosophila melanogaster. A. The schematic diagram of the locomotor assay; B. A photo of the dish with a fly inside; C. The motion trial of a recorded fly; D. A photo of the setup.
Video 1. Recording of the locomotor behavior of the fliesData analysis
- Analyze the photos with MATLAB (MathWorks, R2013a; For MATLAB code for video analysis, see Supplemental file 1) (Figure 1C). Figure out the coordinate of the fly on each photo, and then calculate the moving time, walking length and speed of the fly, and analyze the locomotor trail of the fly.
- Individuals with the walk length less than half of the fly body length during recording should be excluded.
- The Wilcoxon signed rank test should be applied to evaluate differences between matched samples.
Acknowledgments
We would like to thank Jingwu Hou for assistance with experimental setup. This protocol was designed by Q. L. and was implemented in the previously published study (Liu et al., 2016). This study was supported by the ‘Strategic Priority Research Program’ of the CAS (XDB02040004), by grants from the 973 Program (2011CBA00400), as well as by the National Science Foundation of China (91232000, 91132709, 31130027, and 31070956). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
References
- Liu, Q., Yang, X., Tian, J., Gao, Z., Wang, M., Li, Y. and Guo, A. (2016). Gap junction networks in mushroom bodies participate in visual learning and memory in Drosophila. Elife 5.
- Analyze the photos with MATLAB (MathWorks, R2013a; For MATLAB code for video analysis, see Supplemental file 1) (Figure 1C). Figure out the coordinate of the fly on each photo, and then calculate the moving time, walking length and speed of the fly, and analyze the locomotor trail of the fly.
Article Information
Copyright
Liu et al. This article is distributed under the terms of the Creative Commons Attribution License (CC BY 4.0).
How to cite
Readers should cite both the Bio-protocol article and the original research article where this protocol was used:
- Liu, Q., Tian, J., Yang, X., Li, Y. and Guo, A. (2017). Locomotor Assay in Drosophila melanogaster. Bio-protocol 7(10): e2283. DOI: 10.21769/BioProtoc.2283.
- Liu, Q., Yang, X., Tian, J., Gao, Z., Wang, M., Li, Y. and Guo, A. (2016). Gap junction networks in mushroom bodies participate in visual learning and memory in Drosophila. Elife 5.
Category
Neuroscience > Behavioral neuroscience > Learning and memory
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










