- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Escherichia coli Infection of Drosophila
Published: Vol 7, Iss 9, May 5, 2017 DOI: 10.21769/BioProtoc.2256 Views: 9467
Reviewed by: Ruth A. FranklinModesto Redrejo-RodriguezFilipa Vaz

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
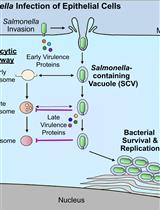
Bacterial Pathogen-mediated Suppression of Host Trafficking to Lysosomes: Fluorescence Microscopy-based DQ-Red BSA Analysis
Mădălina Mocăniță [...] Vanessa M. D'Costa
Mar 5, 2024 2932 Views
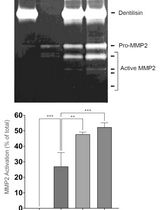
Purification of Native Dentilisin Complex from Treponema denticola by Preparative Continuous Polyacrylamide Gel Electrophoresis and Functional Analysis by Gelatin Zymography
Pachiyappan Kamarajan [...] Yvonne L. Kapila
Apr 5, 2024 2125 Views
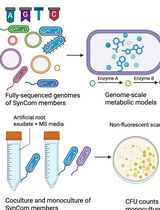
In Silico Prediction and In Vitro Validation of Bacterial Interactions in the Plant Rhizosphere Using a Synthetic Bacterial Community
Arijit Mukherjee [...] Sanjay Swarup
Nov 5, 2025 1710 Views
Abstract
Following septic insults, healthy insects, just like vertebrates, mount a complex immune response to contain and destroy pathogens. The failure to efficiently clear bacterial infections in immuno-compromised fly mutants leads to higher mortality rates which provide a powerful indicator for genes with important roles in innate immunity. The following protocol is designed to reproducibly inject a known amount of non-pathogenic E. coli into otherwise sterile flies and to measure the survival of flies after infection. The protocol can be easily adapted to different types of bacteria.
Keywords: DrosophilaBackground
Classic infection experiments involve infecting Drosophila orally (Chakrabarti et al., 2016) or with a needle dipped in a concentrated bacterial solution (Romeo and Lemaitre, 2008). Unlike these protocols, our experimental procedure allows us to determine the site of infection and precisely control the dose of bacteria injected into each fly. This provides homogeneity and reproducibility, and allows us to adapt bacterial load for different experiments (Akbar et al., 2011 and 2016).
Materials and Reagents
- 15 ml Falcon tubes (Corning, catalog number: 352196 )
- 1.5 ml Eppendorf tubes (USA Scientific, catalog number: 1615-5500 )
- Kimwipes
- 26 G needle (BD, catalog number: 305111 )
- Spin columns (e.g., Thermo Fisher Scientific, Thermo ScientificTM, catalog number: 69700 )
- Empty and clean Drosophila vials (Genesee Scientific, catalog number: 32-109 )
- Microloader tips (Eppendorf, catalog number: 930001007 )
- Vials (Genesee Scientific, catalog number: 32-109 ) with standard Drosophila food (lightly yeasted)
- Microslide (Corning, catalog number: 2948-75X25 )
- 50 Drosophila melanogaster adult virgins (Romeo and Lemaitre, 2008), aged five to seven days post eclosion (see Note 1)
- E. coli-DH5α containing any ampicillin-resistant plasmid (e.g., expressing GFP)
- pEGFP (https://www.addgene.org/vector-database/2485/)
- pET-GFP-C11 (http://www.addgene.org/30183/)
- LB broth (Fisher Scientific, catalog number: BP1426-500 )
- Ampicillin (200 mg/ml) (Sigma-Aldrich, catalog number: A9518 )
- 70% ethanol–diluted from 100% ethanol (Pharmco-AAPER, catalog number: 111000200 )
- Mineral oil (Fisher Scientific, catalog number: O121-1 )
- Sodium chloride (NaCl) (Fisher Scientific, catalog number: S271 )
- Potassium chloride (KCl) (Sigma-Aldrich, catalog number: P9541 )
- Sodium phosphate dibasic heptahydrate (Na2HPO4·7H2O) (Sigma-Aldrich, catalog number: S9390 )
- Potassium phosphate monobasic (KH2PO4) (Fisher Scientific, catalog number: S397-500 )
- 1x PBS (see Recipes)
Equipment
- 37 °C incubator with shaking (Eppendorf, New BrunswickTM, model: Innova® 44 )
- Spectrophotometer (Molecular Devices, model: SpectraMax M2 )
- Borosilicate glass capillaries (WPI, catalog number: TW100-4 )
- Centrifuge capable of spinning Falcon tubes (Eppendorf, model: 5804 R )
- Flaming/Brown Micropipette puller (Sutter Instrument, model: P-97 )
- Mini benchtop centrifuge (Fisher Scientific, model: FisherbrandTM Standard Mini Centrifuge , catalog number: 05-090-100)
- Pico-Injector (Nikon Instruments, catalog number: PLI-188 )–requires nitrogen gas
Note: Should have foot pedal for injecting. - 25 °C incubator (BioCold Environment, model: BC49-IN )
- Anesthetizing fly pad (Genesee Scientific, catalog number: 59-119 )
- Small brush
- Dissecting microscope (Leica)
Software
- Prism (GraphPad) software
Procedure
- Preparation of E. coli suspension
- Inoculate 50 ml LB broth with an E. coli colony from an LB agar plate under sterile conditions.
- Add ampicillin (200 µg/ml final).
- Incubate overnight at 37 °C with shaking. Use spectrophotometer to measure optical density at 600 nm (OD600).
- Dilute 500 µl aliquot of bacterial over-night suspension 5-fold with 2 ml of LB broth
- Measure OD600 of diluted bacteria with broth, and broth alone.
- Subtract OD600 of fresh LB broth from bacteria with broth and calculate OD600 of over-night culture.
- Dilute over-night 50 ml bacterial culture to OD600 = 0.1 with LB broth.
- Spin 50 ml of bacterial suspension 4,000 x g for 10 min in Falcon tubes at room temperature.
- Decant LB broth.
- Re-suspend in 50 ml of 1x PBS by vortexing.
- Adjust with PBS to OD600 = 0.1.
- Heat-Kill bacteria (optional; see Note 2)
- Aliquot 1 ml bacterial suspension into 1.5 ml Eppendorf tubes.
- Incubate at 65 °C for 20 min.
- Bacterial solution is now ready for injections.
- Preparation of injection needles
- Place glass capillary in Micropipette puller.
- Run program to create needles tapered like the needle pictured (Figure 1).
- Ramp + 5, Pull = 90, Velocity = 70, Delay = 70
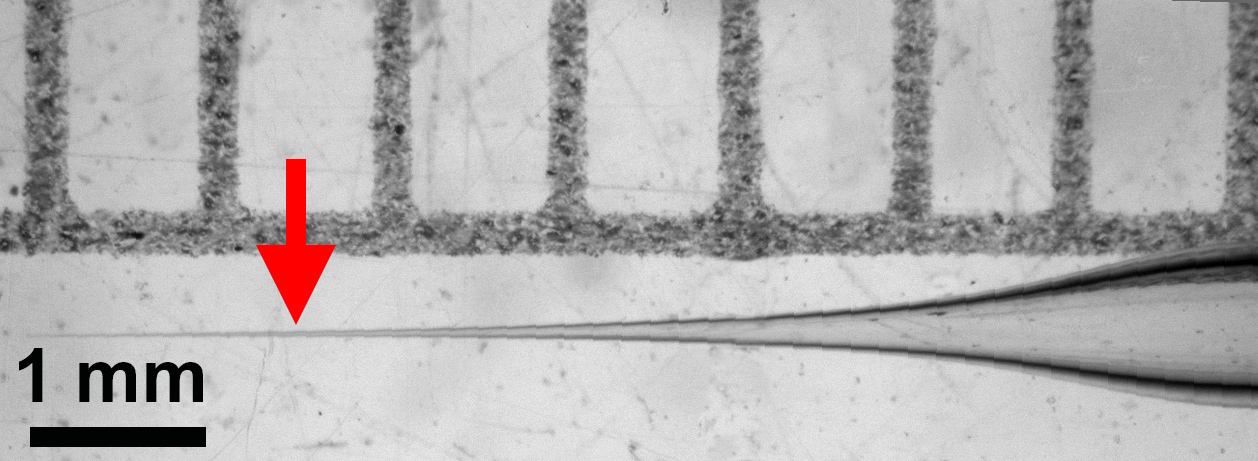
Figure 1. Tapered injection needle. Arrow indicates preferred place to break it. - Stretch a Kimwipe taut over a Falcon tube and stab needle through the Kimwipe to break the very tip of the needle.
- Fly sterilization
- Use a small needle (26 G) to poke three to four holes in the paper filter of the spin columns and place back into collection tube.
- Add 25 anesthetized flies to one spin column (Figure 2).
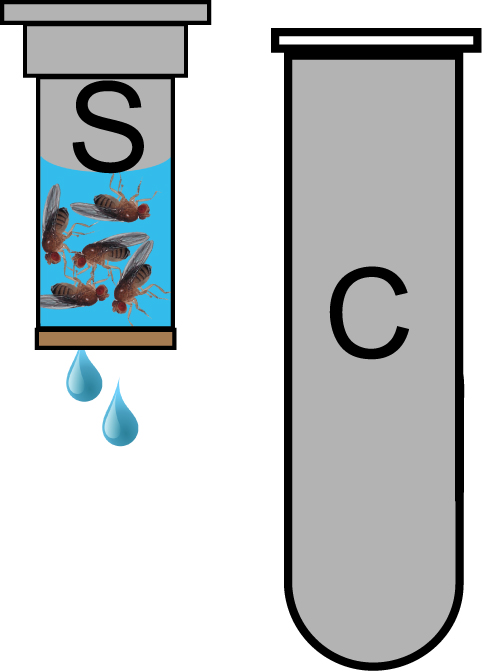
Figure 2. Spin column for Fly sterilization. 25 flies are loaded into a spin column (S) and covered with 70% ethanol. Spin columns are placed in a collection tube (C) to contain the flow through of wash solutions in the centrifuge during the quick spin. Flies are retained in the spin column by the paper filter at its bottom. - Add 700 μl 70% ethanol
- Quick spin on mini benchtop centrifuge (~3-4 sec).
- Remove spin column from collection tube and pour out flow through and place spin column back into collection tube.
- Repeat ethanol wash steps 3c-3e.
- Wash with 700 μl sterile water.
- Quickspin on mini benchtop centrifuge (~3-4 sec).
- Repeat water rinse steps 3g and 3h.
- Fold Kimwipe and place in empty fly vial.
- Put flies into vial with Kimwipe, plug and place at 25 °C until flies have recovered.
- Calibration of injection volume (see Note 3)
- Load 1 μl bacterial solution into the pulled needle using Microloader tips.
- Attach needle to Pico-injector.
- Adjust Pico-injector pressures, injection time to achieve 50 nl per injection.
- Typical starting values for our set-up:
1) Balance pressure = -0.2 psi–Should be neutral .
2) Injection pressure = 15 psi.
3) Injection time = 2 msec. - For calibration: put a drop of mineral oil on a microslide.
- Inject bacterial solution into oil to determine nl per injection (a needle loaded with 1 µl should yield 20 equal sized droplets of 50 nl each).
- Adjust injection time and pressure until desired amount is injected.
- Optimal Pico-injector values will vary slightly between needles.
- Injection of sterilized flies (see Notes 4 and 5)
- Sterilize fly pad with 70% ethanol, allow to dry.
- Place sterilized flies on fly pad.
- Under dissecting microscope hold a fly in place by its legs with a small brush.
- Hold injecting rod and needle with other hand.
- Insert needle into the notum of the immobilized fly (Figure 3).
- Inject bacterial solution into fly using foot pedal–Audible beep will confirm.
- Repeat until all flies have been injected.
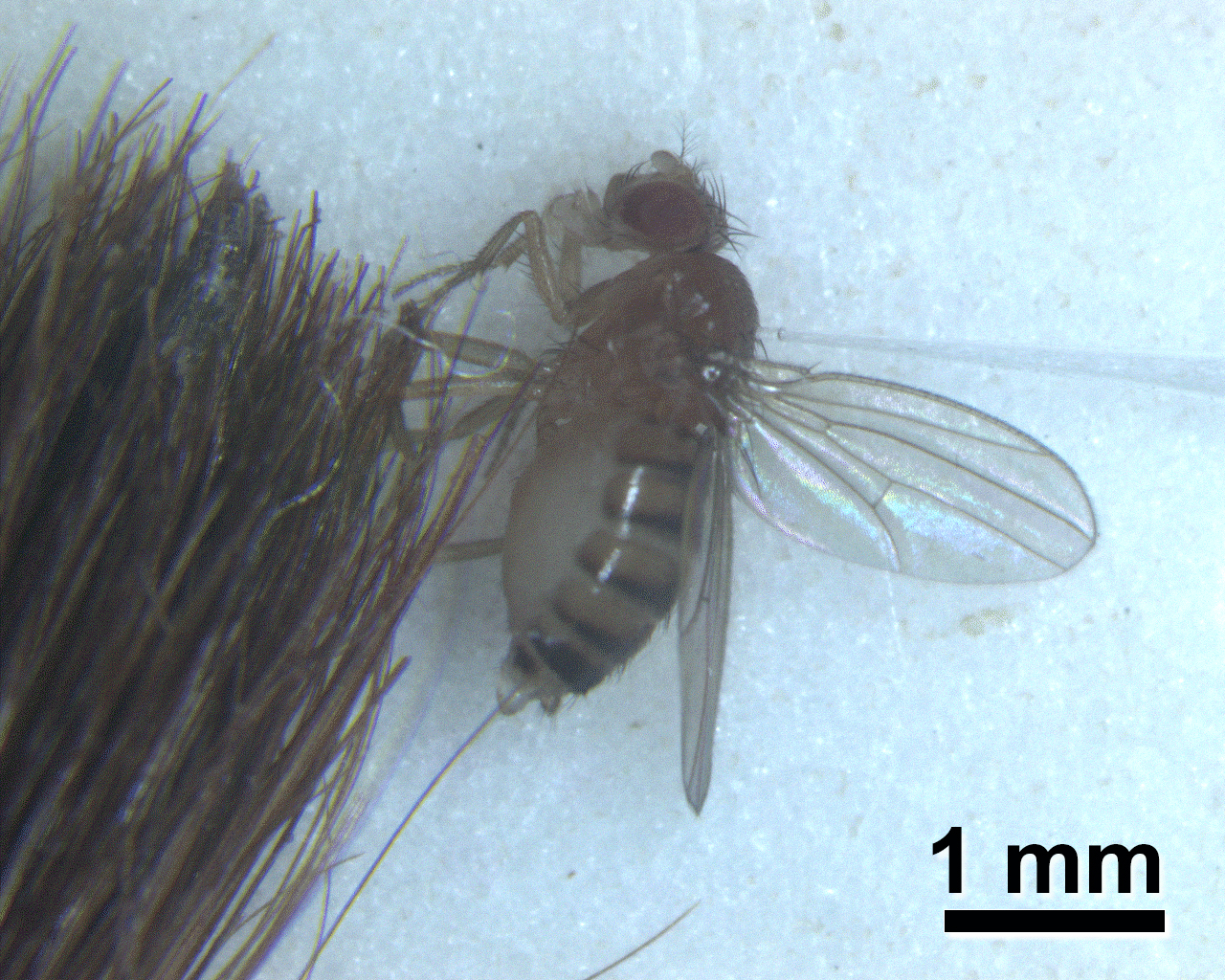

Figure 3. Injection of flies with bacterial suspension. Anesthetized flies are held with a brush as they are injected using a pulled glass needle (see Figure 1) filled with a bacterial suspension. - Incubate flies and monitor survival
- Place ≤ 15 injected flies into vial containing fresh standard Drosophila media.
Vials should be lightly yeasted and contain no more than 1.5 ml of fly food. This minimizes the danger of flies getting stuck in yeast or between the side of the vial and the food as the food tends to dry at later days and separate from the vial. - Place vials with flies into 25 °C incubator and note how many flies are alive 2 h after injection. Flies that do not recover from injections should be discarded from the dataset and subsequent analysis.
- Count dead and living flies every day, once a day until all are dead. For bacteria more pathogenic than E. coli, shorter time intervals will be important.
- Flip flies into fresh vials every 5 days.
Data analysis
Recorded data are entered into Prism (GraphPad) software which is used to display survival curves (Figure 4) and analyze statistical significance of differences in survival between genotypes. Log-Rank (Mantel-Cox) test was used to assess statistical significance of differences between survival curves (Figure 4). Alternatively, multiple Log-rank analysis software packages are available online at: http://astatsa.com/ and http://bioinf.wehi.edu.au/software/russell/logrank. Both sides offer detailed information on how to enter data.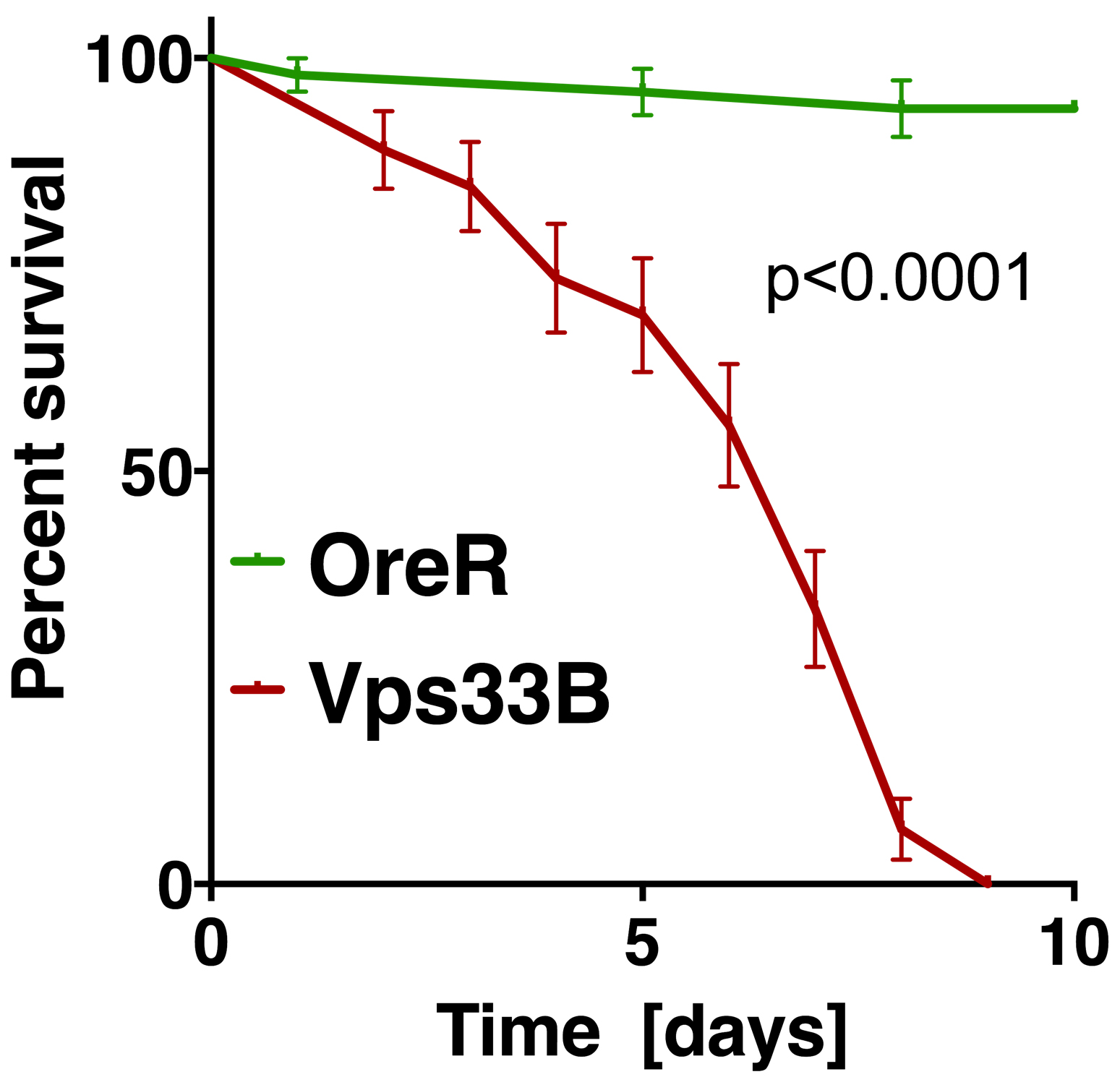
Figure 4. Example of statistical analysis of fly survival post bacterial infection. The diagram shows the fraction of surviving wild-type flies (OreR) or Vps33B mutants after injection with heat-killed E. coli, similar to experiments previously published (Akbar et al., 2016). Statistical analysis was performed in Prism using the Log-rank (or Mantel-Cox) test to determine a P value smaller than 0.0001.
Notes
- A wild-type Drosophila strain (Oregon R or Canton S) should be included as a negative control group for injections. Use sibling and/or parent flies as controls when appropriate. Three replicates of fifty flies per genotype are sufficient for proper statistical analysis.
- It is important to note that heat inactivation can change the potential of bacteria to engage different innate immunity pathways as normally hidden epitopes may be exposed (Chung and Kocks, 2011).
- Most difficulties in this experimental procedure are encountered during the needle calibration and injection steps. The balance pressure is the source of most problems. If the balance pressure is set too high, the result is a constant bleeding of the bacterial solution resulting in difficulty measuring injection droplets, emptying of the needle, and dripping bacterial solution onto the fly. No liquid should escape the needle until the injection foot pedal has been pressed. Problems also arise if the balance pressure is too low. Low pressure will result in sucking up liquid that had just been injected resulting in little to no bacteria being injected (this should also be part of your calibration). Furthermore, once the needle is inserted into the fly, the negative pressure will pull fly tissues into the needle tip resulting in blockage that can often be seen through the microscope. This will prevent any further injections when the foot pedal has been pushed. Placing the needle tip into the bacterial solution and pressing the ‘clear’ button on the Pico-injector can usually solve this by high positive pressure clearing the needle. Once the blockage is cleared, leave the needle tip in the bacterial solution and hold the ‘fill’ button to refill the needle using negative pressure. Once the needle is calibrated it is also useful to inject a few droplets on or near a fly or paintbrush to get a good sense of the size of a 50 nl droplet. A 50 nl droplet is roughly the size of a fly eye.
- Prior to commencing bacterial injections, it is important to have a series of experiments showing flies do not die from inflammation resulting from the needle injury or from the injection media. Mock injections should be performed on wild-type and experimental flies to determine if injury alone is sufficient to kill flies. Furthermore, control injections of sterile PBS should be done to ensure flies die from infection. These mock injections should be analyzed in the same manner as injections of bacteria as described in the data analysis section.
- The experimental procedure described above has been optimized for DH5α E. coli. Each 50 nl injection should deliver around 2,000 CFU to each fly. If other bacterial species are to be used, bacterial loads should be optimized to ensure survival of wild-type flies when virulence allows.
Recipes
- 1x PBS
137 mM NaCl
2.7 mM KCl
10 mM Na2HPO4
1.8 mM KH2PO4
Acknowledgments
The work herein was supported by NIH Grants EY010199, EY021922.
This protocol has been adapted and modified from our previously published work (Akbar et al., 2016; Akbar et al., 2011).
The authors declare no conflicts of interest.
References
- Akbar, M. A., Mandraju, R., Tracy, C., Hu, W., Pasare, C. and Kramer, H. (2016). ARC syndrome-linked Vps33B protein is required for inflammatory endosomal maturation and signal termination. Immunity 45(2): 267-279.
- Akbar, M. A., Tracy, C., Kahr, W. H. and Kramer, H. (2011). The full-of-bacteria gene is required for phagosome maturation during immune defense in Drosophila. J Cell Biol 192(3): 383-390.
- Chakrabarti, S., Dudzic, J. P., Li, X., Collas, E. J., Boquete, J. P. and Lemaitre, B. (2016). Remote control of intestinal stem cell activity by haemocytes in Drosophila. PLoS Genet 12(5): e1006089.
- Chung, Y. S. and Kocks, C. (2011). Recognition of pathogenic microbes by the Drosophila phagocytic pattern recognition receptor Eater. J Biol Chem 286(30): 26524-26532.
- Romeo, Y. and Lemaitre, B. (2008). Drosophila immunity: methods for monitoring the activity of Toll and Imd signaling pathways. Methods Mol Biol 415: 379-394.
Article Information
Copyright
© 2017 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Tracy, C. and Krämer, H. (2017). Escherichia coli Infection of Drosophila. Bio-protocol 7(9): e2256. DOI: 10.21769/BioProtoc.2256.
Category
Immunology > Animal model > Drosophila (Fruit fly)
Microbiology > in vivo model > Bacterium
Microbiology > Microbe-host interactions > Bacterium
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










